Gene expression and DNA microarrays No lab on

Gene expression and DNA microarrays • No lab on Thursday. • No class on Tuesday or Thursday next week – NCBI training Monday and Tuesday – Feb. 5 during class time - questions and answers period. • There will be a lab on Feb. 5. • Powerpoint lectures will be posted on website. • Reading assignment - two handouts – Chapter 3 has information regarding arrays

Genome of the week • E. coli O 157: H 7. • Causes haemorrhagic colitis – Initially identified in 1982 during an outbreak of severe bloody diarrhea. – Linked to contaminated ground beef from Michigan • 75, 000 cases per year

• Two different strains sequenced - link is to the U. S. sequence. • Major findings: – Comparison of E. coli O 157: H 7 with E. coli K-12 (common lab strain) found that the O 157: H 7 genome is ~ 1 Mb larger than K-12 and contains 1, 387 genes specific for O 157: H 7. – Genomes share a 4. 1 Mb backbone with species specific DNA interspersed throughout the genome • K-islands - specific to K-12 (0. 53 Mb) • O-islands - specific – Lateral transfer of DNA occurs much more frequently than previously thought. Especially high for enterobacteria. • O-island specific DNA encoded genes required for virulence and a large number of phage and phage associated genes.

Gene expression • What is gene expression? • Methods for measuring a single gene. – Northern Blots – Reporter genes – Quantitative RT-PCR • Operons, regulons, and stimulons. • DNA microarrays. – Expression profiling – Identifying protein binding sites. – Comparing gene content of different strains.

What is gene expression? • The amount of RNA produced from a gene. • Level of RNA produced from a gene is controlled by: – Transcription – Degradation • Transcriptome - Expressed transcripts in a cell under defined experimental conditions. – m. RNA(5 -10% of total RNA). – r. RNA, t. RNA - make up most of total RNA – sc. RNA (protein secretion), tm. RNA (rescue stalled ribosomes).

Analysis of gene expression at the single gene level. • Northern Blots – Measure RNA levels by hybridization of a labeled probe to total RNA. • Reporter Genes – Use of an enzyme to measure the amount of transcription from a promoter. • Quantitative RT-PCR. • Brief review pages - 158 -160.

Regulons and Stimulons • Operon - group of genes co-expressed on a single transcript. – One location of the genome • Regulon - genes that are regulated by a single transcription factor. – Genes and operons throughout the genome • Stimulon - collection of genes that are regulated in response to environmental changes. – Can be multiple regulons affected at once. • Regulatory network - alternative term for regulon.

Assaying the regulation of 1000 s of genes in a single experiment • DNA microarrays – DNA molecules printed at high density used to determine the level of RNA or DNA in a sample. – Can be thought of a “reverse Northern blots” • Other technologies (described in chapter 3). – SAGE – Microbeads

DNA Microarrays -Introduction • Spotted DNA arrays (glass slides) – Competitive binding of samples – Fluorescent detection - Cy 3 and Cy 5 – Small sample sizes (10 -30µl). – PCR or c. DNA arrays – Long oligonucleotide arrays • Better specificity, cheaper, easier to work with. • Short oligonucleotide arrays – ex. Affymetrix • DNA spotted onto nylon membranes (macroarrays)

Microarray experimental overview

Hybridization: basic concept The ability of two strands to hybridize is dependent on their complementarity. More complementarity=better hybridization

Labeling RNA or DNA with Cy 3 or Cy 5. • Cy 3 and Cy 5 - most often used fluorescent molecules used to label samples for microarray analysis. – Absorb light at one wavelength and emit at another. – Emission and Excitation spectra do not overlap significantly. – In arrays Cy 3 and Cy 5 are usually false colored green (Cy 3) and red (Cy 5) for ease of visualization.

More labeling • Direct incorporation - incorporates Cy 3 -or Cy 5 d. NTP directly into c. DNA – RNA to c. DNA - reverse transcriptase – DNA to DNA - DNA polymerase – Big problem - Cy 3 and Cy 5 are not incorporated with same efficiency. • Indirect incorporation - preferred method. – Incorporate an aminoallyl-d. UTP molecule during reverse transcription of RNA to c. DNA. – Chemically couple Cy 3 or Cy 5 dye after c. DNA is made. – Coupling is efficient with both dyes.

Applications of DNA microarrays • Expression profiling – Determining the relative levels of RNA in two or more samples. • DNA/DNA hybridizations – Investigate gene content between different strains – Determine gene dosage – 16 S arrays - microbial communities (being developed). • Identification of protein binding sites – Ch. IP-Chip. Immunoprecipitation of protein/DNA complexes. Assaying those interactions with microarrays.

B. subtilis DNA microarrays • PCR generated microarrays using custom primers (Sigma-Genosys). • Each PCR product represents a single gene. • 4074 genes of 4101 on the array. • Printed on Corning CMT-GAPS slides. • 4 E. coli controls, each represented 15 -20 times on the array.

How a DNA microarray works • Comparing the genome content of two B. subtilis strains. • The two strains differ only by the fact that JH 642 is lysogenized with the bacteriophage SPb. • JH 642 vs PY 79 genomic DNA hybridization. – PY 79 does not contain SPb. – SPb spots will be red.

JH 642 PY 79 Array size = 16 mm x 16 mm Spot size = 150 m. M
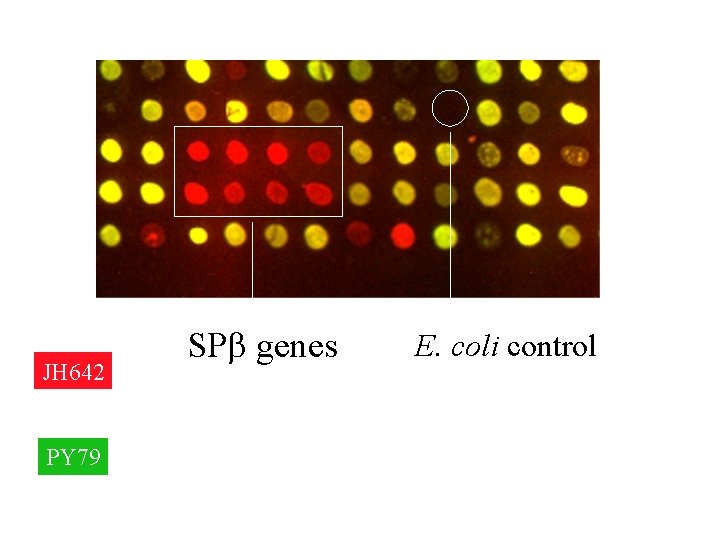
JH 642 PY 79 SPb genes E. coli control
- Slides: 18