Bioconductor Packages for Preprocessing DNA Microarray Data affy

Bioconductor Packages for Pre-processing DNA Microarray Data affy and marray Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course Winter 2002 © Copyright 2002, all rights reserved
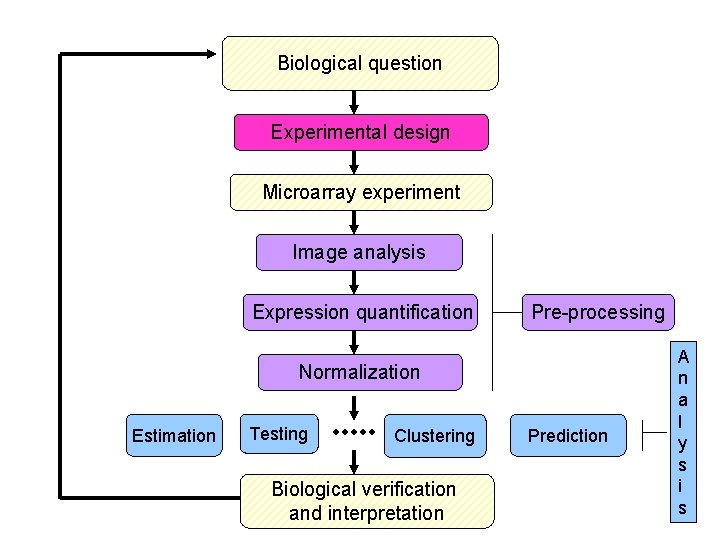
Biological question Experimental design Microarray experiment Image analysis Expression quantification Pre-processing Normalization Estimation Testing Clustering Biological verification and interpretation Prediction A n a l y s i s

Pre-processing • affy: Affymetrix oligonucleotide chips • marray: Spotted DNA microarrays Reading in intensity data, diagnostic plots, normalization, expression measures. Both suites of packages start with very different data types, but produce similar objects of class expr. Set. One can then use other Bioconductor packages, e. g. , genefilter, geneplotter.
Pre-processing: spotted DNA microarrays

marray: Pre-processing spotted DNA microarray data • marray. Classes: – class definitions for c. DNA microarray data (MIAME); – basic methods for manipulating microarray objects: printing, plotting, subsetting, class conversions, etc. • marray. Input: – reading in intensity data and textual data describing probes and targets; – automatic generation of microarray data objects; – widgets for point & click interface. • marray. Plots: diagnostic plots. • marray. Norm: robust adaptive location and scale normalization procedures.
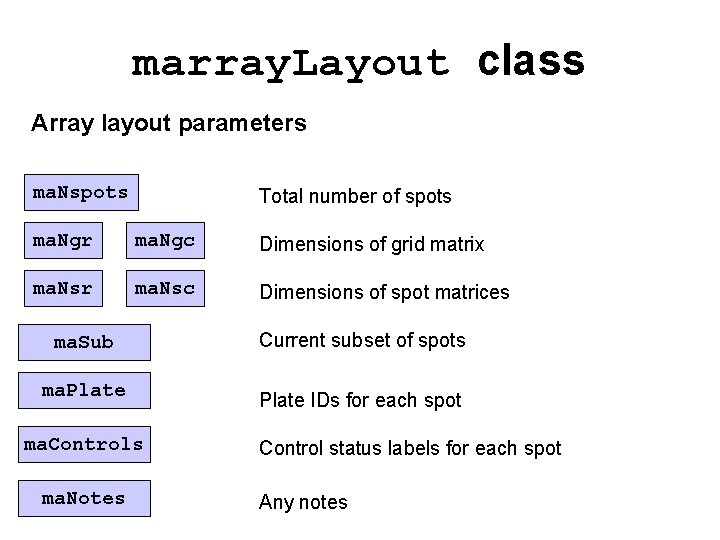
marray. Layout class Array layout parameters ma. Nspots Total number of spots ma. Ngr ma. Ngc Dimensions of grid matrix ma. Nsr ma. Nsc Dimensions of spot matrices ma. Sub Current subset of spots ma. Plate IDs for each spot ma. Controls ma. Notes Control status labels for each spot Any notes
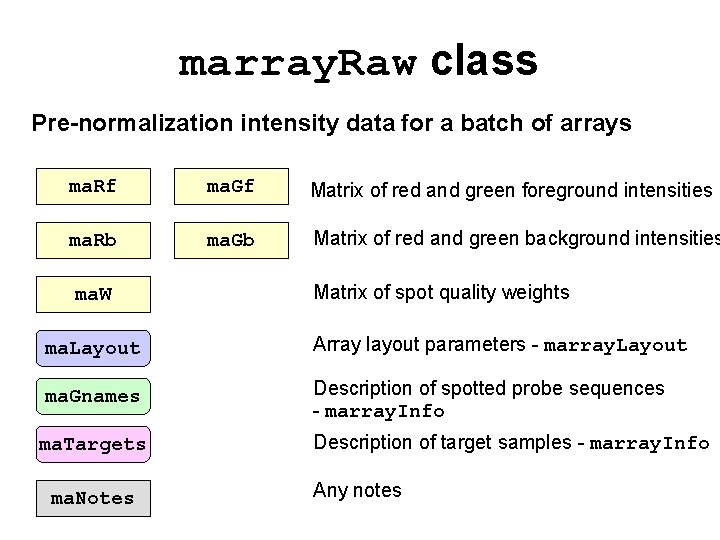
marray. Raw class Pre-normalization intensity data for a batch of arrays ma. Rf ma. Gf Matrix of red and green foreground intensities ma. Rb ma. Gb Matrix of red and green background intensities ma. W Matrix of spot quality weights ma. Layout Array layout parameters - marray. Layout ma. Gnames Description of spotted probe sequences - marray. Info ma. Targets ma. Notes Description of target samples - marray. Info Any notes
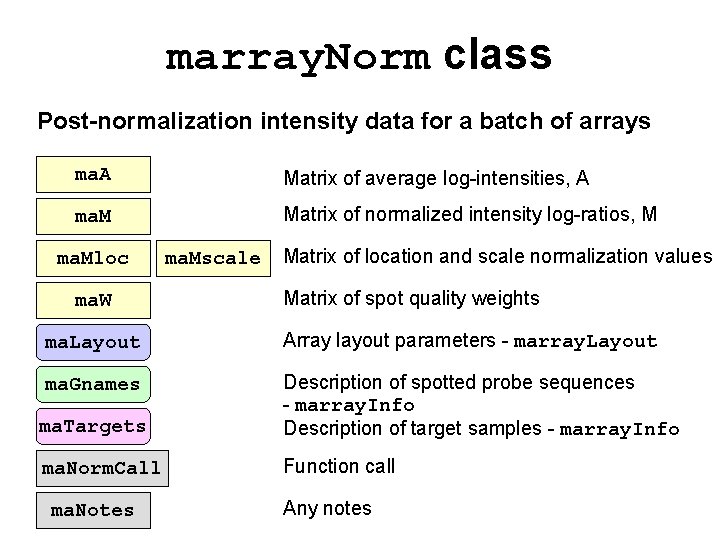
marray. Norm class Post-normalization intensity data for a batch of arrays ma. A Matrix of average log-intensities, A ma. M Matrix of normalized intensity log-ratios, M ma. Mloc ma. W ma. Mscale Matrix of location and scale normalization values Matrix of spot quality weights ma. Layout Array layout parameters - marray. Layout ma. Gnames ma. Targets Description of spotted probe sequences - marray. Info Description of target samples - marray. Info ma. Norm. Call Function call ma. Notes Any notes

marray. Input package • marray. Input provides functions for reading microarray data into R and creating microarray objects of class marray. Layout, marray. Info, and marray. Raw. • Input – Image quantitation data, i. e. , output files from image analysis software. E. g. . gpr for Gene. Pix, . spot for Spot. – Textual description of probe sequences and target samples. E. g. gal files, god lists.
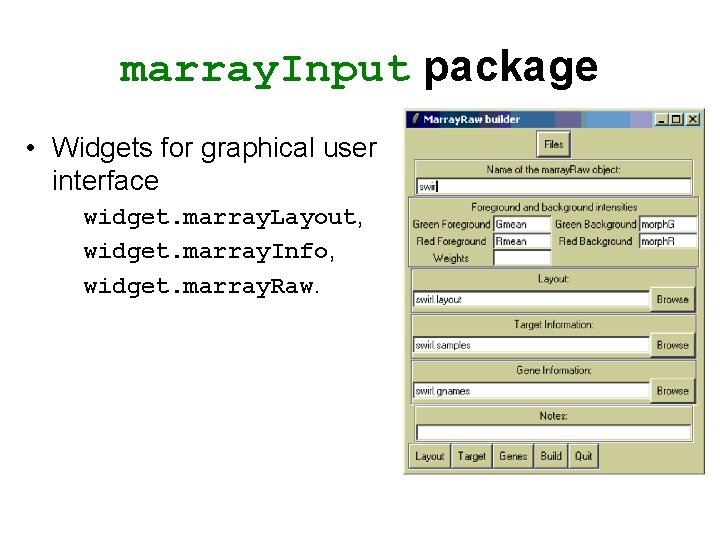
marray. Input package • Widgets for graphical user interface widget. marray. Layout, widget. marray. Info, widget. marray. Raw.

marray. Plots package • See demo(marray. Plots). • Diagnostic plots of spot statistics. E. g. red and green log intensities, intensity log ratios M, average log intensities A, spot area. – ma. Image: 2 D spatial color images. – ma. Boxplot: boxplots. – ma. Plot: scatter-plots with fitted curves and text highlighted. • Stratify plots according to layout parameters such as print-tip-group, plate. E. g. MA-plots with loess fits by print-tip-group.

2 D spatial images ma. Image Cy 3 background intensity Cy 5 background intensity
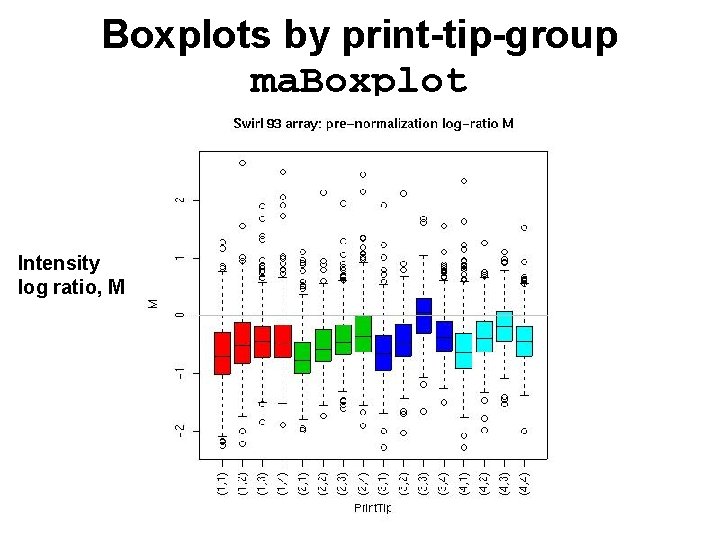
Boxplots by print-tip-group ma. Boxplot Intensity log ratio, M
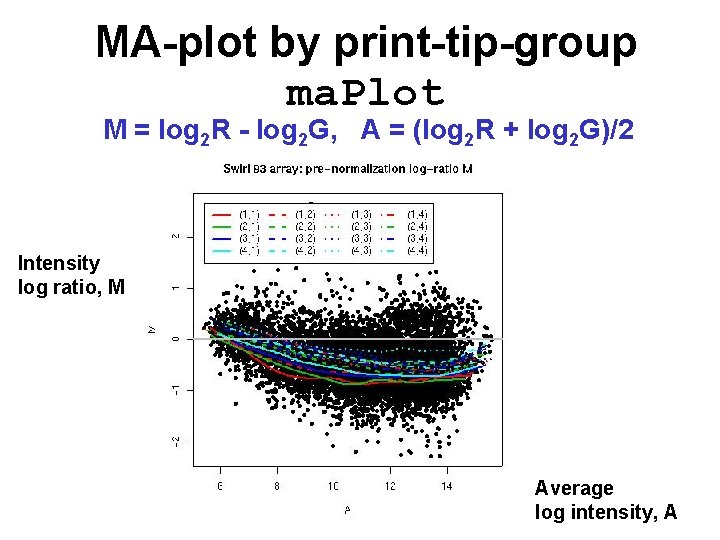
MA-plot by print-tip-group ma. Plot M = log 2 R - log 2 G, A = (log 2 R + log 2 G)/2 Intensity log ratio, M Average log intensity, A

marray. Norm package • ma. Norm. Main: main normalization function, allows robust adaptive location and scale normalization for a batch of arrays – intensity or A-dependent location normalization (ma. Norm. Loess); – 2 D spatial location normalization (ma. Norm 2 D); – median location normalization (ma. Norm. Med); – scale normalization using MAD (ma. Norm. MAD); – composite normalization; – your own normalization function. • ma. Norm: simple wrapper function. ma. Norm. Scale: simple wrapper function for scale normalization.
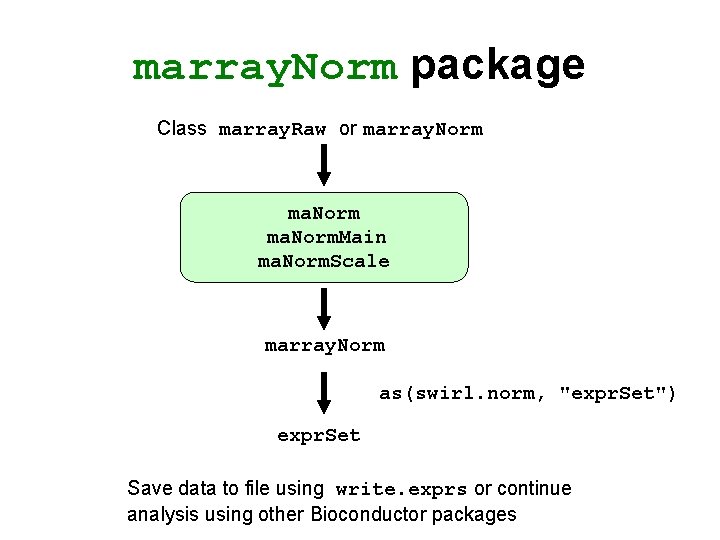
marray. Norm package Class marray. Raw or marray. Norm ma. Norm. Main ma. Norm. Scale marray. Norm as(swirl. norm, "expr. Set") expr. Set Save data to file using write. exprs or continue analysis using other Bioconductor packages

swirl dataset • Microrrays: – 8, 448 probes (768 controls); – 4 x 4 grid matrix; – 22 x 24 spot matrices. • 4 hybridizations: swirl mutant and wild type m. RNA • Data stored in object of class marray. Raw: data(swirl). • > ma. Info(ma. Targets(swirl))[, 3: 4] experiment Cy 3 experiment Cy 5 1 swirl wild type 2 wild type swirl 3 swirl wild type 4 wild type swirl

Oligonucleotide chips
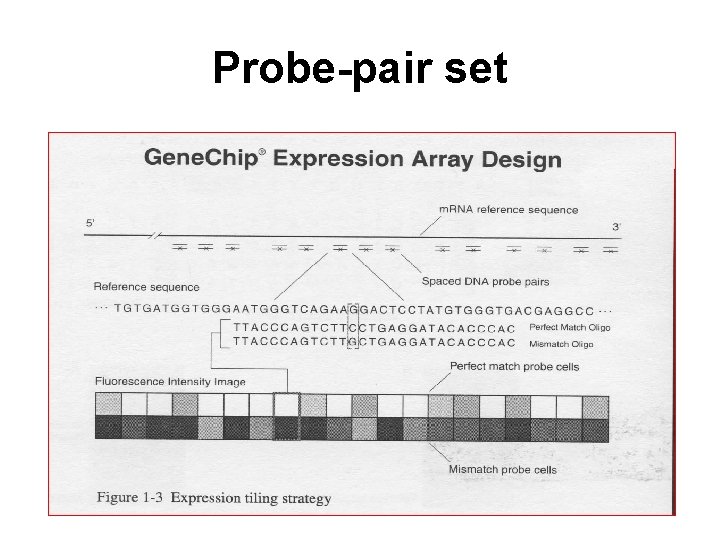
Probe-pair set

Terminology • Each gene or portion of a gene is represented by 16 to 20 oligonucleotides of 25 base-pairs. • Probe: an oligonucleotide of 25 base-pairs, i. e. , a 25 -mer. • Perfect match (PM): A 25 -mer complementary to a reference sequence of interest (e. g. , part of a gene). • Mismatch (MM): same as PM but with a single homomeric base change for the middle (13 th) base (transversion purine <-> pyrimidine, G <->C, A <->T). • Probe-pair: a (PM, MM) pair. • Probe-pair set: a collection of probe-pairs (16 to 20) related to a common gene or fraction of a gene. • Affy ID: an identifier for a probe-pair set. • The purpose of the MM probe design is to measure non-specific binding and background noise.

Affymetrix files • Main software from Affymetrix company Micro. Array Suite - MAS, now version 5. • DAT file: Image file, ~10^7 pixels, ~50 MB. • CEL file: Cell intensity file, probe level PM and MM values. • CDF file: Chip Description File. Describes which probes go in which probe sets and the location of probe-pair sets (genes, gene fragments, ESTs).

affy: Pre-processing Affymetrix data • Class definitions for probe-level data: Affy. Batch, Prob. Set, Cdf, Cel. • Basic methods for manipulating microarray objects: printing, plotting, subsetting. • Functions and widgets for data input from CEL and CDF files, and automatic generation of microarray data objects. • Diagnostic plots: 2 D spatial images, density plots, boxplots, MA-plots, etc.

affy: Pre-processing Affymetrix data • Background estimation. • Probe-level normalization: quantile and curvefitting normalization (Bolstad et al. , 2002). • Expression measures: MAS 4. 0 Av. Diff, MAS 5. 0 Signal, MBEI (Li & Wong, 2001), RMA (Irizarry et al. , 2003). • Main functions: Read. Affy, rma, expresso, express.
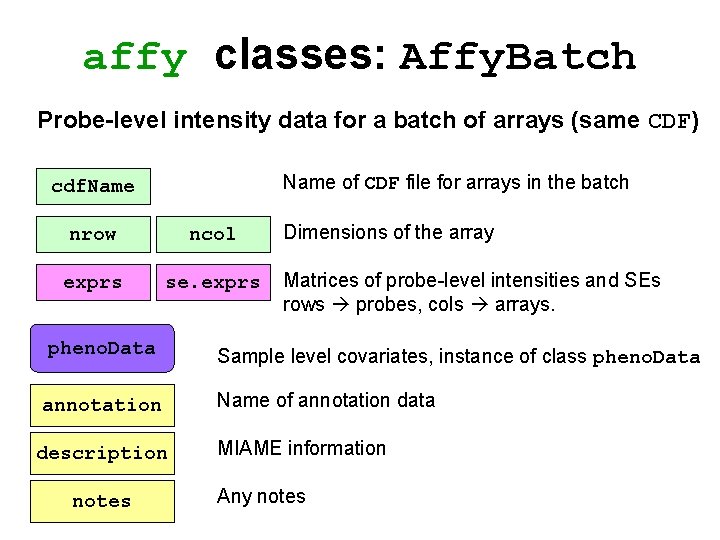
affy classes: Affy. Batch Probe-level intensity data for a batch of arrays (same CDF) Name of CDF file for arrays in the batch cdf. Name nrow ncol exprs se. exprs pheno. Data annotation description notes Dimensions of the array Matrices of probe-level intensities and SEs rows probes, cols arrays. Sample level covariates, instance of class pheno. Data Name of annotation data MIAME information Any notes

affy classes • Probe. Set: PM, MM intensities for individual probe sets. – pm: matrix of PM intensities for individual probe sets, rows probes, cols arrays. – mm: matrix of MM intensities for individual probe sets, rows probes, cols arrays. Apply probeset to Affy. Batch object to get list of Probe. Set objects. • Cel: Single array cel intensity data. • Cdf: Information contained in a CDF file.

CDF data packages • Data packages containing necessary CDF information are available at www. bioconductor. org. • Packages contain environment objects, which provide mappings between Affy. IDs and matrices of probe locations, rows probe-pairs, cols PM, MM (e. g. , 20 X 2 matrix for hu 6800). • cdf. Name slot of Affy. Batch. • HGU 95 Av 2 and HGU 133 A provided in package.

Reading in data: Read. Affy Creates object of class Affy. Batch

Accessing PM and MM data • probe. Names: method for accessing Affy. IDs corresponding to individual probes. • pm, mm: methods for accessing probe-level PM and MM intensities probes x arrays matrix. • Can use on Affy. Batch objects.

Diagnostic plots • See demo(affy). • Diagnostic plots of probe-level intensities, PM and MM. – image: 2 D spatial color images of log intensities (Affy. Batch, Cel). – boxplot: boxplots of log intensities (Affy. Batch). – mva. pairs: scatter-plots with fitted curves (apply exprs, pm, or mm to Affy. Batch object). – hist: density plots of log intensities (Affy. Batch).

image
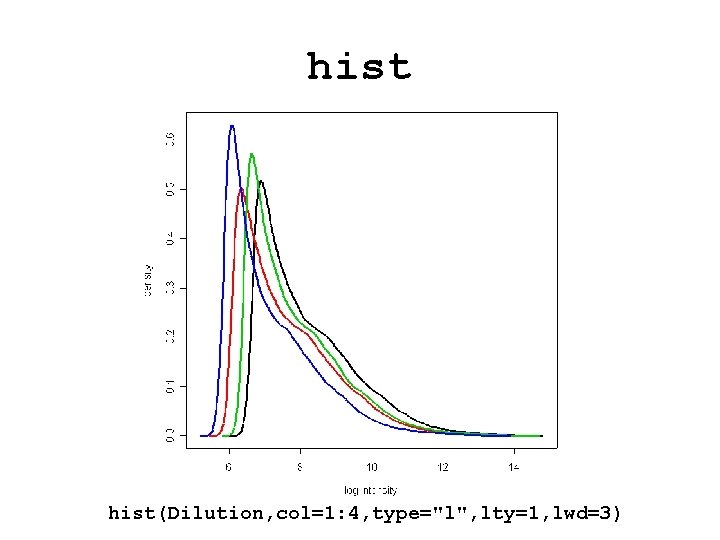
hist(Dilution, col=1: 4, type="l", lty=1, lwd=3)
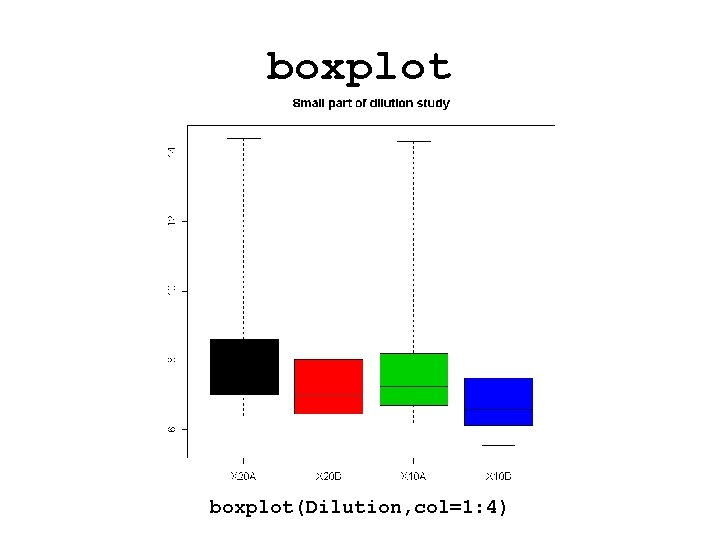
boxplot(Dilution, col=1: 4)
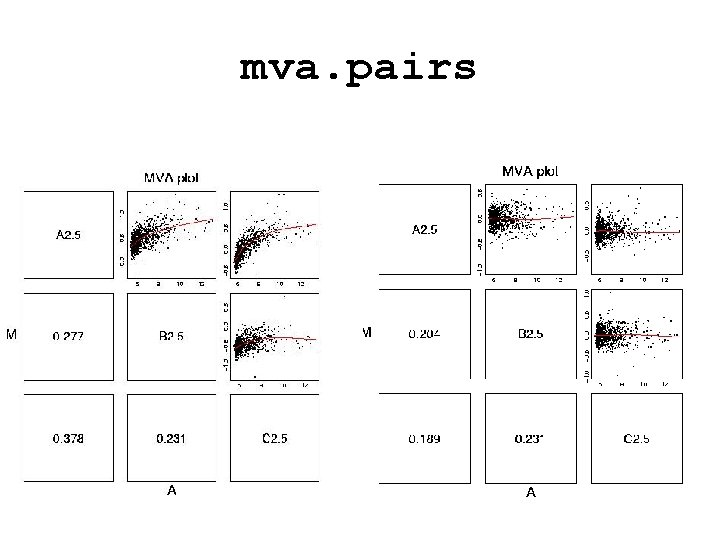
mva. pairs

Expression measures • expresso: Choice of common methods for – – background correction: bgcorrect. methods normalization: normalize. Affy. Batch. methods probe specific corrections: pmcorrect. methods expression measures: express. summary. stat. methods. • rma: Fast implementation of RMA (Irizarry et al. , 2003): model-based background correction, quantile normalization, median polish expression measures. • express: Implementing your own expression measures. • normalize: Normalization procedures in normalize. Affy. Batch. methods or normalize. methods(object).

Expression meassures: expresso(widget=TRUE)
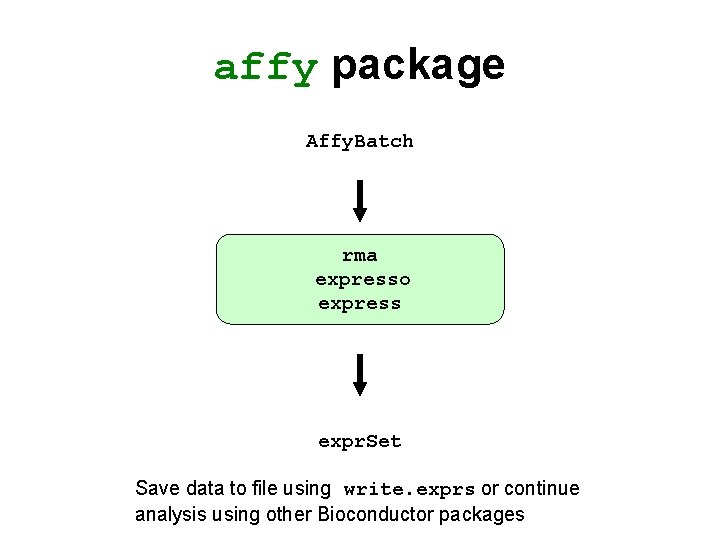
affy package Affy. Batch rma expresso express expr. Set Save data to file using write. exprs or continue analysis using other Bioconductor packages

Probe sequence analysis • Examine probe intensity based on location relative to 5’ end of RNA sequence of interest. • Expect probe intensities to be lower at 5’ end compared to 3’ of m. RNA. • E. g. deg<-Affy. RNAdeg(Dilution) plot. Affy. RNAdeg(deg)

Dilution dataset • HGU 95 A chip • 4 arrays: Human liver m. RNA – 2 concentrations: 10 and 20 mg; – 2 scanners: 1 and 2. • Data stored in object of class Affy. Batch: data(Dilution). • > p. Data(Dilution) liver sn 19 scanner 20 A 20 0 1 20 B 20 0 2 10 A 10 0 1 10 B 10 0 2

Combining data across slides Data on G genes for n hybridizations G x n genes-by-arrays data matrix Arrays Array 1 Array 2 Genes Gene 1 Gene 2 Gene 3 Gene 4 Gene 5 … 0. 46 -0. 10 0. 15 -0. 45 -0. 06 … 0. 30 0. 49 0. 74 -1. 03 1. 06 … Array 3 Array 4 0. 80 0. 24 0. 04 -0. 79 1. 35 … 1. 51 0. 06 0. 10 -0. 56 1. 09 … Array 5 … 0. 90 0. 46 0. 20 -0. 32 -1. 09 … M = log 2( Red intensity / Green intensity) expression measure, e. g, RMA . . . .
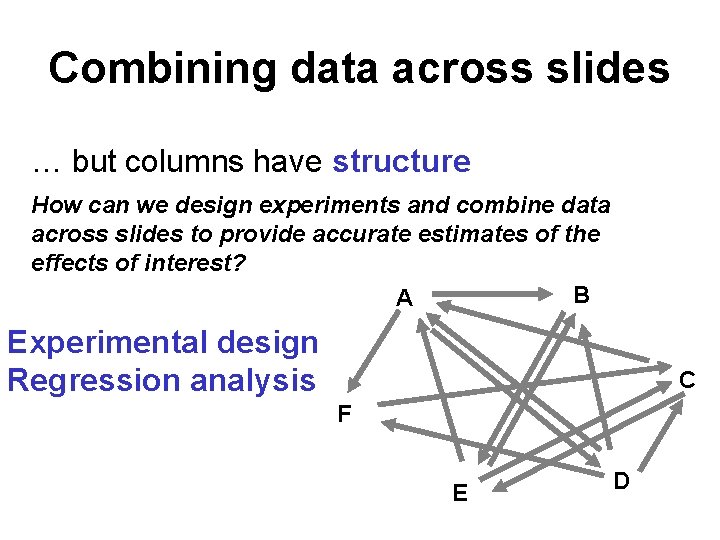
Combining data across slides … but columns have structure How can we design experiments and combine data across slides to provide accurate estimates of the effects of interest? B A Experimental design Regression analysis C F E D
expr. Set class exprs se. exprs pheno. Data annotation description notes Matrix of expression measures, genes x samples Matrix of SEs for expression measures Sample level covariates, instance of class pheno. Data Name of annotation data MIAME information Any notes

Reading in pheno. Data tk. Sample. Names tkpheno. Data tk. MIAME
- Slides: 42