Computational Analysis of the Taxanomical Classification of Short











- Slides: 22

Computational Analysis of the Taxanomical Classification of Short 16 S r. RNA Sequences Christel Chehoud Mentor: Brian Haas

Overview l l l Human Microbiome Project 16 S r. RNA Reference and Test Sets Classifiers Accuracy of Classifications Results

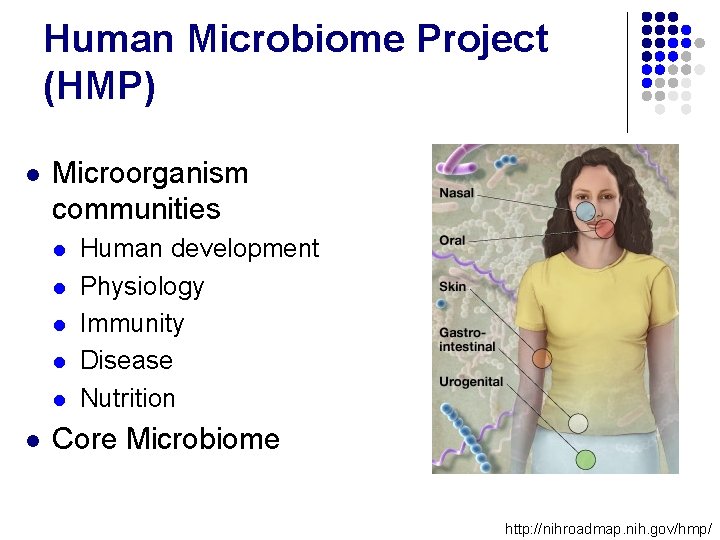
Human Microbiome Project (HMP) l Microorganism communities l l l Human development Physiology Immunity Disease Nutrition Core Microbiome http: //nihroadmap. nih. gov/hmp/

16 S r. RNA l 16 S l Ribosomal RNA l Large RNA component of the small subunit of the ribosome l Phylogenetic Markers l l Species Identification 1542 bp

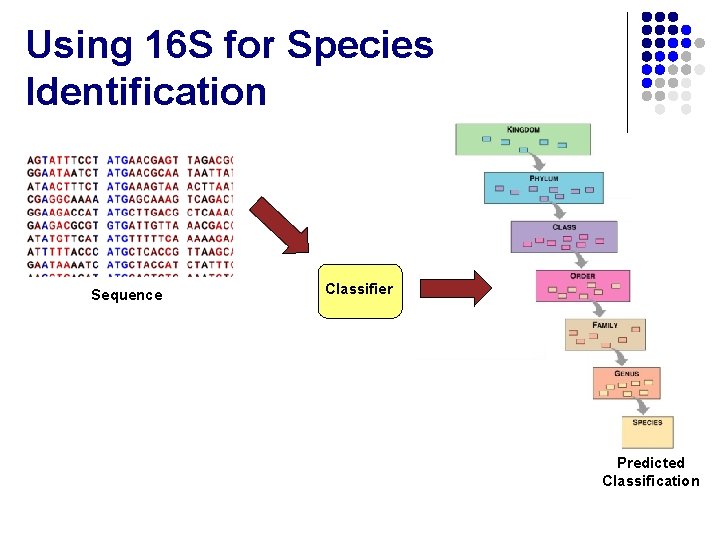
Using 16 S for Species Identification Sequence Classifier Predicted Classification

Project Goal l l New Sequencing Technology Evaluate the accuracy of the classification of the 16 S r. RNA across different: l l l Classifiers Regions of the sequence Phylogeny

Reference Dataset l RDP Core Set l l l l Trusted Taxonomies 6, 621 sequences Phylum: 27 Class: 43 Order: 97 Family: 258 Genus: 1352

Green. Genes’s Full Collection of Sequences l l Full Collection used by Green. Genes High phylogenetic diversity l 188, 073 sequences

Comparison of Taxonomy Predictions by Method l Classified Green. Genes Core Set Using: l l All Match l 188, 073 135, 269 RDP (Naïve Bayesian) kmer. Rank Blast 135, 269 sequences l. Phylum: 27 l. Class: 43 l. Order: 96 l. Family: 257 l. Genus: 1335

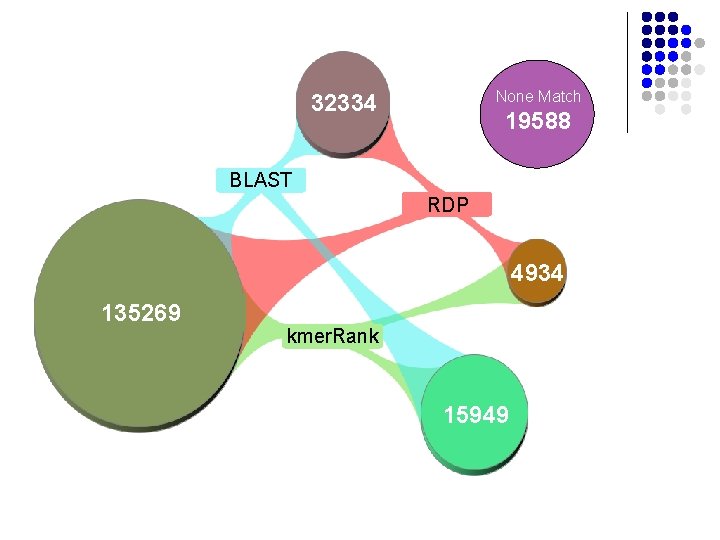
None Match: 19588 None Match 32334 19588 BLAST RDP 4934 135269 kmer. Rank 15949

CD-hit: Normalizing Genus Representation l 3% difference between genera l l l 188, 073 l 21, 179 sequences Phylum: 27 Class: 43 Order: 96 Family: 235 Genus: 1241 135, 269 21, 179 Li, 2006

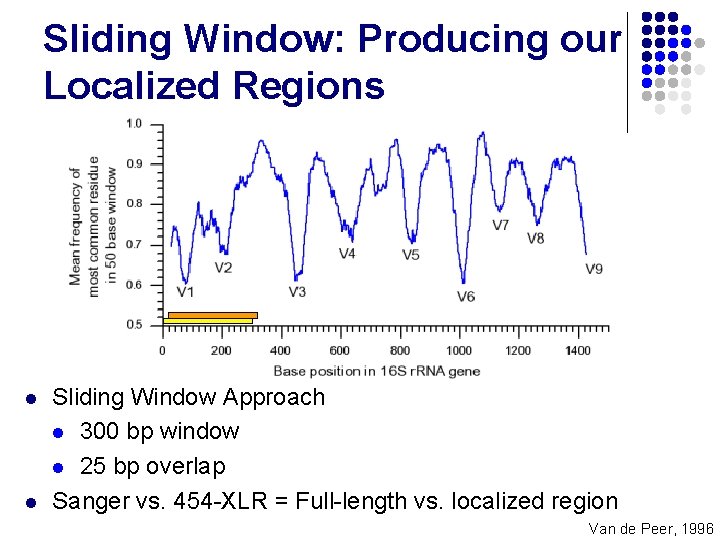
Sliding Window: Producing our Localized Regions l l Sliding Window Approach l 300 bp window l 25 bp overlap Sanger vs. 454 -XLR = Full-length vs. localized region Van de Peer, 1996

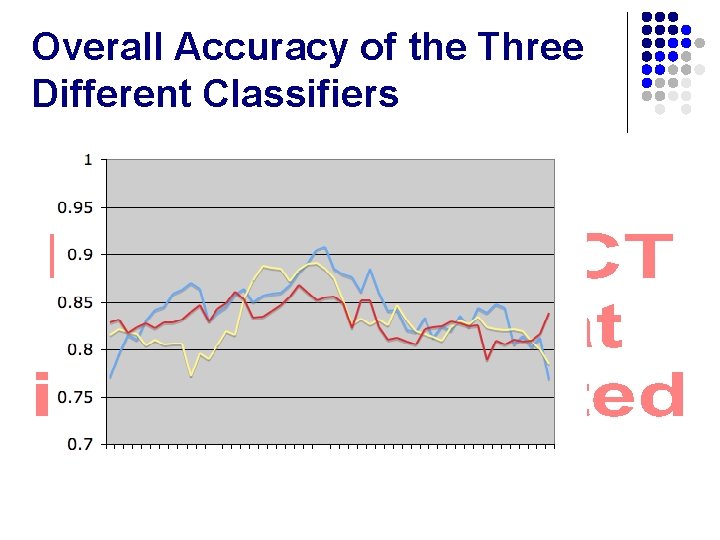
Overall Accuracy of the Three Different Classifiers

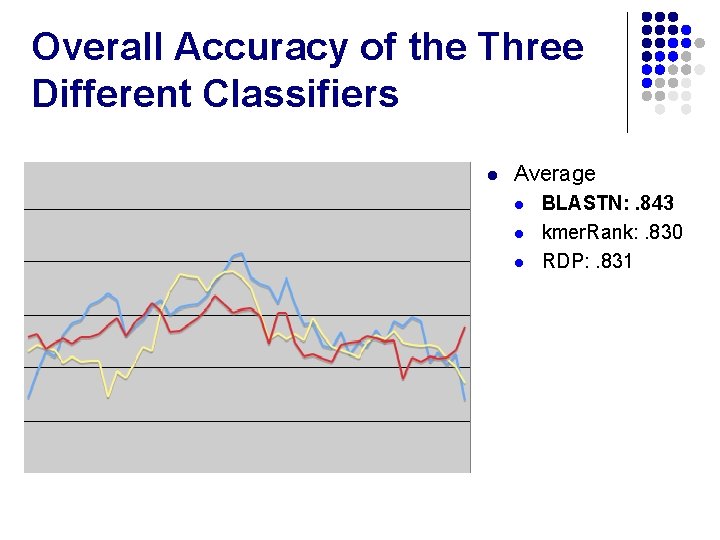
Overall Accuracy of the Three Different Classifiers l Average l l l BLASTN: . 843 kmer. Rank: . 830 RDP: . 831

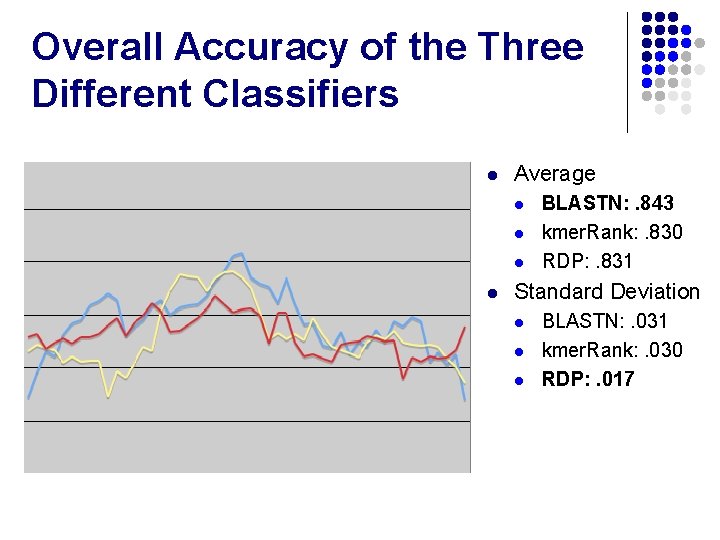
Overall Accuracy of the Three Different Classifiers l Average l l BLASTN: . 843 kmer. Rank: . 830 RDP: . 831 Standard Deviation l l l BLASTN: . 031 kmer. Rank: . 030 RDP: . 017

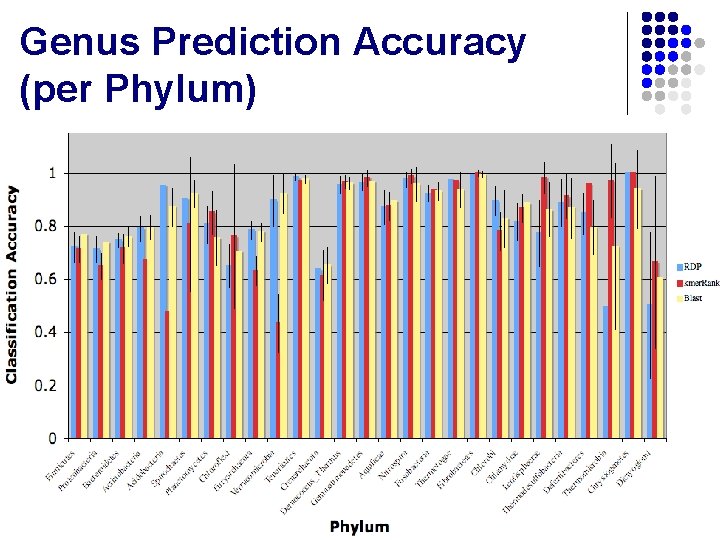
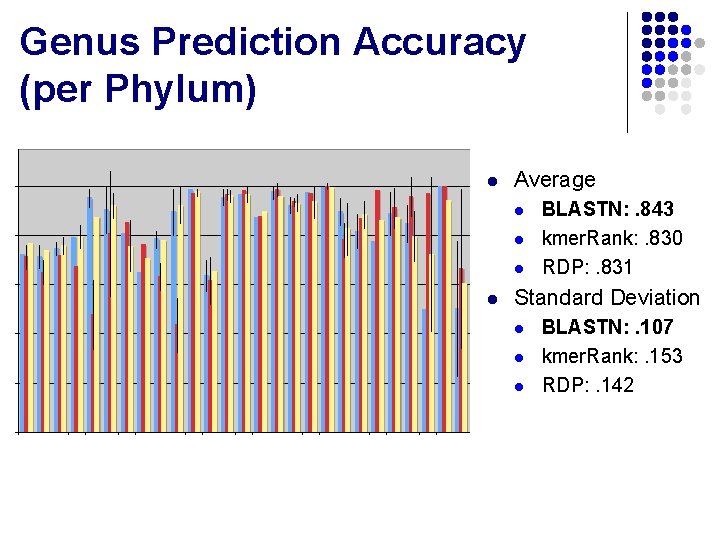
Genus Prediction Accuracy (per Phylum)

Genus Prediction Accuracy (per Phylum) l Average l l BLASTN: . 843 kmer. Rank: . 830 RDP: . 831 Standard Deviation l l l BLASTN: . 107 kmer. Rank: . 153 RDP: . 142

Finding the 16 S Region Providing the Most Reliable Prediction Accuracy

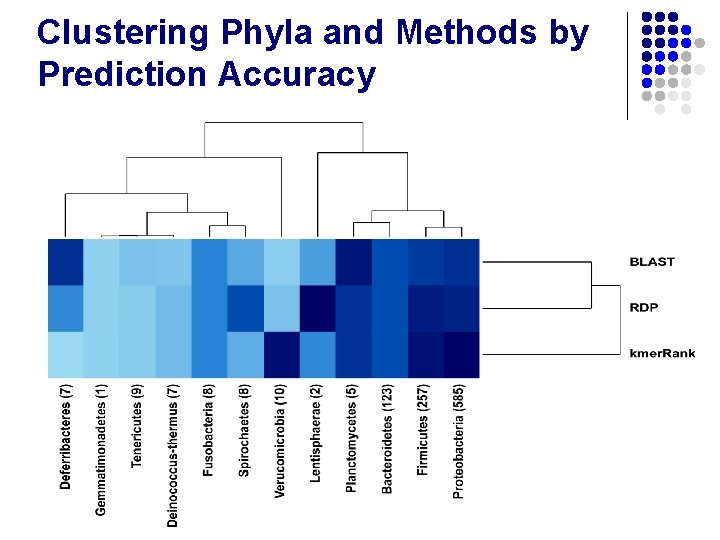
Clustering Phyla and Methods by Prediction Accuracy

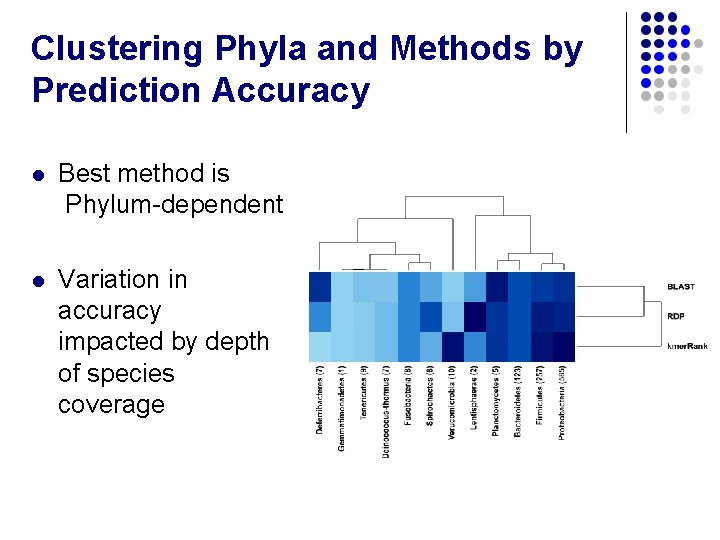
Clustering Phyla and Methods by Prediction Accuracy l Best method is Phylum-dependent l Variation in accuracy impacted by depth of species coverage

Summary l l Central region of 16 S is the most accurate, on average Of the methods examined, BLAST is most accurate across all 16 S regions and all phyla, on average RDP-bayes is least variable across short sequence regions Best short sequence classification method is phylum-dependent

Acknowledgements l Genome Sequencing and Analysis Program l l l l Brian Haas Dirk Gevers Michael Feldgarden Doyle Ward Chad Nusbaum Bruce Birren Administration l l l Shawna Young Lucia Vielma Maura Silverstein