Statistical Genomics Lecture 19 SUPER Zhiwu Zhang Washington















![Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr] Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr]](https://slidetodoc.com/presentation_image/28076e9b84215eb211d504fe90219363/image-27.jpg)
![Plots of power vs. FDR the. Color=rainbow(4) plot(fdr. mean[, 1], power , type="b", col=the. Plots of power vs. FDR the. Color=rainbow(4) plot(fdr. mean[, 1], power , type="b", col=the.](https://slidetodoc.com/presentation_image/28076e9b84215eb211d504fe90219363/image-28.jpg)

- Slides: 29

Statistical Genomics Lecture 19: SUPER Zhiwu Zhang Washington State University

Administration Homework 5, due April 13, Wednesday, 3: 10 PM Final exam: May 3, 120 minutes (3: 10 -5: 10 PM), 50

Read material Statistics (lecture slides) R programming(lecture slides) Genetics: GBS, populations structure, kinship Imputation GWAS: GLM, MLM, CMLM, ECMLM, SUPER, MLMM, EMMA, EMMAx/P 3 D, Farm. CPU, PC+K GS: g. BLUP

Outline Kinship based on QTN Confounding between QTN and kinship Complimentary kinship SUPER

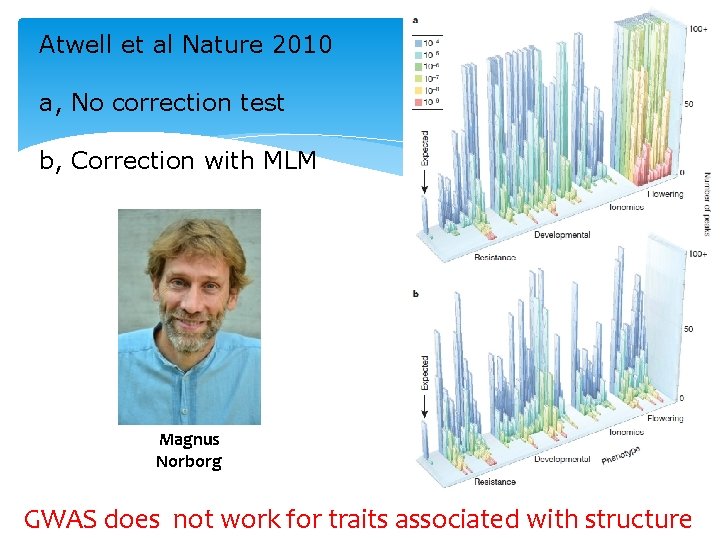
Atwell et al Nature 2010 a, No correction test b, Correction with MLM Magnus Norborg GWAS does not work for traits associated with structure

MLM Tree John Pollak Ivan Brian Kennedy. Mao Dick Quaas Larry Schaeffer Charles Henderson Jim Wilton 1983

Growing tree: Method and software Meng Li CMLM Xiaolei Liu Farm. CPU Yumei Yang i. BLUP Meng Huang BLINK Qishan Wang SUPER Alex Lipka GAPIT g. BLUP CMLM P 3 D Jiabo Wang c. BLUP

More covariates observation mean b= [ b 0 y [1 PC 2 b 1 x 1 SNP b 2 ] u= x 2 ] =X y = Xb + Zu +e Ind 1 [ u 1 1 0 Ind 2 u 2 0 1 … … 0 0 … … Z Ind 9 Ind 10 u 9 u 10 ] 0 0 1

Variance in MLM y = Xb + Zu + e Var(y)=V=Var(u)+Var(e) u prediction: Best Linear Unbiased Prediction, BLUP) b prediction: Best Linear Unbiased Estimate, BLUE)

Kinship defined by single marker Sensitive Resistance S 1 S 2 S 3 S 4 R 1 R 2 R 3 R 4 S 1 1 1 0 0 S 2 1 1 0 0 S 3 1 1 0 0 S 4 1 1 0 0 R 1 0 0 1 1 R 2 0 0 1 1 R 3 0 0 1 1 R 4 0 0 1 1 Adding additional markers bluer the picture

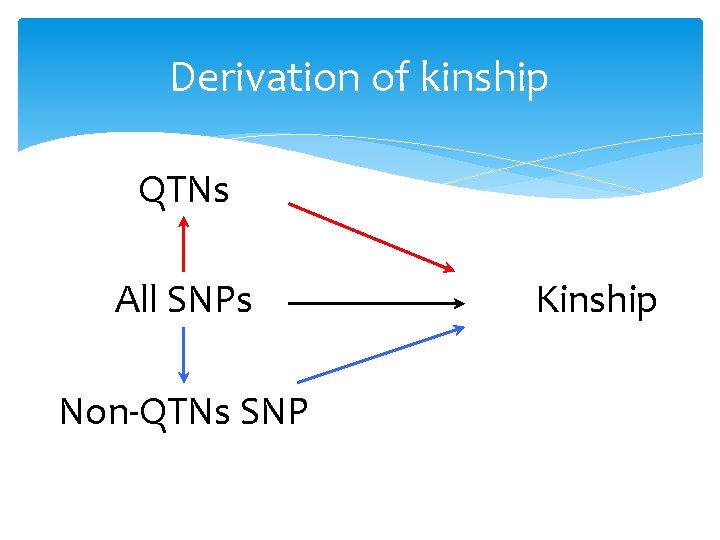
Derivation of kinship QTNs All SNPs Non-QTNs SNP Kinship

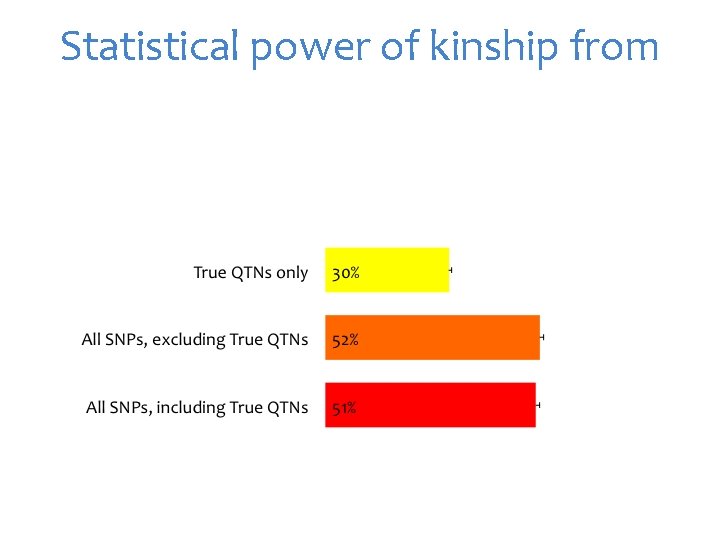
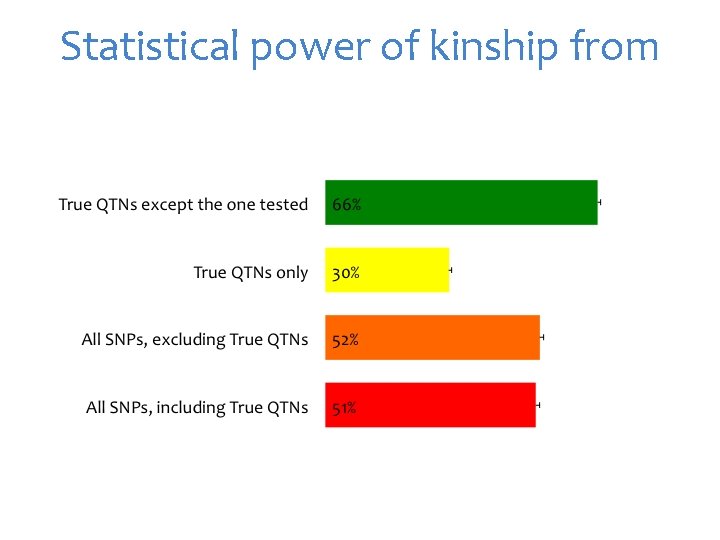
Statistical power of kinship from

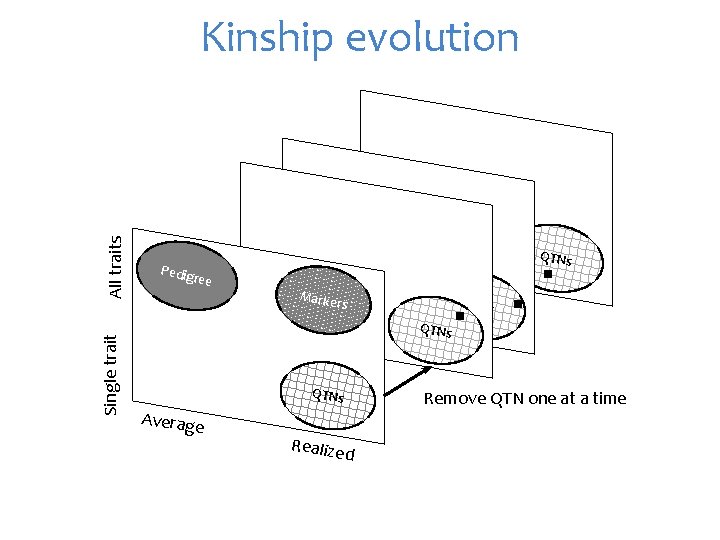
Single trait All traits Kinship evolution QTNs Pedig ree Marke rs QTNs Average Realize Remove QTN one at a time d

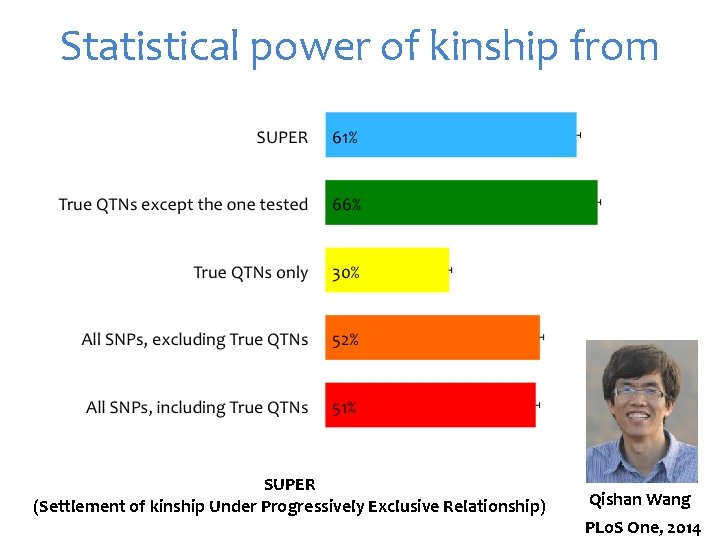
Statistical power of kinship from

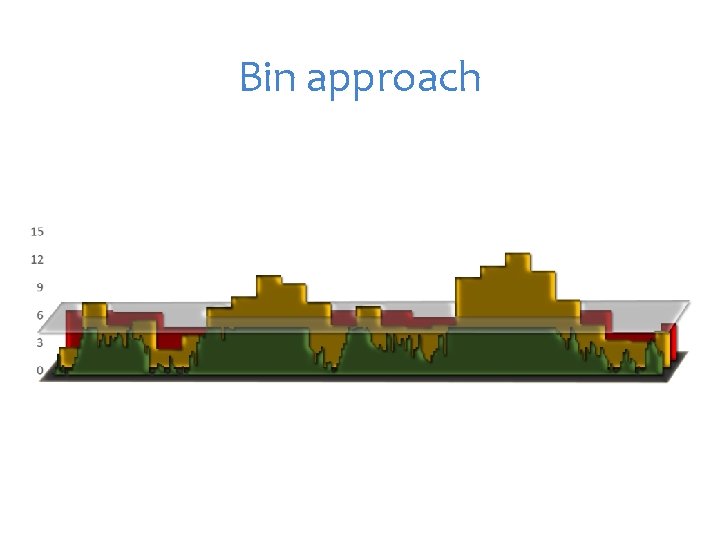
Bin approach

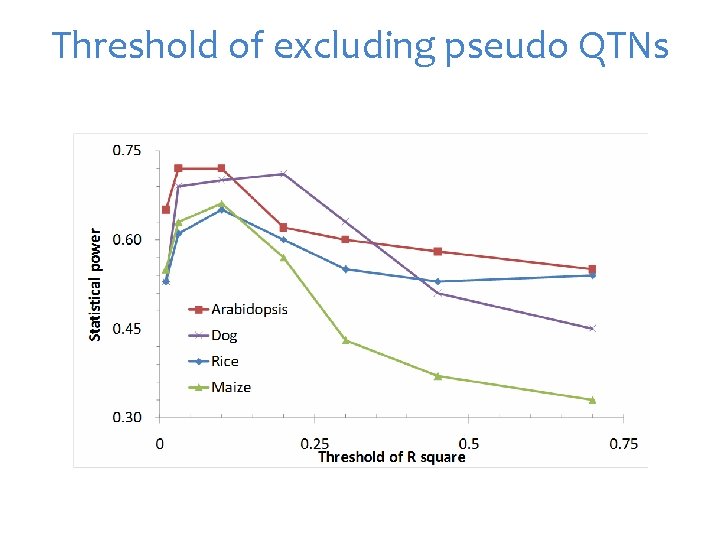
Mimic QTN-1 1. Choose t associated SNPs as QTNs each represent an interval of size s. 2. Build kinship from the t QTNs 3. Optimization on t and s 4. For a SNP, remove the QTNs in LD with the SNP, e. g. R square > 1% 5. Use the remaining QTNs to build kinship for testing the SNP

Statistical power of kinship from SUPER (Settlement of kinship Under Progressively Exclusive Relationship) Qishan Wang PLo. S One, 2014

Threshold of excluding pseudo QTNs

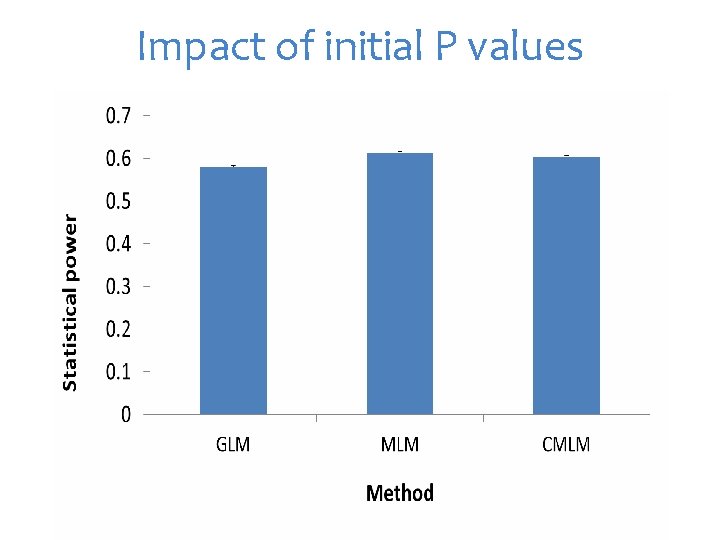
Impact of initial P values

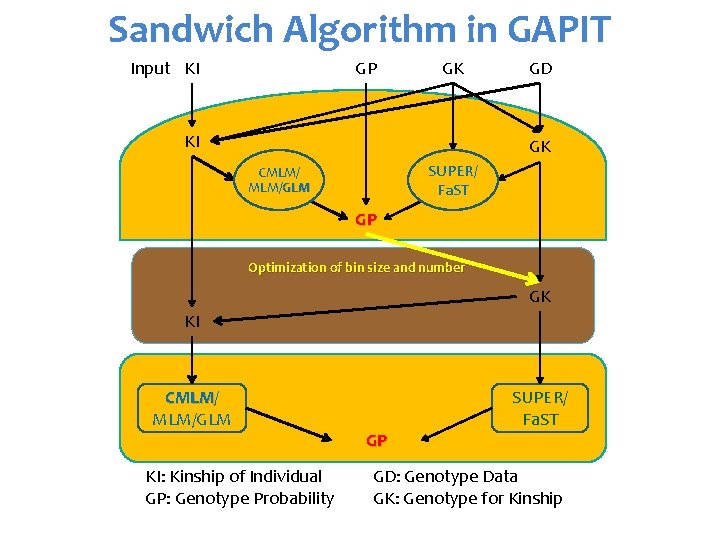
Sandwich Algorithm in GAPIT Input KI GP GK KI GD GK SUPER/ Fa. ST CMLM/GLM GP Optimization of bin size and number GK KI CMLM/ CMLM MLM/GLM KI: Kinship of Individual GP: Genotype Probability GP SUPER/ Fa. ST GD: Genotype Data GK: Genotype for Kinship

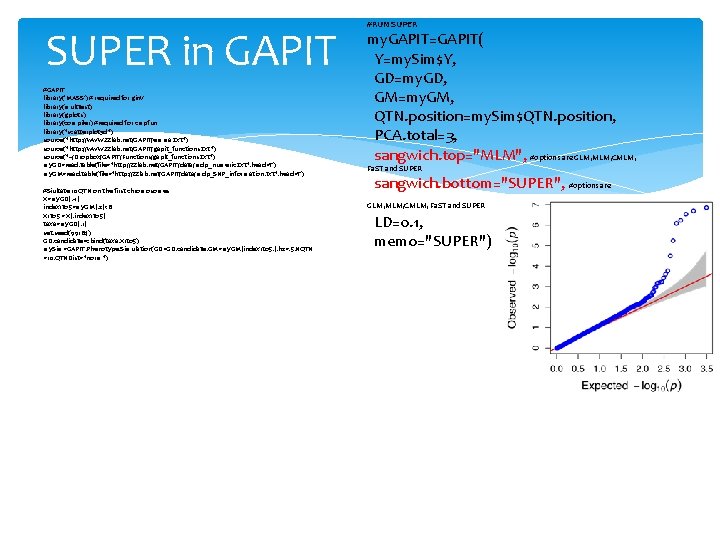
SUPER in GAPIT #GAPIT library('MASS') # required for ginv library(multtest) library(gplots) library(compiler) #required for cmpfun library("scatterplot 3 d") source("http: //www. zzlab. net/GAPIT/emma. txt") source("http: //www. zzlab. net/GAPIT/gapit_functions. txt") source("~/Dropbox/GAPIT/Functions/gapit_functions. txt") my. GD=read. table(file="http: //zzlab. net/GAPIT/data/mdp_numeric. txt", head=T) my. GM=read. table(file="http: //zzlab. net/GAPIT/data/mdp_SNP_information. txt", head=T) #Siultate 10 QTN on the first chromosomes X=my. GD[, -1] index 1 to 5=my. GM[, 2]<6 X 1 to 5 = X[, index 1 to 5] taxa=my. GD[, 1] set. seed(99164) GD. candidate=cbind(taxa, X 1 to 5) my. Sim=GAPIT. Phenotype. Simulation(GD=GD. candidate, GM=my. GM[index 1 to 5, ], h 2=. 5, NQTN =10, QTNDist="norm") #RUN SUPER my. GAPIT=GAPIT( Y=my. Sim$Y, GD=my. GD, GM=my. GM, QTN. position=my. Sim$QTN. position, PCA. total=3, sangwich. top="MLM", #options are GLM, MLM, CMLM, Fa. ST and SUPER sangwich. bottom="SUPER", #options are GLM, MLM, CMLM, Fa. ST and SUPER LD=0. 1, memo="SUPER")

GAPIT. FDR. Type. I Function my. Stat=GAPIT. FDR. Type. I(WS=c(1 e 0, 1 e 3, 1 e 4, 1 e 5), GM=my. GM, seq. QTN=my. Sim$QTN. position, GWAS=my. GAPIT$GWAS)

Return

Area Under Curve (AUC) par(mfrow=c(1, 2), mar = c(5, 2, 5, 2)) plot(my. Stat$FDR[, 1], my. Stat$Power, type="b") plot(my. Stat$Type. I[, 1], my. Stat$Power, type="b")

Replicates nrep=3 set. seed(99164) stat. Rep=replicate(nrep, { my. Sim=GAPIT. Phenotype. Simulation(GD=GD. candidate, GM=my. GM[index 1 to 5, ], h 2=. 5, NQTN=10, QTNDist="norm") my. GAPIT=GAPIT( Y=my. Sim$Y, GD=my. GD, GM=my. GM, QTN. position=my. Sim$QTN. position, PCA. total=3, sangwich. top="MLM", #options are GLM, MLM, CMLM, Fa. ST and SUPER sangwich. bottom="SUPER", #options are GLM, MLM, CMLM, Fa. ST and SUPER LD=0. 1, memo="SUPER") my. Stat=GAPIT. FDR. Type. I(WS=c(1 e 0, 1 e 3, 1 e 4, 1 e 5), GM=my. GM, seq. QTN=my. Sim$QT N. position, GWAS=my. GAPIT$GWAS) })

str(stat. Rep)
![Means over replicates powerstat Rep2 FDR s fdrseq3 lengthstat Rep 7 fdrstat Reps fdr Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr]](https://slidetodoc.com/presentation_image/28076e9b84215eb211d504fe90219363/image-27.jpg)
Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr] fdr. mean=Reduce ("+", fdr) / length(fdr) #AUC: power vs. FDR s. auc. fdr=seq(6, length(stat. Rep), 7) auc. fdr=stat. Rep[s. auc. fdr] auc. fdr. mean=Reduce ("+", auc. fdr) / length(auc. fdr)
![Plots of power vs FDR the Colorrainbow4 plotfdr mean 1 power typeb colthe Plots of power vs. FDR the. Color=rainbow(4) plot(fdr. mean[, 1], power , type="b", col=the.](https://slidetodoc.com/presentation_image/28076e9b84215eb211d504fe90219363/image-28.jpg)
Plots of power vs. FDR the. Color=rainbow(4) plot(fdr. mean[, 1], power , type="b", col=the. Color [1], xlim=c(0, 1)) for(i in 2: ncol(fdr. mean)){ lines(fdr. mean[, i], power , type="b", col= the. Color [i]) }

Highlight Kinship based on QTN Confounding between QTN and kinship Complimentary kinship SUPER