Other genomic arrays Methylation ch IP on chip















- Slides: 21

Other genomic arrays: Methylation, ch. IP on chip… UBio Training Courses

SNP-arrays and copy number Genotyping arrays can detect CNVs

Copy numbers from SNP arrays

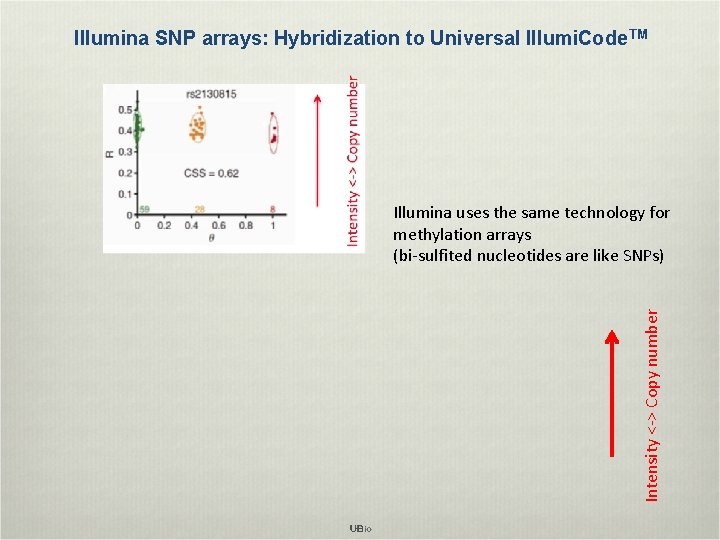
Illumina SNP arrays: Hybridization to Universal Illumi. Code TM Intensity <-> Copy number Illumina uses the same technology for methylation arrays (bi-sulfited nucleotides are like SNPs)

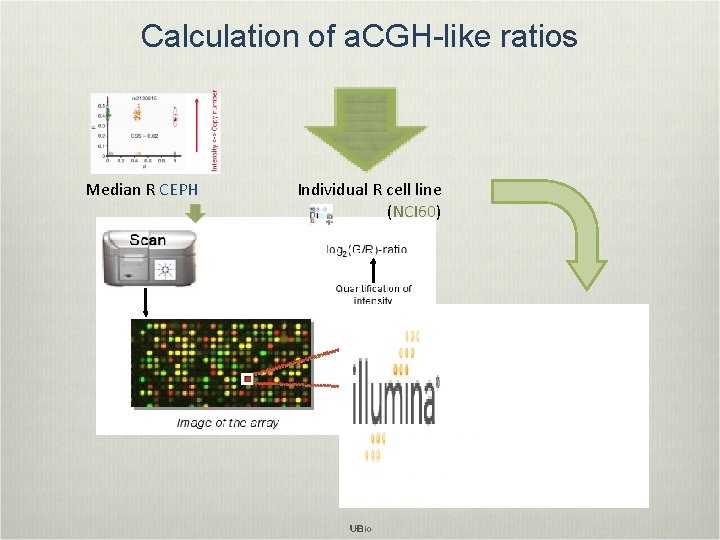
Calculation of a. CGH-like ratios Median R CEPH Individual R cell line (NCI 60)

Methylation arrays

METHYLATION MICROARRAYS Bead. Arrays Infinium Human. Methylation 27 Bead. Chip o Until 12 samples per chip. o 27, 578 Cp. G loci, >14. 000 genes o 2 beads per locus (methylated/no methylated) o Random distribution (50 mer) o Input: Bisulphyted DNA o Includes probes for the promoter regions of mi. RNA 110 genes

METHYLATION MICROARRAYS Illumina Golden Gate Assay • Until 147, 456 DNA methylation measures simultaneously. • Resolution: 1 Cp. G • Until 96 samples simultaneously • Golden. Gate Methylation Cancer Panel I 1, 505 Cp. G loci selected from 807 gene • Allows custom designs

METHYLATION MICROARRAYS SOFTWARE Bead Studio Genome Studio Methylation module http: //www. illumina. com/pages. ilmn? ID=196 Lumi package (Import, background correction, normalization) Beadarray package (Import, QC) Methylumi (Import, QC , normalization, differential meth. )

METHYLATION MICROARRAYS DIFFERENTIAL METHYLATION Bead Studio Genome Studio Methylation module http: //www. illumina. com/pages. ilmn? ID=196 Beta values: Hypermethylated Hypomethylated β= Imethylated/Imethylated+Ino_methylated 1 0. 7 β 0. 3 0

METHYLATION MICROARRAYS NORMALIZATION Methylumi normalization 1) Calculate medians for Cy 3 and Cy 5 at high an low betas 2) Cy 5 medians adjusted to Cy 3 channel (dye bias) 3) Recalculate betas with new intensities

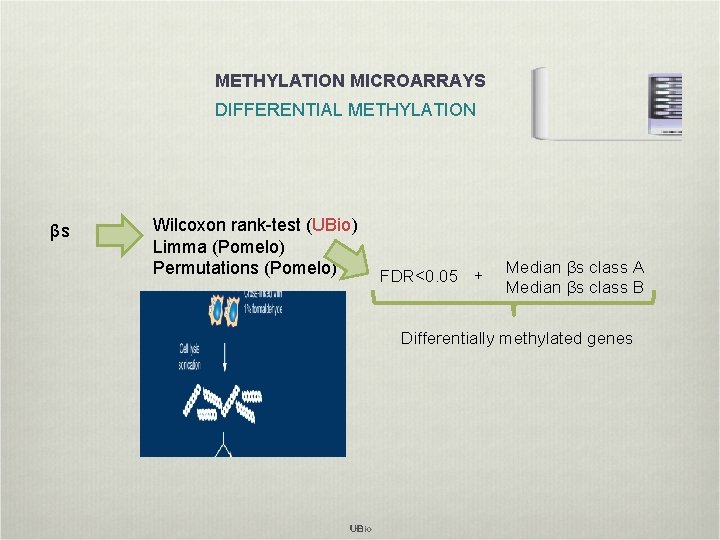
METHYLATION MICROARRAYS DIFFERENTIAL METHYLATION βs Wilcoxon rank-test (UBio) Limma (Pomelo) Permutations (Pomelo) FDR<0. 05 + Median βs class A Median βs class B Differentially methylated genes

Ch. IP on chip

Ch. IP on Chip We thank Chris Glass lab, UCSD, for the original slide

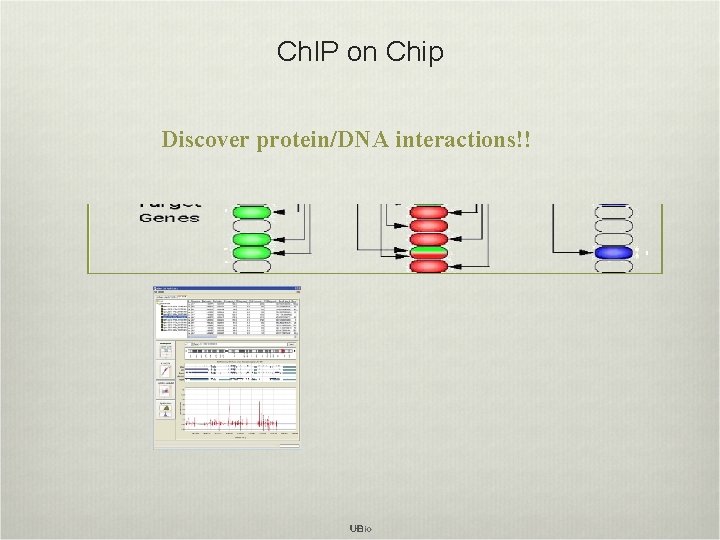
Ch. IP on Chip Discover protein/DNA interactions!!

Ch. IP on Chip software Chip Analytics WORKFLOW I. 1. Pre-normalization. Background substraction: Foreground – background Default: Median blank substraction Each channel – median negative controls 2. Normalization (dye-byas and interarray normalization) Default : Median dye-byas, median interarray. Recommended: Loess

Ch. IP on Chip software Chip Analytics WORKFLOW II. 3. Error modelling To identify which probes are most representative of binding events: P(X)=P-value of a single probe matching event P(X neighb)= Positive signals in a probe should be corroborated by the signals of probes that are its genomic neighbors, provided they are close enough P(Xneighb) follows a Gaussian distribution Both the P(X) and the P(Xneighb) values of a probe need to satisfy significance thresholds in order for a probe to be considered as representing a binding event

Ch. IP on Chip software Chip Analytics WORKFLOW III. 4. Segment identification (clusters of enriched probes) bp 5. Gene identification -Segment, Gene or Probe report (Gene or probe ID, Chr, Start, End, p(X)…)

Co. Cas http: //www. ciml. univ-mrs. fr/software/cocas/index. html Agilent platform Normalization QC Report Genome Visualization Peak Finder Benoukraf et al. Bioinformatics 2009.

Weeder: Motif discovery in sequences from co-regulated genes (single specie). Weeder. H: Motif discovery in sequences from homologous genes. Pscan: Motif discovery in sequences from co-regulated genes (JASPAR, TRANSFAC matrices) UBio training courses: See “Course on Introduction to Sequence Analysis”

Thanks ! Visit UBio web ! http: //bioinfo. cnio. es/