Bioinformatics challenges in nextgeneration sequencing era Valentina Boeva

Bioinformatics challenges in next-generation sequencing era Valentina Boeva
Plan § a little bit about cell biology § a little bit about sequencing § a little bit about NGS applications § bioinformatical challenges in the primary steps data analysis • assembly or alignment of reads • normalization of read counts
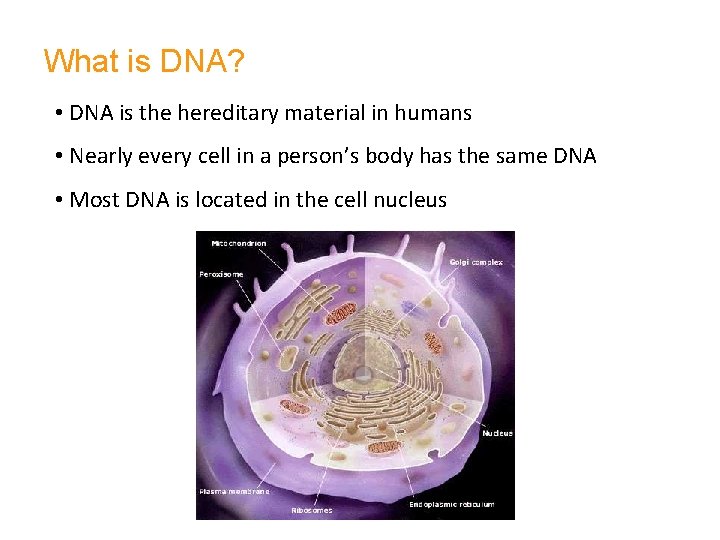
What is DNA? • DNA is the hereditary material in humans • Nearly every cell in a person’s body has the same DNA • Most DNA is located in the cell nucleus

What is DNA? • the information in DNA is stored as a code made up of four chemical bases: A, G, C and T • Human DNA consists of about 3 billion bases organized in chromosomes (46 in Humans) Each chromosome is a very long word in 4 letter alphabet
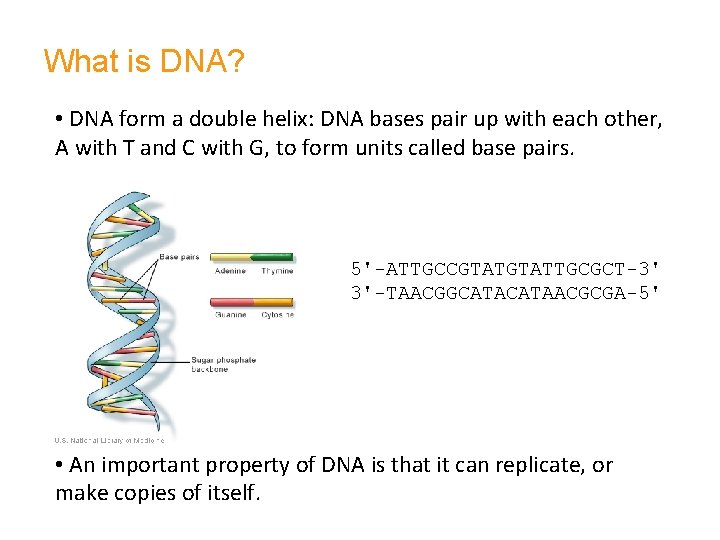
What is DNA? • DNA form a double helix: DNA bases pair up with each other, A with T and C with G, to form units called base pairs. 5'-ATTGCCGTATTGCGCT-3' 3'-TAACGGCATAACGCGA-5' • An important property of DNA is that it can replicate, or make copies of itself.
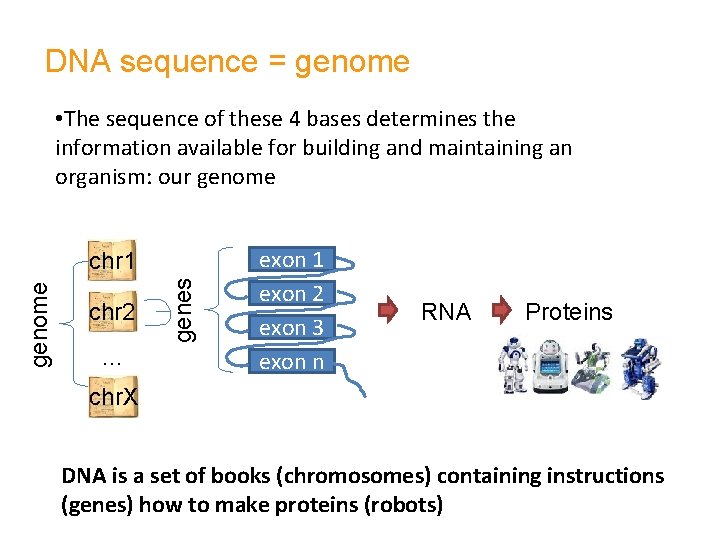
DNA sequence = genome • The sequence of these 4 bases determines the information available for building and maintaining an organism: our genome chr 2 … genes genome chr 1 exon 2 exon 3 exon n RNA Proteins chr. X DNA is a set of books (chromosomes) containing instructions (genes) how to make proteins (robots)
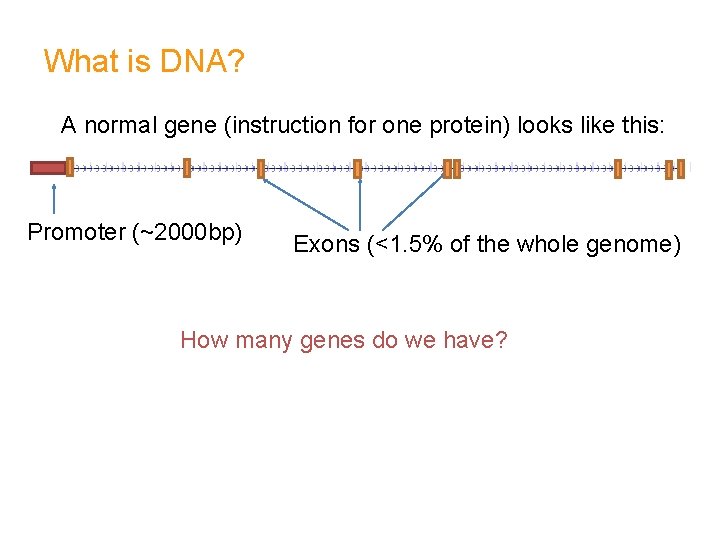
What is DNA? A normal gene (instruction for one protein) looks like this: Promoter (~2000 bp) Exons (<1. 5% of the whole genome) How many genes do we have?
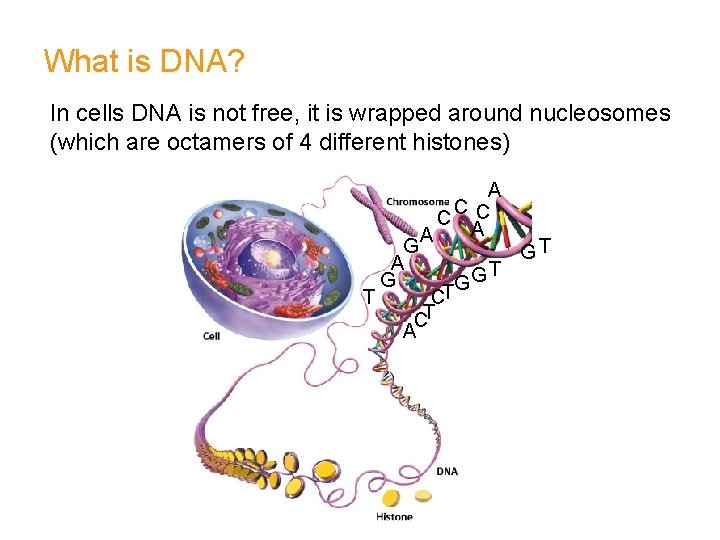
What is DNA? In cells DNA is not free, it is wrapped around nucleosomes (which are octamers of 4 different histones) T G A C C C A A G G CT CT A T GT
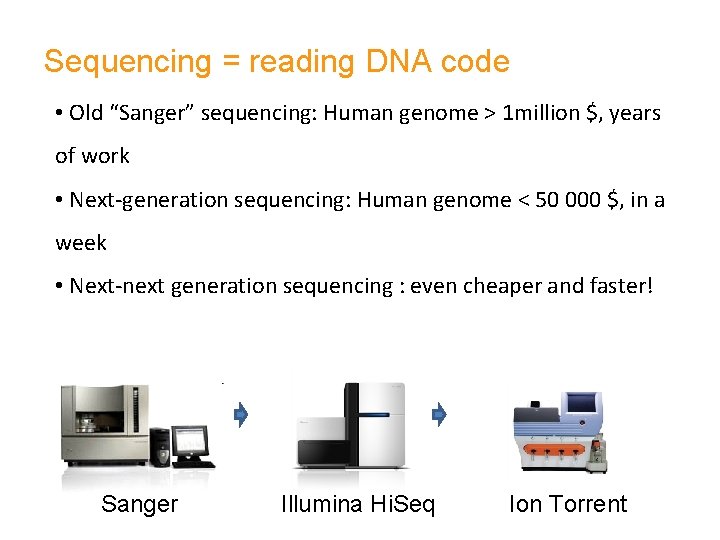
Sequencing = reading DNA code • Old “Sanger” sequencing: Human genome > 1 million $, years of work • Next-generation sequencing: Human genome < 50 000 $, in a week • Next-next generation sequencing : even cheaper and faster! Sanger Illumina Hi. Seq Ion Torrent

Why are we interested to read DNA code? Example 1: to understand the cause of a disease related to a genomic change (cancer, development abnormalities, autism, schizophrenia) Single Nucleotide Polymorphism SNP mutation 5'-ATTGCGGTATAGCGCT-3' Mutated DNA 5'-ATTGCCGTATTGCGCT-3' 5'-ATTGCGGTATTGCGCT-3' Normal DNA Mutation in an exon may lead to a dysfunctional protein

Why are we interested to read DNA code? Example 2: Sequence RNA to see which genes are active and which exons are transcribed. brain breast
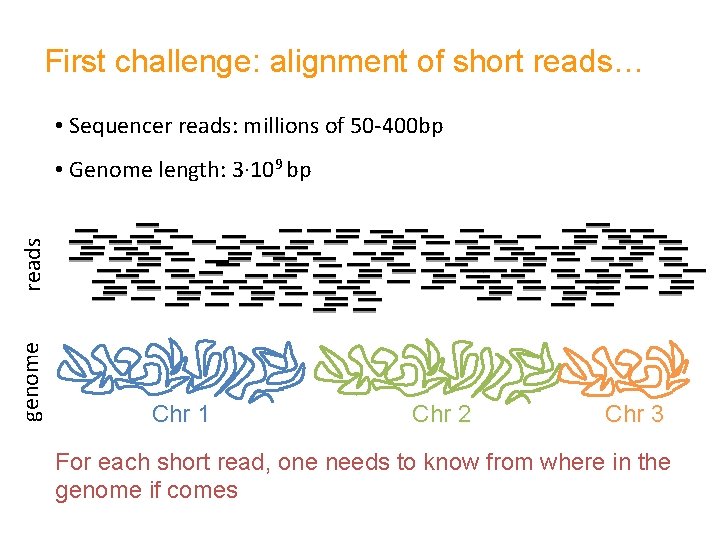
First challenge: alignment of short reads… • Sequencer reads: millions of 50 -400 bp genome reads • Genome length: 3∙ 109 bp Chr 1 Chr 2 Chr 3 For each short read, one needs to know from where in the genome if comes

Alignement should be done with mismatches • Read is short sequence that can contain errors and mutations/SNPs 5'-ATTGCGGTATAGCTCT-3' Read 5'-…ATTGCCGTATTGCGCT…-3' Genomic sequence 5'-…ATTGCGGTATTGCGCT…-3' mutation SNP Sequencing P(SNP)≈1/1000 errors Reference sequence When you align ATCG-reads you cannot tell SNPs from errors. .
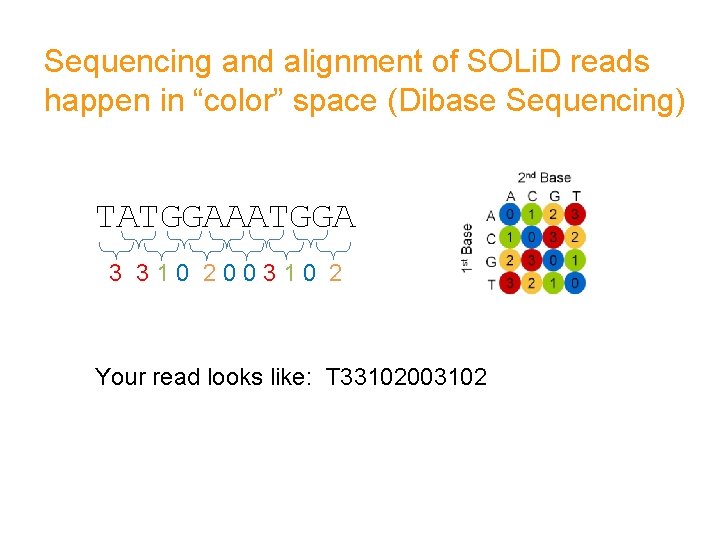
Sequencing and alignment of SOLi. D reads happen in “color” space (Dibase Sequencing) TATGGAAATGGA 3 310 200310 2 Your read looks like: T 33102003102
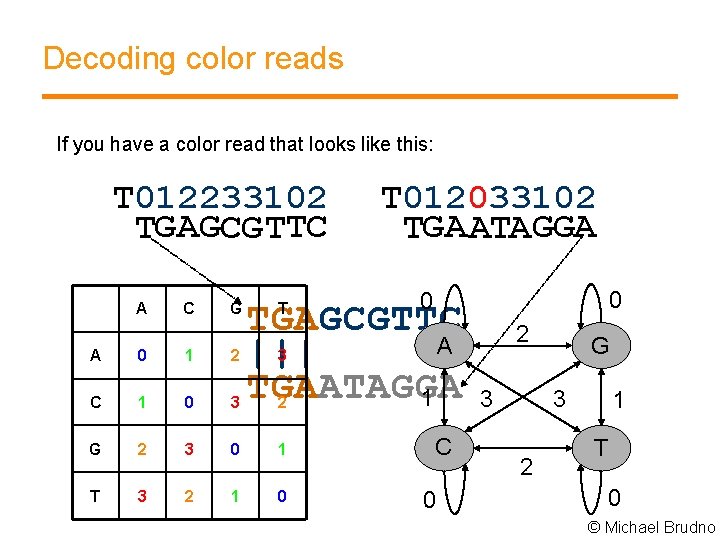
Decoding color reads If you have a color read that looks like this: T 012233102 TGAGCGTTC A 15 C G T 012033102 TGAATAGGA A 0 1 2 C 1 0 3 G 2 3 0 1 T 3 2 1 0 0 0 TGAGCGTTC A ||| TGAATAGGA 1 T 2 3 2 C 0 3 G 3 2 1 T 0 © Michael Brudno
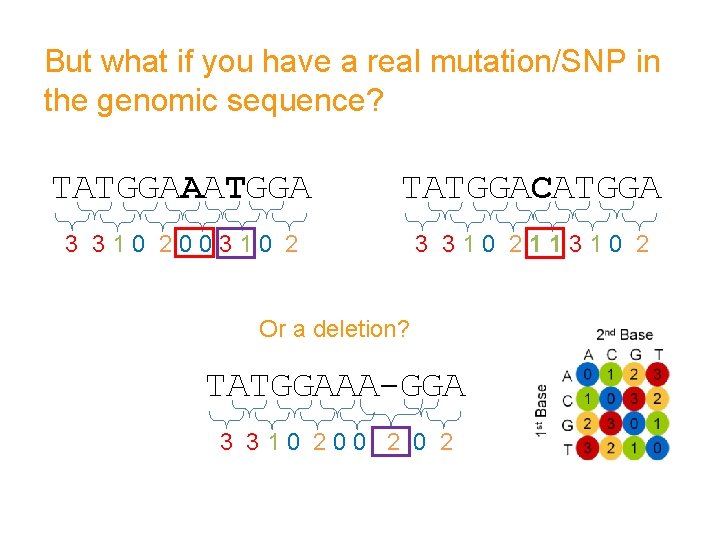
But what if you have a real mutation/SNP in the genomic sequence? TATGGAAATGGA TATGGACATGGA 3 310 200310 2 3 310 211310 2 Or a deletion? TATGGAAA-GGA 3 310 200 2

Alignment issues - reference genome is very large (Human ≈ 3 Gb) and millions of reads…. (Sensitivity vs Speed) - with mismatches, insertions and deletions … - work with colors in the case of SOLi. D reads - take into account qualities. . since for each sequenced base (ATCG) or color we know the probability that we misread it. Read: AAACTCTTCAAAAAATCAATGAATCCAGGAGCTGG Quality (q): hhhhhhhhhhhhhhhh. Mhhe Qi = ord(qi) – 64 quality at position i Pi = 10 Q/10/(1+ 10 Q/10) probability of error at position i
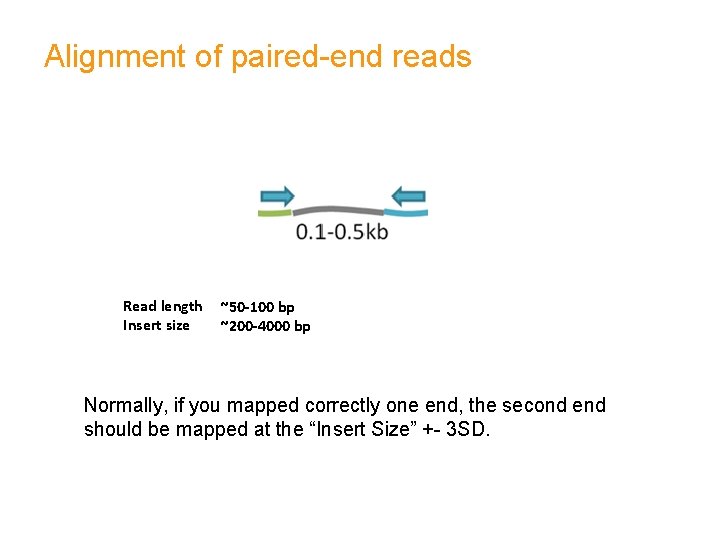
Alignment of paired-end reads Read length Insert size ~50 -100 bp ~200 -4000 bp Normally, if you mapped correctly one end, the second end should be mapped at the “Insert Size” +- 3 SD.
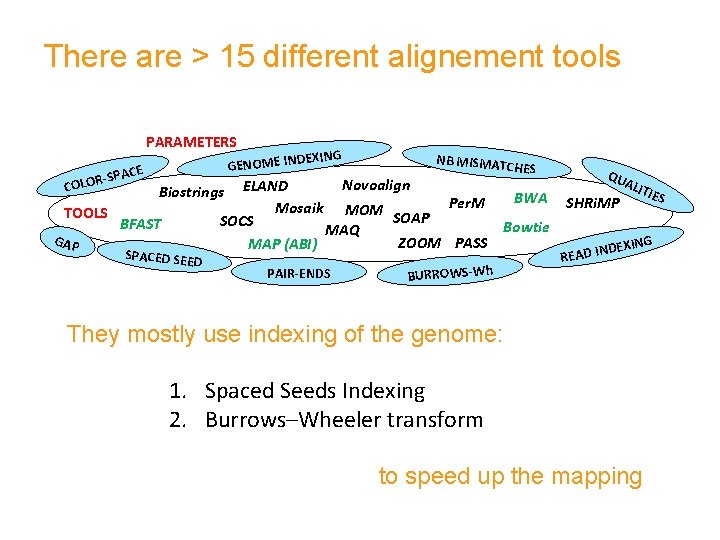
There are > 15 different alignement tools PARAMETERS DEXING GENOME IN SPACE LOR- NB MISMAT C HES Novoalign Biostrings ELAND BWA Per. M Mosaik MOM TOOLS SOAP SOCS BFAST Bowtie MAQ GAP ZOOM PASS MAP (ABI) CO SPACED SEE D PAIR-ENDS BURROWS-Wh QU SHRi. MP ALI TIE S EXING ND READ I They mostly use indexing of the genome: 1. Spaced Seeds Indexing 2. Burrows–Wheeler transform to speed up the mapping

Spaced Seeds Indexing Read ACATGCTAGTGCGTAGCTTAACAGTCGAGGCTAG Seed 1: care ! 0: don’t care ! Masks The four spaced seeds of weight 13 each span the 33 bp read, and guarantee that every location with at most two mismatches is found (sensitivity)

Burrows–Wheeler transform (BWT) …also called block-sorting compression We transform first the reference sequence (e. g. , k-mers): The output of BWT is easier to compress because it has many repeated characters. Using BWT significantly reduces memory requirement.

Burrows–Wheeler transform
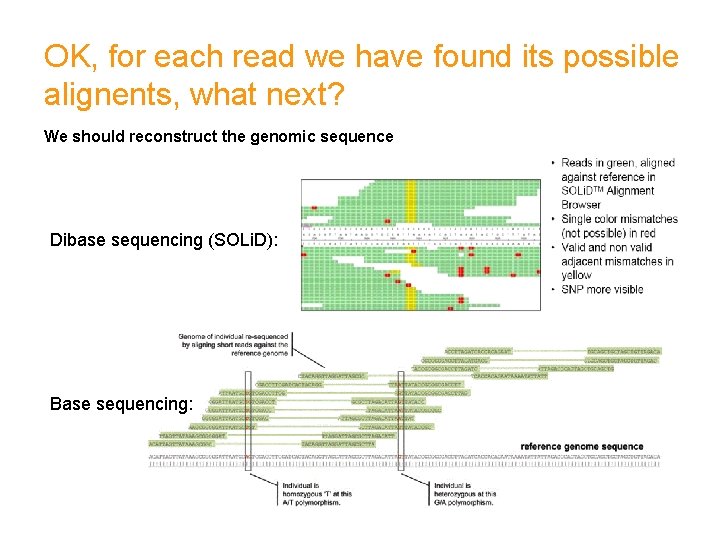
OK, for each read we have found its possible alignents, what next? We should reconstruct the genomic sequence Dibase sequencing (SOLi. D): Base sequencing:
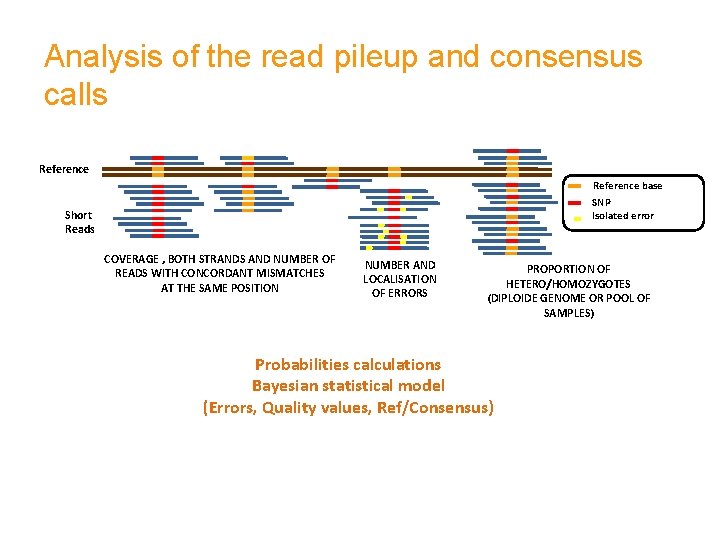
Analysis of the read pileup and consensus calls Reference base SNP Isolated error Short Reads COVERAGE , BOTH STRANDS AND NUMBER OF READS WITH CONCORDANT MISMATCHES AT THE SAME POSITION NUMBER AND LOCALISATION OF ERRORS PROPORTION OF HETERO/HOMOZYGOTES (DIPLOIDE GENOME OR POOL OF SAMPLES) Probabilities calculations Bayesian statistical model (Errors, Quality values, Ref/Consensus)
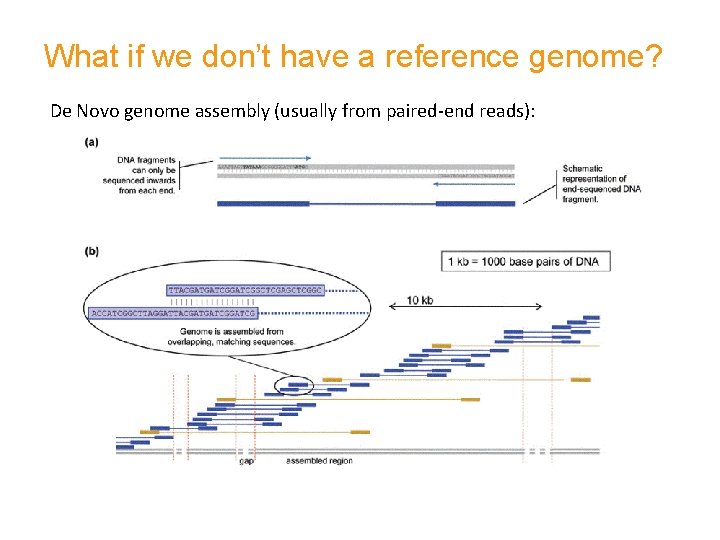
What if we don’t have a reference genome? De Novo genome assembly (usually from paired-end reads):

De Novo genome assembly most often uses • conservative seed and extension approaches • Eulerian path through the de Bruijn graph

Conservative seed and extension approaches (SSAKE, Warren et al. , 2007) hash table: (sequence, # of occurrences in the set) Reads prefix tree for first eleven 5' end bases of reads. in order to limit the search space 27

Conservative seed and extension approaches (SSAKE, Warren et al. , 2007) Start from most abundant reads: for each unassembled read u: for its possible 3' most k-mer: search for a read r with a perfect match of its 5’ with this 3' k-mer u. 5’-AAGAATACAGGATCTAGAATCTCAC 5’-AGAATACAGGATCTAGAATCTCAC 5’-AATACAGGATCTAGAATCTCAC etc 5’-AATACAGGATCTAGAATCTCACCCT If so, u is extended by the unmatched 3' end of r, and r is removed from the hash table and prefix tree. (minimum overlap, e. g. , 16) The process of cycling is repeated after every extension of u. 28
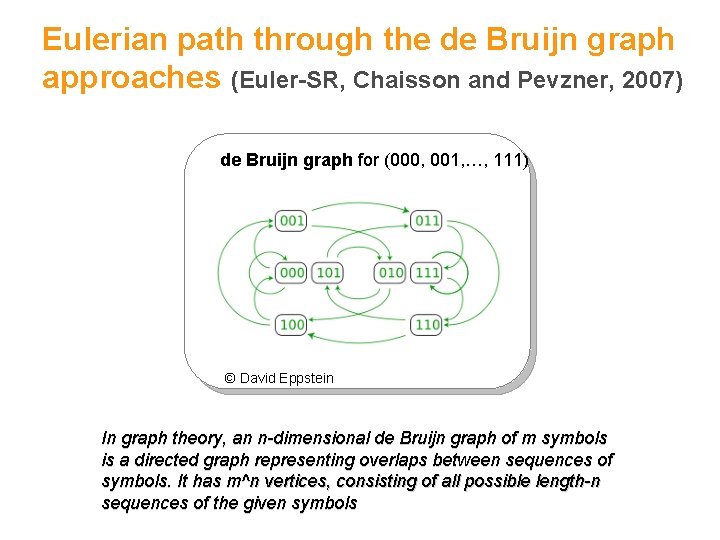
Eulerian path through the de Bruijn graph approaches (Euler-SR, Chaisson and Pevzner, 2007) de Bruijn graph for (000, 001, …, 111) © David Eppstein 29 In graph theory, an n-dimensional de Bruijn graph of m symbols is a directed graph representing overlaps between sequences of symbols. It has m^n vertices, consisting of all possible length-n sequences of the given symbols
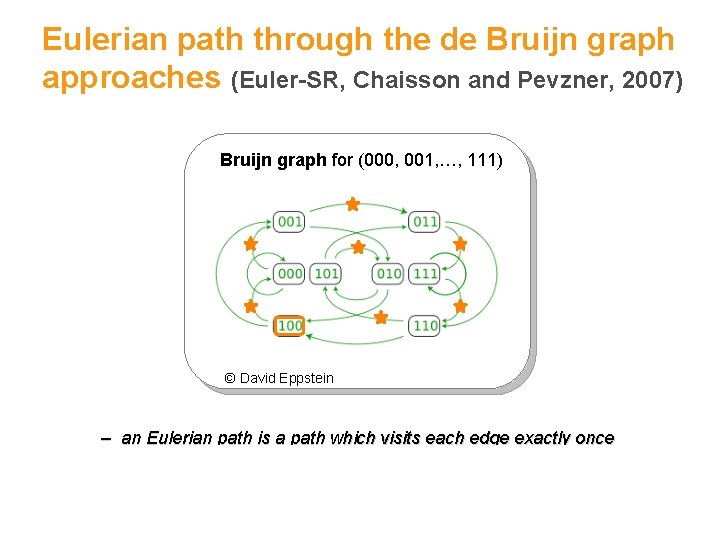
Eulerian path through the de Bruijn graph approaches (Euler-SR, Chaisson and Pevzner, 2007) Bruijn graph for (000, 001, …, 111) © David Eppstein – an Eulerian path is a path which visits each edge exactly once 30

Eulerian path through the de Bruijn graph approaches (Euler-SR, Chaisson and Pevzner, 2007) Maximum branching optimization on de Bruijn graphs error correction graph construction graph correction assembly by transforming paths in the corrected graph Spectral Alignment de Bruijn graph for reads replacing the de Bruijn graph with its Maximum branching find the Eulerian path through the de Bruijn graph that corresponds to the assembled genome define a path of edges for each read, and to assign a weight to every edge equal to the number of reads mapped to the edge. Maximum Branching - contains no cycles and has all vertices with indegree at most 1, with largest total weight 31

Alignemnt and assembling reads are not the only challenges in NGS • Prediction of genomic structural variances using pairedend data • Calculation of copy number profiles (gain and losses of peaces of chromosomes) • …. .

Prediction of genomic structural variances using paired-end data

Prediction of genomic structural variances using paired-end data Select “abnormal” reads Cluster “abnormal” reads For each cluster infer its type of structural variance What if there are some ‘noise’ abnormal reads?

Copy-number profiles Assessing copy numbers
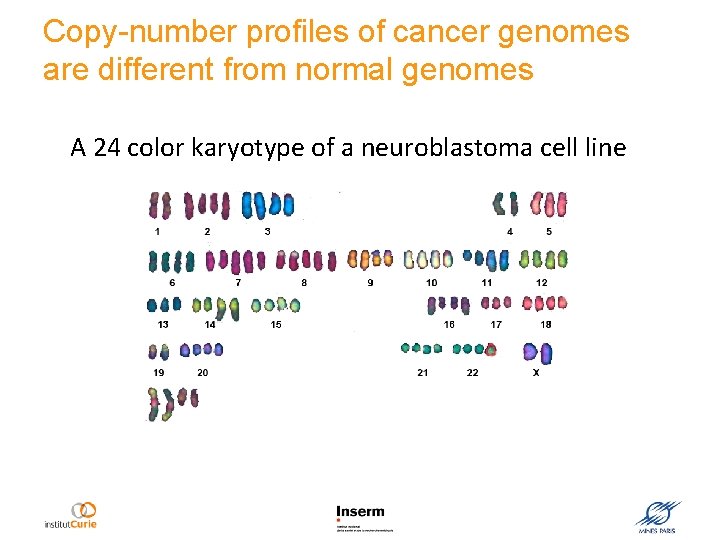
Copy-number profiles of cancer genomes are different from normal genomes A 24 color karyotype of a neuroblastoma cell line

Identification of regions of gain and loss helps to predict the aggressiveness of cancer
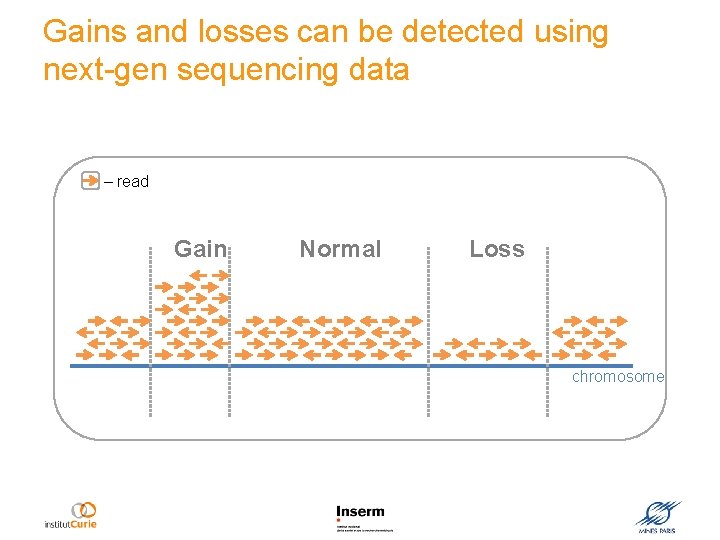
Gains and losses can be detected using next-gen sequencing data – read Gain Normal Loss chromosome
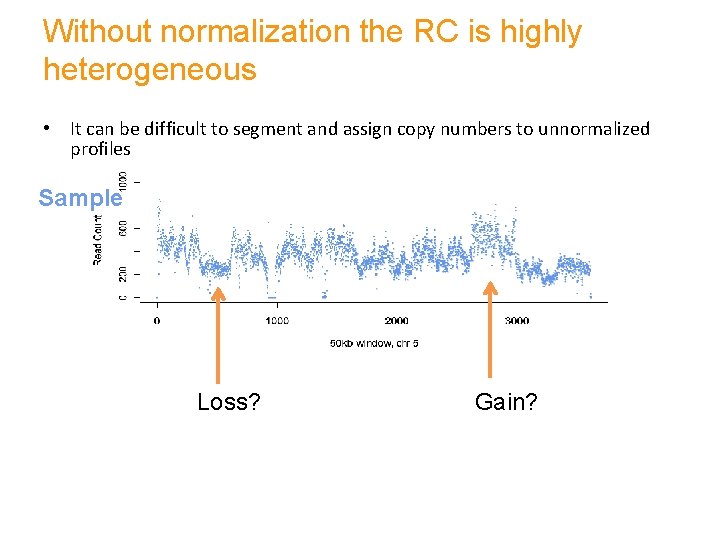
Without normalization the RC is highly heterogeneous • It can be difficult to segment and assign copy numbers to unnormalized profiles Sample Control Loss? Gain?
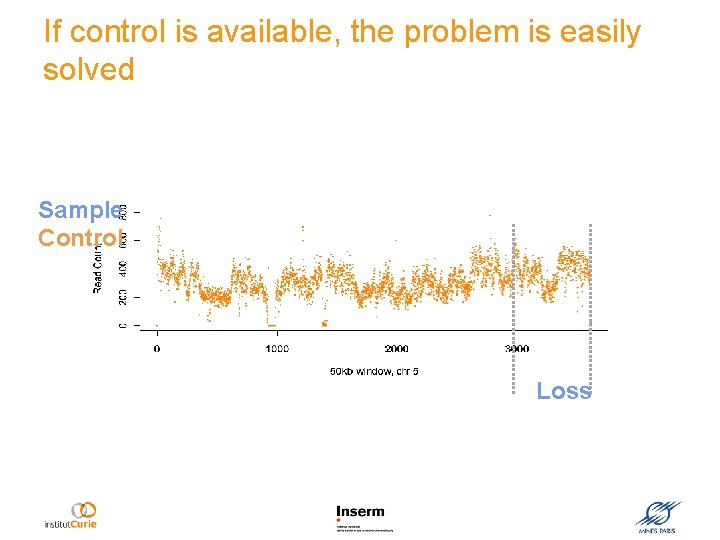
If control is available, the problem is easily solved Sample Control Loss

If control is available, the problem is easily solved Loss
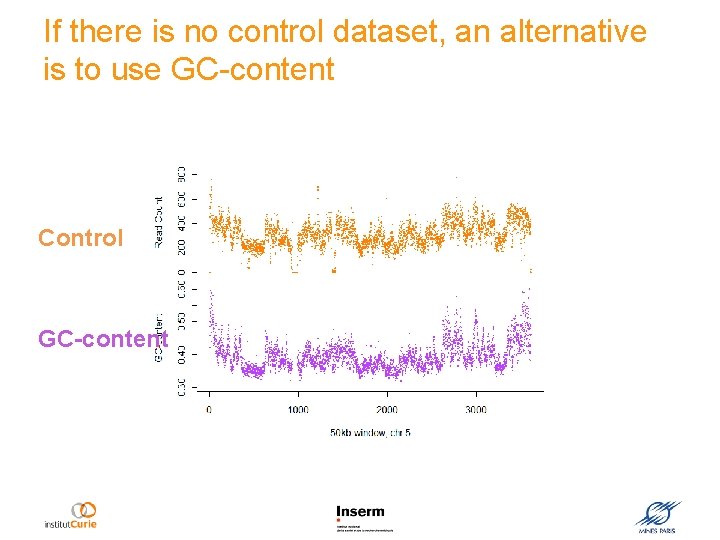
If there is no control dataset, an alternative is to use GC-content Control GC-content
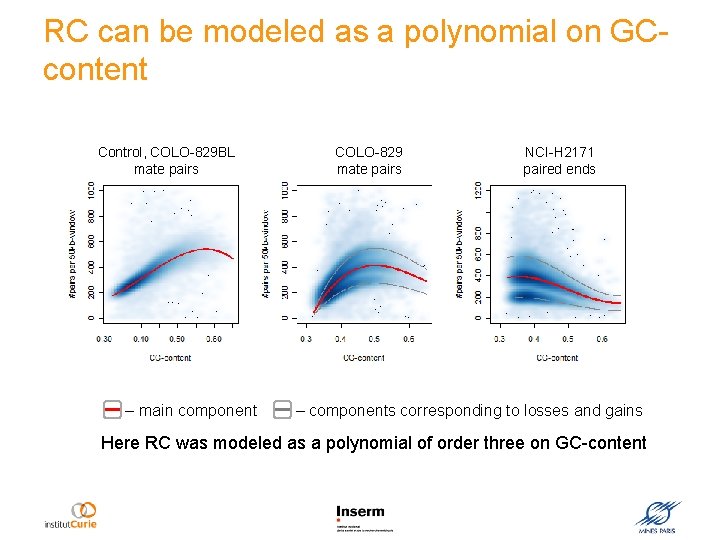
RC can be modeled as a polynomial on GCcontent Control, COLO-829 BL mate pairs – main component COLO-829 mate pairs NCI-H 2171 paired ends – components corresponding to losses and gains Here RC was modeled as a polynomial of order three on GC-content
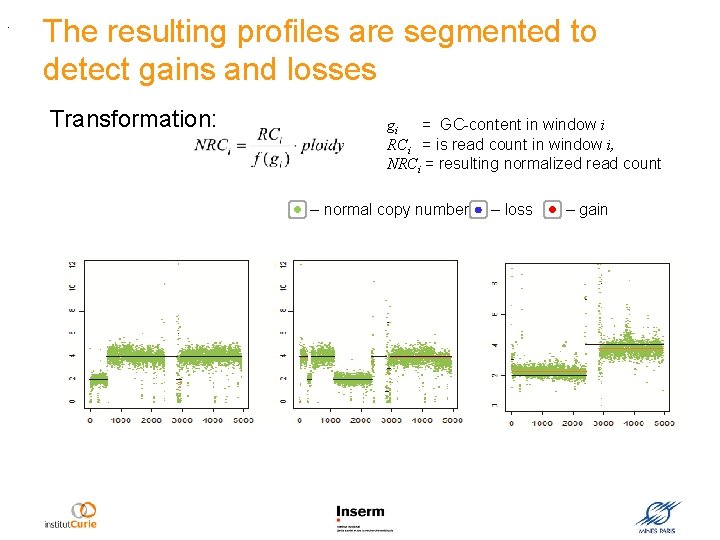
The resulting profiles are segmented to detect gains and losses Transformation: gi = GC-content in window i RCi = is read count in window i, NRCi = resulting normalized read count – normal copy number Normalized copy number . Genomic position (50 -kb window) – loss – gain
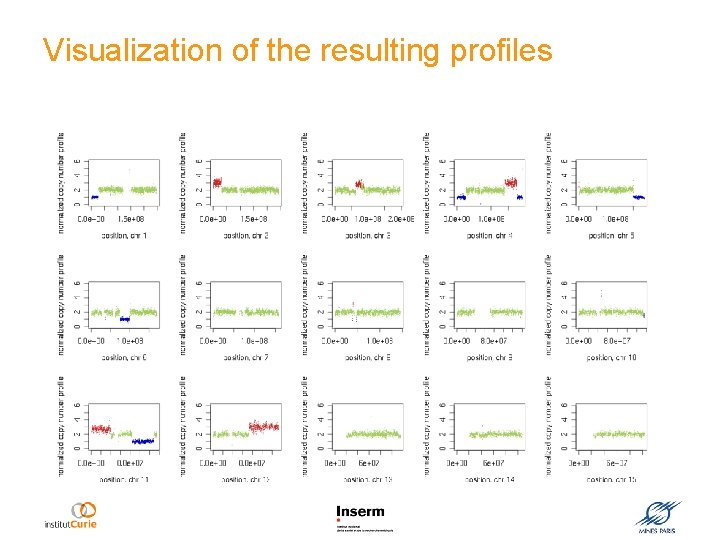
Visualization of the resulting profiles

Conclusion • Next generation sequencing produces large amount of data that should be analyzed by bioinformaticians • elaboration of efficient tools is needed to treat the data • Alignment • De novo assembly • Detection of genomic structural variants • Calculation of copy number profiles (normalization and segmentation) • Etc. .
![S] High number of other applications Already experimented at the NGS platform of the S] High number of other applications Already experimented at the NGS platform of the](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-47.jpg)
S] High number of other applications Already experimented at the NGS platform of the IC
![R] Visualisation of SV deletion interchromosomal and large Intrachromosomal SVs in Circos (SVDetect tool) R] Visualisation of SV deletion interchromosomal and large Intrachromosomal SVs in Circos (SVDetect tool)](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-48.jpg)
R] Visualisation of SV deletion interchromosomal and large Intrachromosomal SVs in Circos (SVDetect tool) interchromosomal translocation & inverted duplication
![T] Ch. IP-Seq method 50 bp Ch. IP-Seq is a technique to identify protein T] Ch. IP-Seq method 50 bp Ch. IP-Seq is a technique to identify protein](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-49.jpg)
T] Ch. IP-Seq method 50 bp Ch. IP-Seq is a technique to identify protein binding sites on DNA Mains steps of Ch. IP-Seq • Example: EWS-FLI 1 oncogenic transcription factor – cause of Ewing sarcoma (O. Delattre, U 830) EWS-activation domain FLI 1 DNA binding domain Age
![T] Types of Ch. IP-Seq analysis H 3 K 36 me 3 Activated Repressed T] Types of Ch. IP-Seq analysis H 3 K 36 me 3 Activated Repressed](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-50.jpg)
T] Types of Ch. IP-Seq analysis H 3 K 36 me 3 Activated Repressed Noresponse H 3 K 9 me 3 RNApol. II SPI-1 Spi-1 binding in preleukemic mouse cells (Christel Guillouf, U 830) Spi-1 is rather an activator that binds around TSSs Total promoters Spi-1 binding in promoters Cp. G islands help Spi-1 binding
![U] Small. RNA as regulators of gene expression How to characterize the small. RNAs U] Small. RNA as regulators of gene expression How to characterize the small. RNAs](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-51.jpg)
U] Small. RNA as regulators of gene expression How to characterize the small. RNAs expression profile ?
![V] Small. RNA-seq analysis • mi. RNA annotation and comparison • Detection of si. V] Small. RNA-seq analysis • mi. RNA annotation and comparison • Detection of si.](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-52.jpg)
V] Small. RNA-seq analysis • mi. RNA annotation and comparison • Detection of si. RNAs cluster • Analyses of small. RNAs in repeated regions • Pipelines for small. RNAs analysis • Sequencing platform comparison • . . . Small. RNA profiling in Mouse ES cells during the differentiation Project in collaboration with E. Heard's lab (C. Ciaudo, J. Toedling, JC. Chen) and O. Voinnet (IBMP, ETH)
![W] 5 C : Measure physical interactions From E. Nora W] 5 C : Measure physical interactions From E. Nora](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-53.jpg)
W] 5 C : Measure physical interactions From E. Nora
![X] 5 C : A new technology Forward Primers • How to manipulate the X] 5 C : A new technology Forward Primers • How to manipulate the](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-54.jpg)
X] 5 C : A new technology Forward Primers • How to manipulate the data ? Forward Primers Chromosomal interaction • How to represent the interaction map ? • How to remove noise and background ? • How to compare samples ? • How to detect interaction domains ? • . . . Detection of chromosomal interaction in the context of X inactivation Project in collaboration with E. Heard's lab (E. Nora, L. Giorgetti, ) and the J. Dekker's lab (B. Lajoie, UMMS, USA)
![Y] The NGS project ending • Meeting for the final presentation of the results Y] The NGS project ending • Meeting for the final presentation of the results](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-55.jpg)
Y] The NGS project ending • Meeting for the final presentation of the results • Report writing • Publication (Material & Methods, Figures) • Presentation of the NGS project at external meetings (NGS meetings, conferences, congress, posters, …) • Definition of new objectives • Next NGS projects Pilot experiment, success > bigger NGS project ? Addition of new samples (ex. Need of more controls)
![Z] Past NGS projects at the IC Biomedical research (30%): • human cancers : Z] Past NGS projects at the IC Biomedical research (30%): • human cancers :](http://slidetodoc.com/presentation_image/3f31d9057b2877ebfcd1f2d58068a785/image-56.jpg)
Z] Past NGS projects at the IC Biomedical research (30%): • human cancers : breast, ovarian, cervix, ewing • human normal cells : infertility Pilot projects for diagnosis (10%): • breast cancers and BRCA genes, • 43 genes implicated in carcinogenesis Fundamental researchs (60%): • yeast (mutations, meiosis and non-coding RNA), • mouse (small RNAs) • human (histone H 3, Common Fragile Sites)

The End NGS 2011 : A color space odyssey
- Slides: 57