Genomewide Functional Linkage Maps Methods for inferring functional












- Slides: 43

Genome-wide Functional Linkage Maps Methods for inferring functional linkages: Complexes, Pathways Rosetta stone Phylogenetic profiles Gene neighbors Operon method (Microarray method) The Genome-wide functional linkage Map in M. tb Assessing accuracy of functional linkages Functional linkages in structural genomics Analyzing parallel pathways The DIP and Pro. Links databases

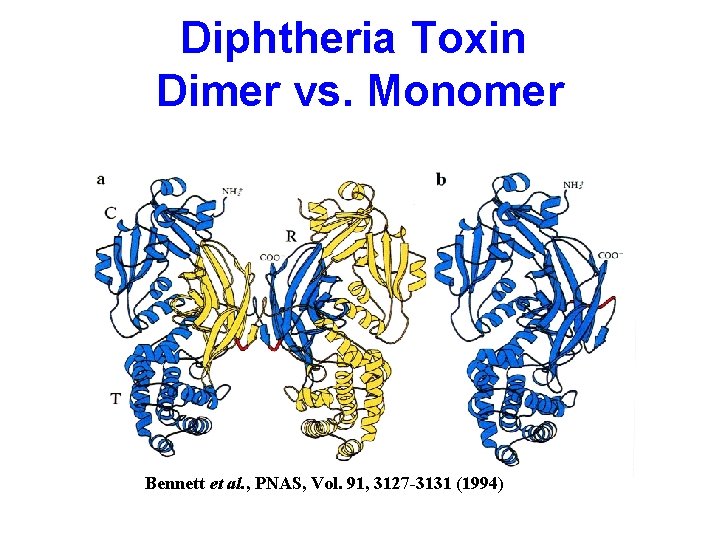
Diphtheria Toxin Dimer vs. Monomer Bennett et al. , PNAS, Vol. 91, 3127 -3131 (1994)


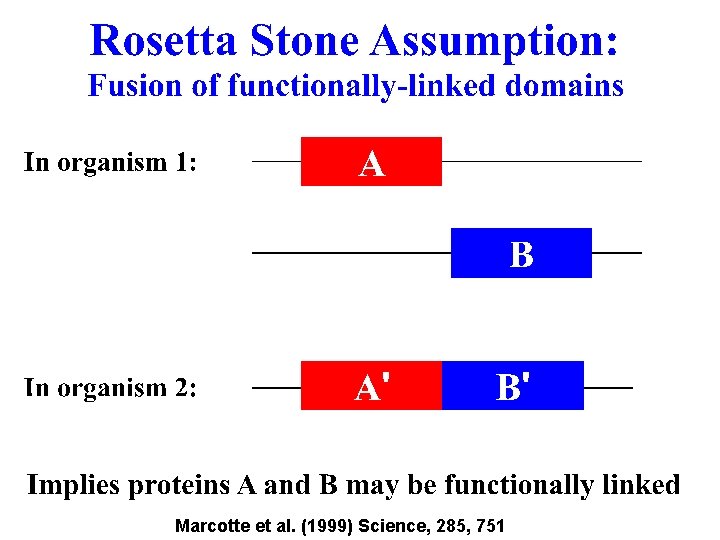
Marcotte et al. (1999) Science, 285, 751

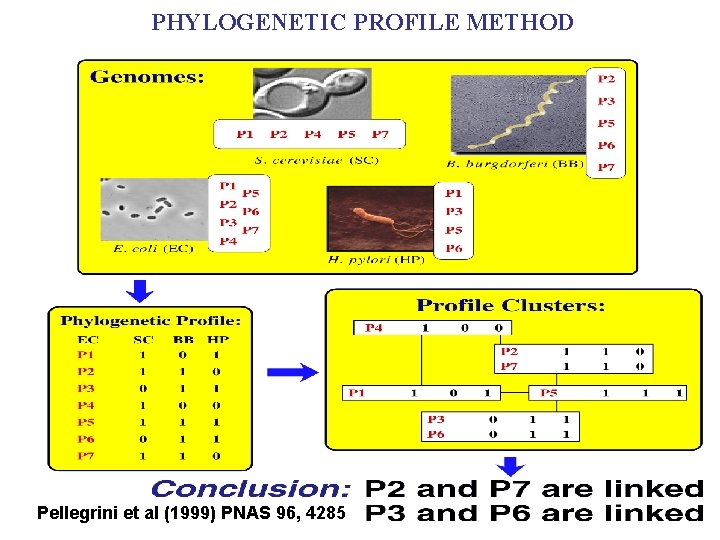
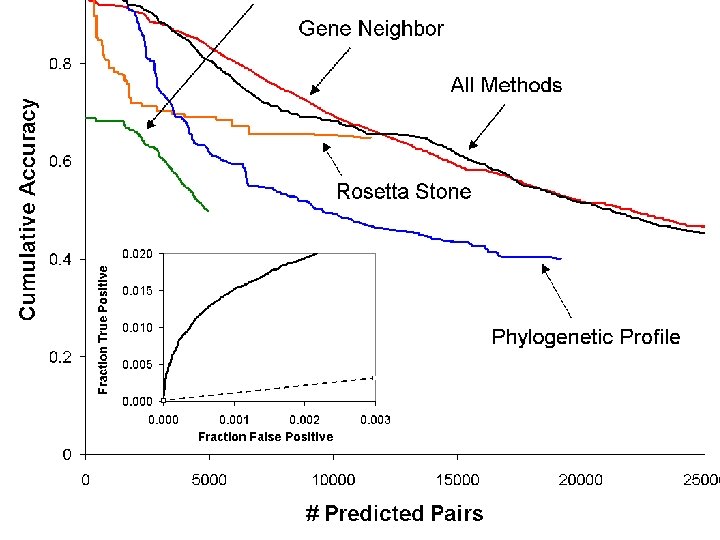
PHYLOGENETIC PROFILE METHOD Pellegrini et al (1999) PNAS 96, 4285

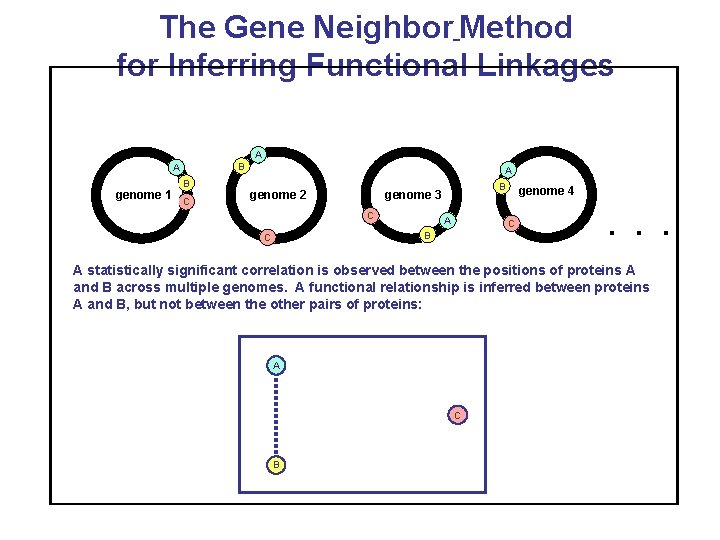
The Gene Neighbor Method for Inferring Functional Linkages A B A genome 1 B C A genome 2 B genome 3 C A C B C genome 4 . . . A statistically significant correlation is observed between the positions of proteins A and B across multiple genomes. A functional relationship is inferred between proteins A and B, but not between the other pairs of proteins: A C B

OPERON or GENE CLUSTER method of inferring functional linkages in the genome of Mycobacterium tuberculosis gene A bbbb gene B gene C The 100 bp threshold is chosen because it gives the broadest coverage consistent with high accuracy Research of Michael Strong

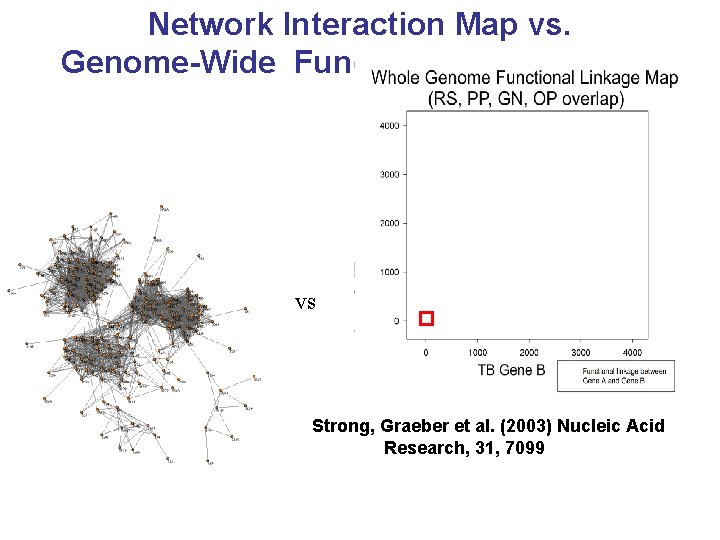
Network Interaction Map vs. Genome-Wide Functional Linkage Map vs Strong, Graeber et al. (2003) Nucleic Acid Research, 31, 7099

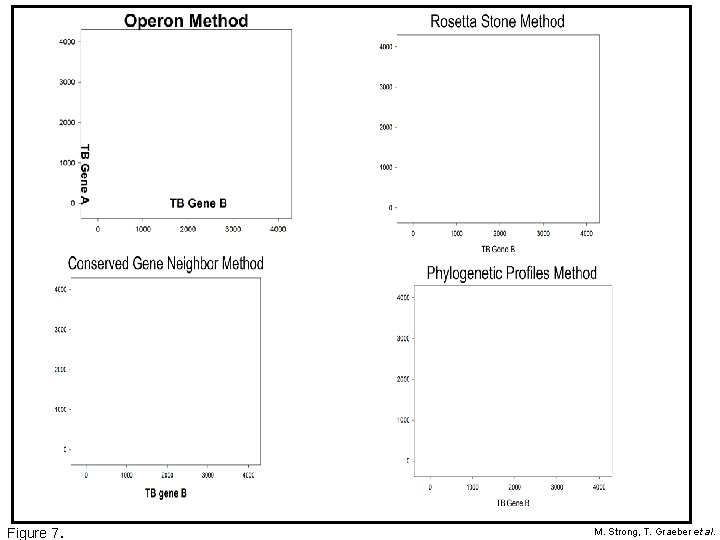
Figure 7. M. Strong, T. Graeber et al.

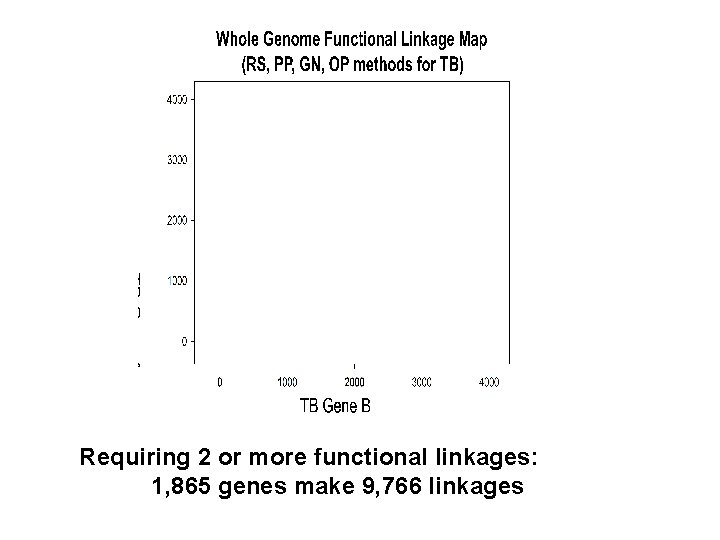
Requiring 2 or more functional linkages: 1, 865 genes make 9, 766 linkages

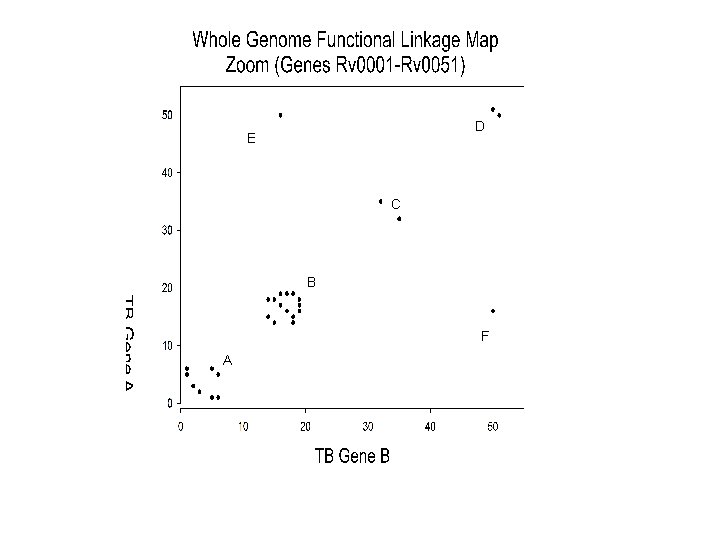
D E C B F A

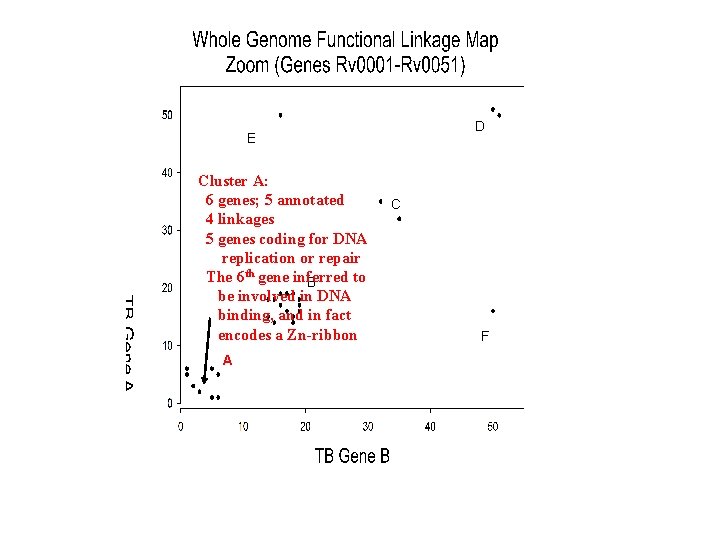
D E Cluster A: 6 genes; 5 annotated 4 linkages 5 genes coding for DNA replication or repair The 6 th gene inferred to B be involved in DNA binding, and in fact encodes a Zn-ribbon A C F

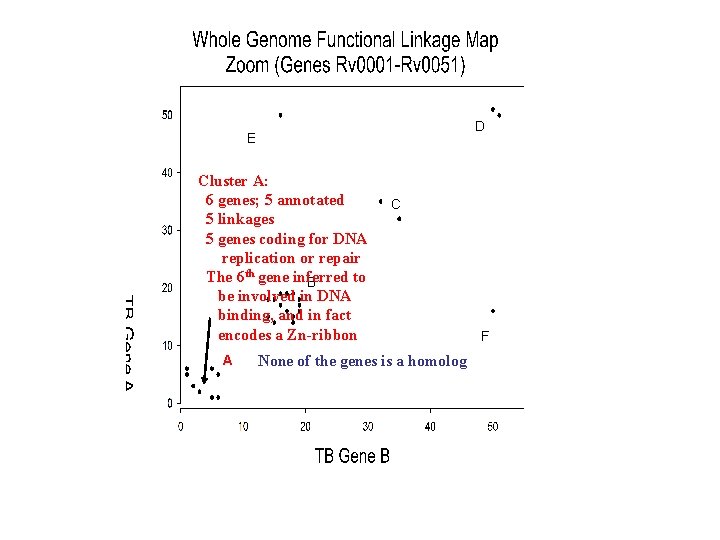
D E Cluster A: 6 genes; 5 annotated 5 linkages 5 genes coding for DNA replication or repair The 6 th gene inferred to B be involved in DNA binding, and in fact encodes a Zn-ribbon A C None of the genes is a homolog F

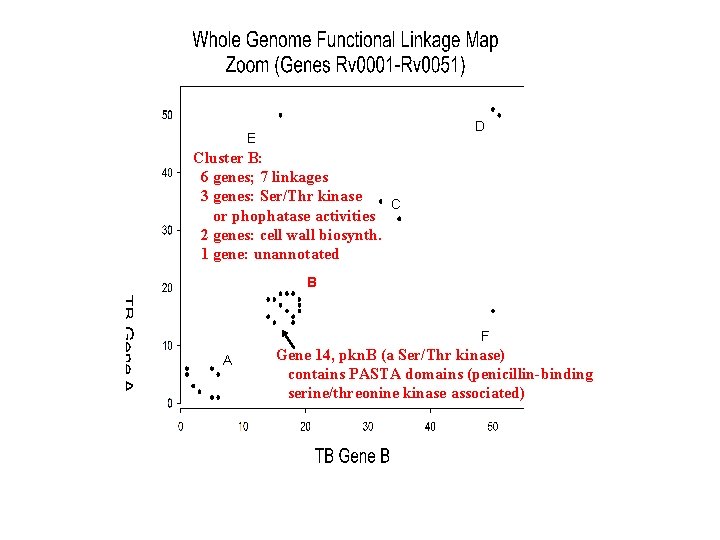
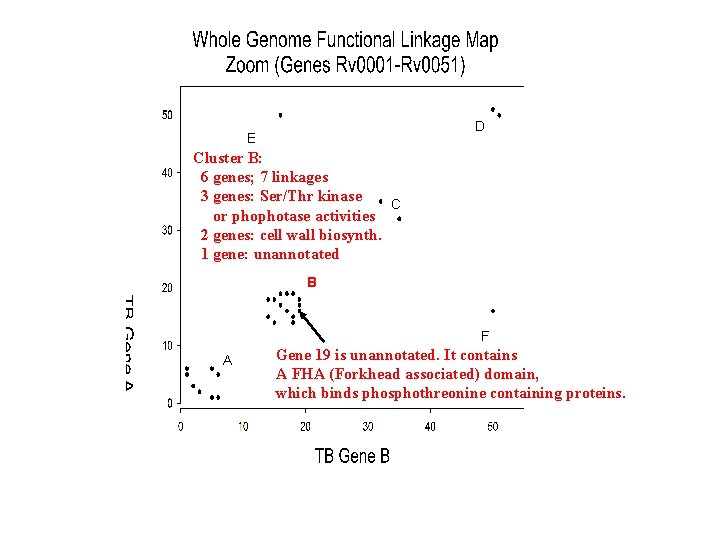
D E Cluster B: 6 genes; 7 linkages 3 genes: Ser/Thr kinase C or phophatase activities 2 genes: cell wall biosynth. 1 gene: unannotated B F A Gene 14, pkn. B (a Ser/Thr kinase) contains PASTA domains (penicillin-binding serine/threonine kinase associated)

D E Cluster B: 6 genes; 7 linkages 3 genes: Ser/Thr kinase C or phophotase activities 2 genes: cell wall biosynth. 1 gene: unannotated B F A Gene 19 is unannotated. It contains A FHA (Forkhead associated) domain, which binds phosphothreonine containing proteins.

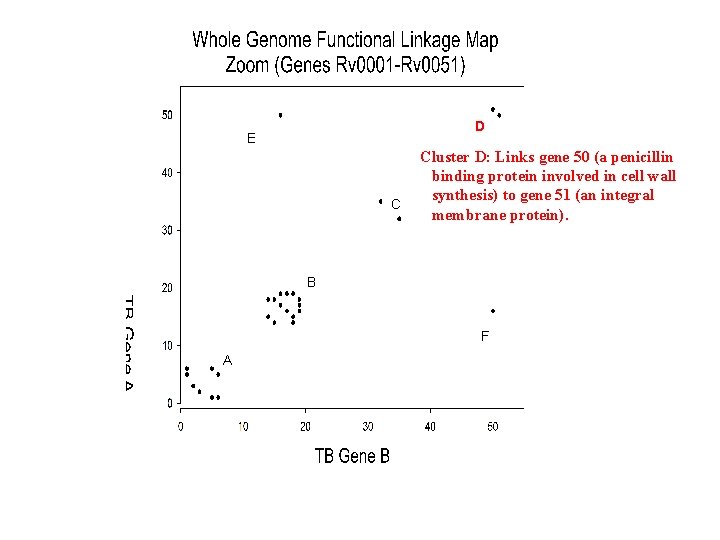
D E C Cluster D: Links gene 50 (a penicillin binding protein involved in cell wall synthesis) to gene 51 (an integral membrane protein). B F A

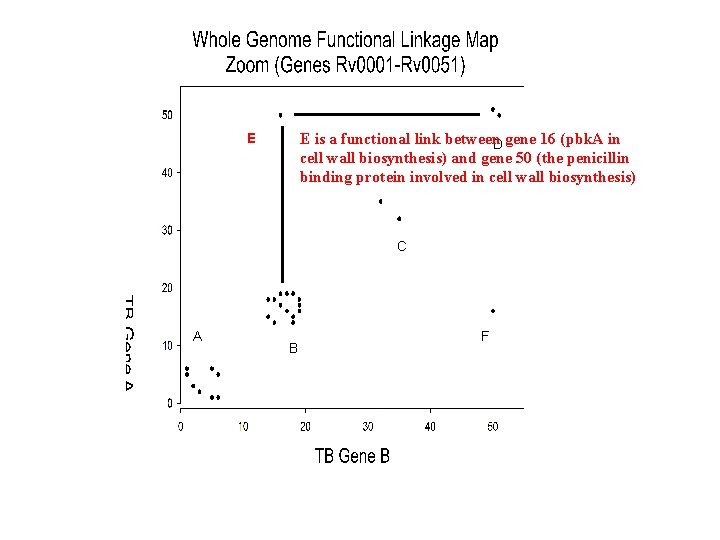
E is a functional link between. D gene 16 (pbk. A in cell wall biosynthesis) and gene 50 (the penicillin binding protein involved in cell wall biosynthesis) E C A B F

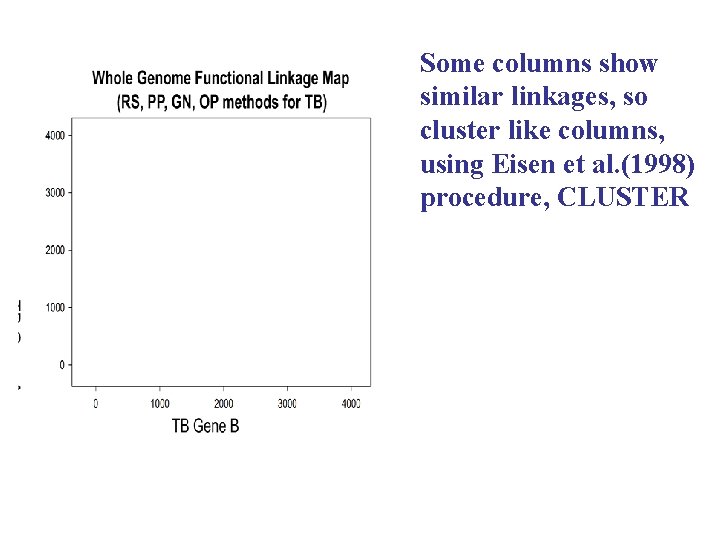
Some columns show similar linkages, so cluster like columns, using Eisen et al. (1998) procedure, CLUSTER

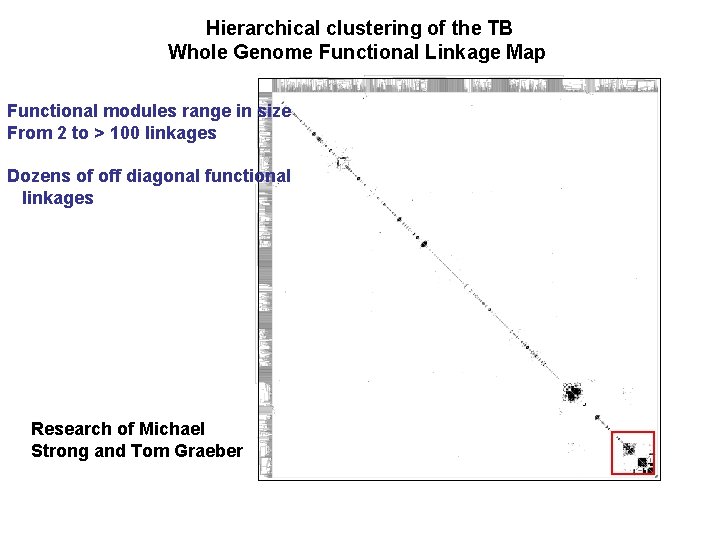
Hierarchical clustering of the TB Whole Genome Functional Linkage Map Functional modules range in size From 2 to > 100 linkages Dozens of off diagonal functional linkages Research of Michael Strong and Tom Graeber

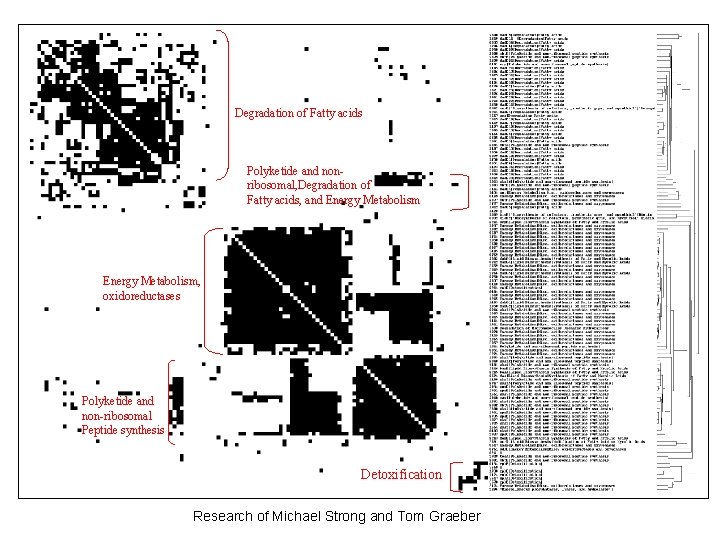
Degradation of Fatty acids Polyketide and nonribosomal, Degradation of Fatty acids, and Energy Metabolism, oxidoreductases Polyketide and non-ribosomal Peptide synthesis Detoxification Research of Michael Strong and Tom Graeber

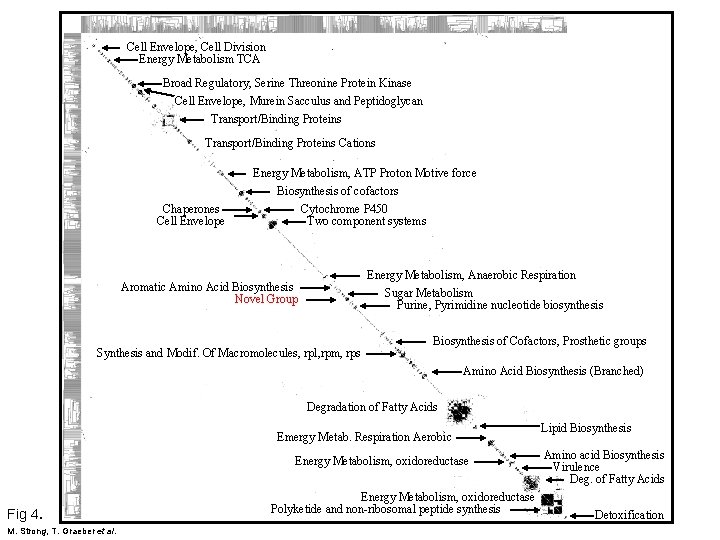
Cell Envelope, Cell Division Energy Metabolism TCA Broad Regulatory, Serine Threonine Protein Kinase Cell Envelope, Murein Sacculus and Peptidoglycan Transport/Binding Proteins Cations Chaperones Cell Envelope Energy Metabolism, ATP Proton Motive force Biosynthesis of cofactors Cytochrome P 450 Two component systems Energy Metabolism, Anaerobic Respiration Sugar Metabolism Purine, Pyrimidine nucleotide biosynthesis Aromatic Amino Acid Biosynthesis Novel Group Synthesis and Modif. Of Macromolecules, rpl, rpm, rps Biosynthesis of Cofactors, Prosthetic groups Amino Acid Biosynthesis (Branched) Degradation of Fatty Acids Emergy Metab. Respiration Aerobic Energy Metabolism, oxidoreductase Fig 4. M. Strong, T. Graeber et al. Energy Metabolism, oxidoreductase Polyketide and non-ribosomal peptide synthesis Lipid Biosynthesis Amino acid Biosynthesis Virulence Deg. of Fatty Acids Detoxification

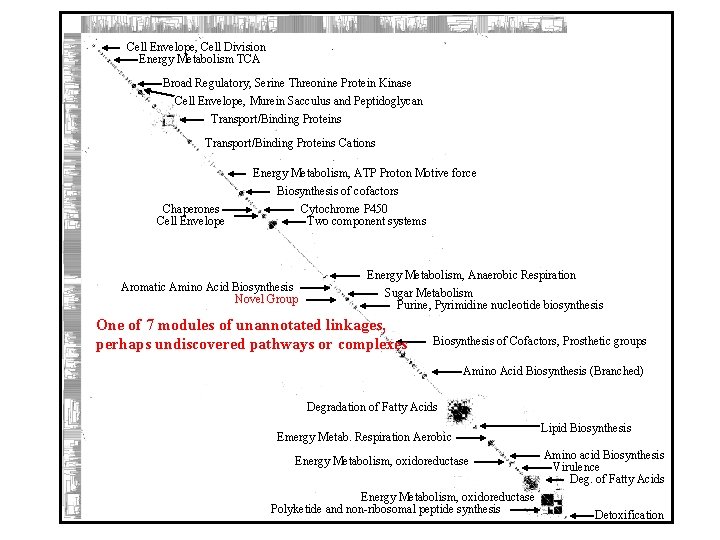
Cell Envelope, Cell Division Energy Metabolism TCA Broad Regulatory, Serine Threonine Protein Kinase Cell Envelope, Murein Sacculus and Peptidoglycan Transport/Binding Proteins Cations Chaperones Cell Envelope Energy Metabolism, ATP Proton Motive force Biosynthesis of cofactors Cytochrome P 450 Two component systems Aromatic Amino Acid Biosynthesis Novel Group Energy Metabolism, Anaerobic Respiration Sugar Metabolism Purine, Pyrimidine nucleotide biosynthesis One of 7 modules of unannotated linkages, perhaps undiscovered pathways or complexes Biosynthesis of Cofactors, Prosthetic groups Amino Acid Biosynthesis (Branched) Degradation of Fatty Acids Emergy Metab. Respiration Aerobic Energy Metabolism, oxidoreductase Polyketide and non-ribosomal peptide synthesis Lipid Biosynthesis Amino acid Biosynthesis Virulence Deg. of Fatty Acids Detoxification

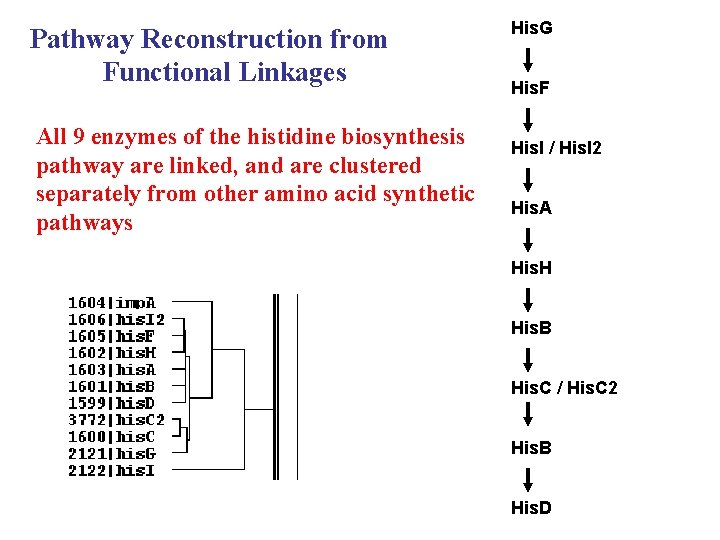
Pathway Reconstruction from Functional Linkages All 9 enzymes of the histidine biosynthesis pathway are linked, and are clustered separately from other amino acid synthetic pathways His. G His. F His. I / His. I 2 His. A His. H His. B His. C / His. C 2 His. B His. D

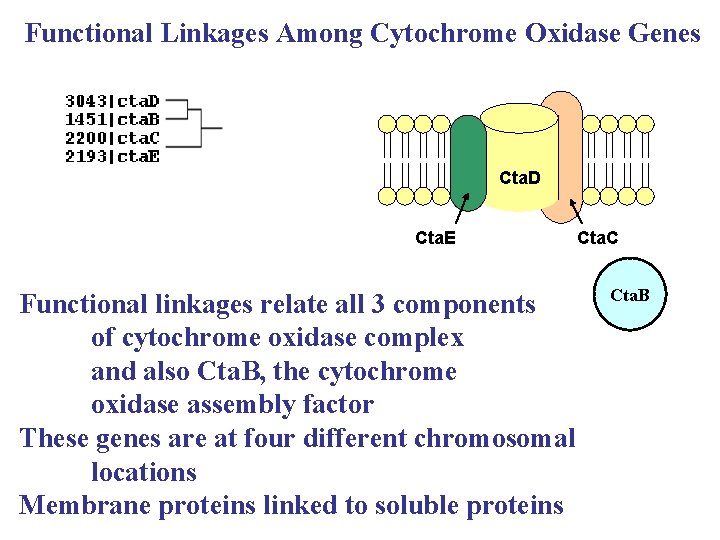
Functional Linkages Among Cytochrome Oxidase Genes Cta. D Cta. E Functional linkages relate all 3 components of cytochrome oxidase complex and also Cta. B, the cytochrome oxidase assembly factor These genes are at four different chromosomal locations Membrane proteins linked to soluble proteins Cta. C Cta. B

Quantitative Assessment of Inferred Protein Complexes Research of Edward Marcotte, Matteo Pellegrini, Michael Thompson and Todd Yeates

Calculating Probabilities of Coevolution Phylogenetic Profile Rosetta Stone N= number of fully sequenced genomes n= number of homologs of protein A m = number of homologs of protein B k = number of genomes shared in common Gene Neighbor X= fractional separation of genes Operon n = intergenic separation

Combining Inferences of Co. Evolution from 4 Methods We use a Bayesian approach to combine the probabilities from the four methods to arrive at a single probability that two proteins co-evolve: where positive pairs are proteins with common pathway annotation and negative pairs are proteins with different annotation

Pro. Links Database www. dip. doembi. ucla. edu/pronav ~ 10, 000 Functional Linkages inferred from 83 fully sequenced genomes

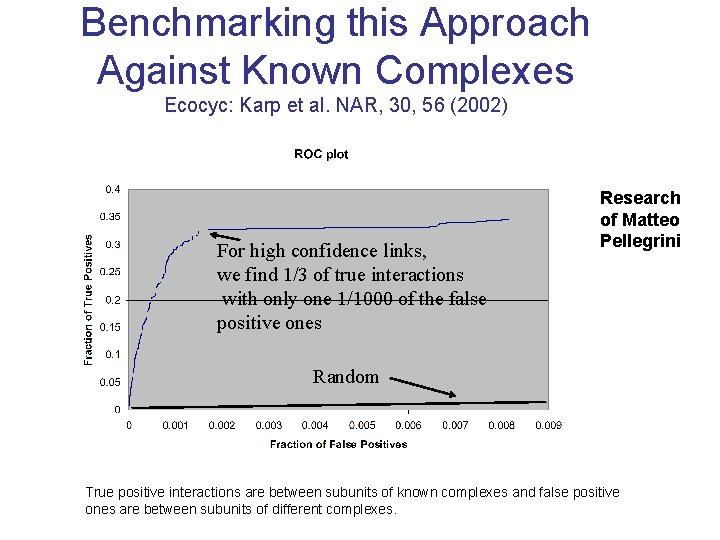
Benchmarking this Approach Against Known Complexes Ecocyc: Karp et al. NAR, 30, 56 (2002) For high confidence links, we find 1/3 of true interactions with only one 1/1000 of the false positive ones Research of Matteo Pellegrini Random True positive interactions are between subunits of known complexes and false positive ones are between subunits of different complexes.

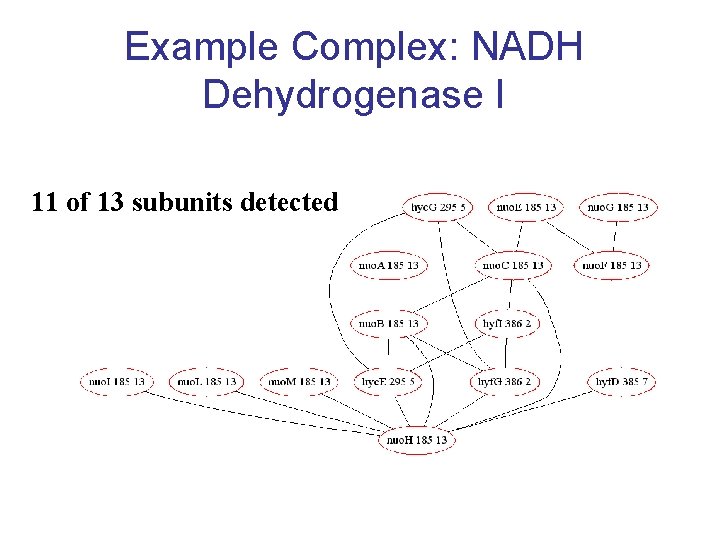
Example Complex: NADH Dehydrogenase I 11 of 13 subunits detected

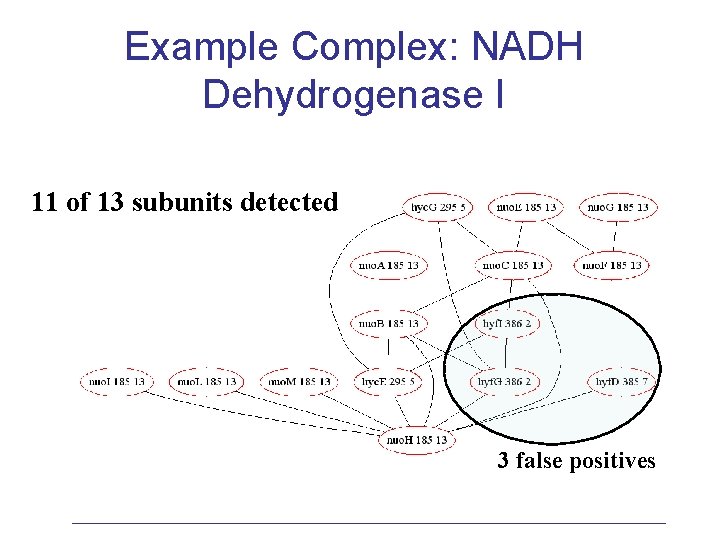
Example Complex: NADH Dehydrogenase I 11 of 13 subunits detected 3 false positives

From Inferred Protein Linkages to Structures of Complexes Research of Michael Strong, Shuishu Wang, Markus Kauffman

The Problem of PE and PPE Proteins in M. tb PE, PE-PGRS, and PPE Proteins in M. tuberculosis 38 PE proteins; 61 PE-PGRS proteins; 68 PPE proteins Together compromise about 5 % of the genome No function is known, but some appear to be membrane bound No structure is known: always insoluble when expressed Goal: use functional linkages to predict a complex between a PE and a PPE protein: express complex, and determine its structure Research of Shuishu Wang and Michael Strong

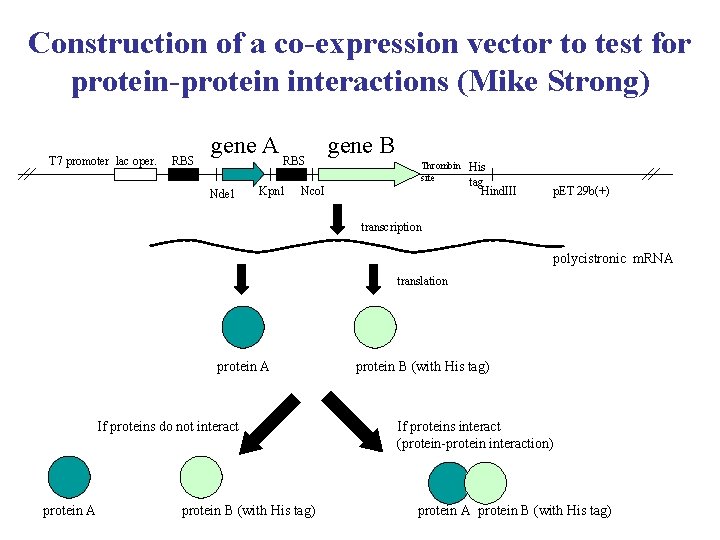
Construction of a co-expression vector to test for protein-protein interactions (Mike Strong) T 7 promoter lac oper. RBS gene A Nde 1 RBS Kpn 1 gene B Thrombin site Nco. I His tag Hind. III p. ET 29 b(+) transcription polycistronic m. RNA translation protein A If proteins do not interact protein A protein B (with His tag) If proteins interact (protein-protein interaction) protein A protein B (with His tag)

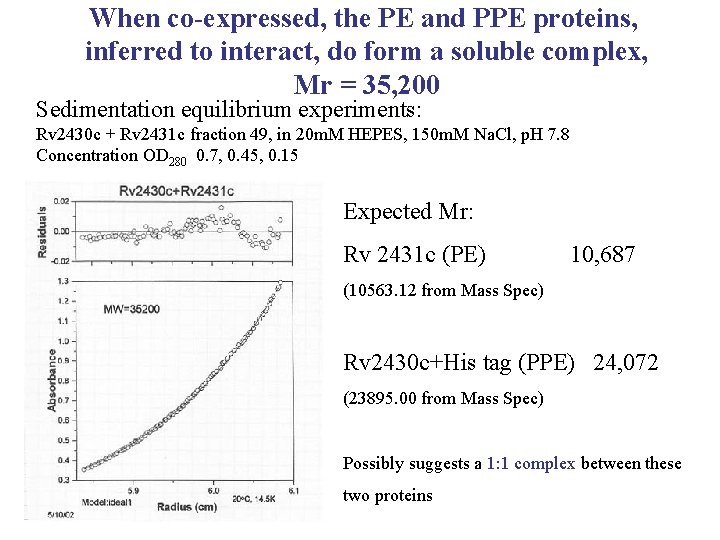
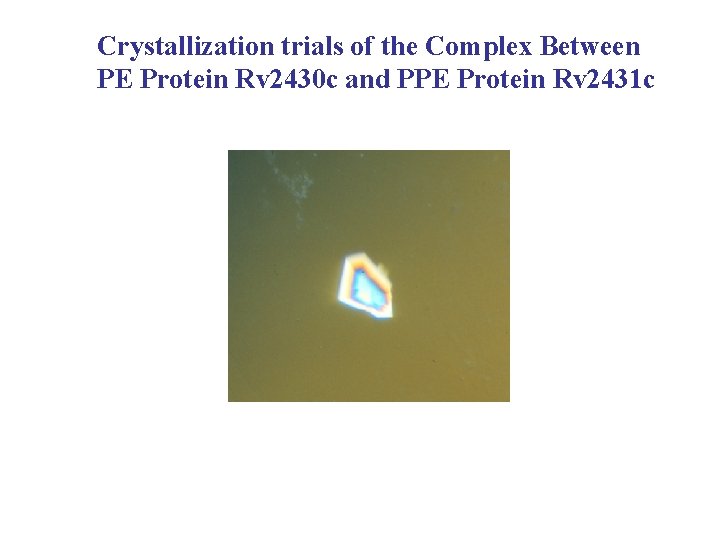
When co-expressed, the PE and PPE proteins, inferred to interact, do form a soluble complex, Mr = 35, 200 Sedimentation equilibrium experiments: Rv 2430 c + Rv 2431 c fraction 49, in 20 m. M HEPES, 150 m. M Na. Cl, p. H 7. 8 Concentration OD 280 0. 7, 0. 45, 0. 15 Expected Mr: Rv 2431 c (PE) 10, 687 (10563. 12 from Mass Spec) Rv 2430 c+His tag (PPE) 24, 072 (23895. 00 from Mass Spec) Possibly suggests a 1: 1 complex between these two proteins

Crystallization trials of the Complex Between PE Protein Rv 2430 c and PPE Protein Rv 2431 c

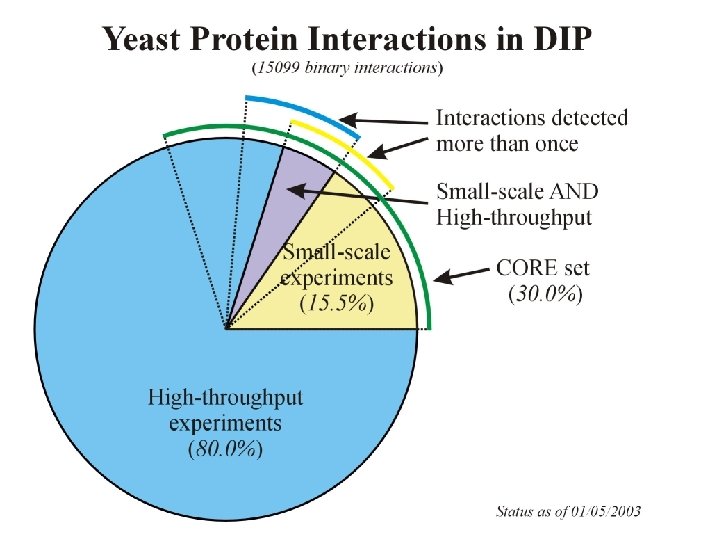
Database of Interacting Proteins www. dip. doe-mbi. ucla. edu Experimentally detected interactions from the scientific literature Currently ~ 44, 000 interactions

The DIP Database DOE-MBI LSBMM, UCLA


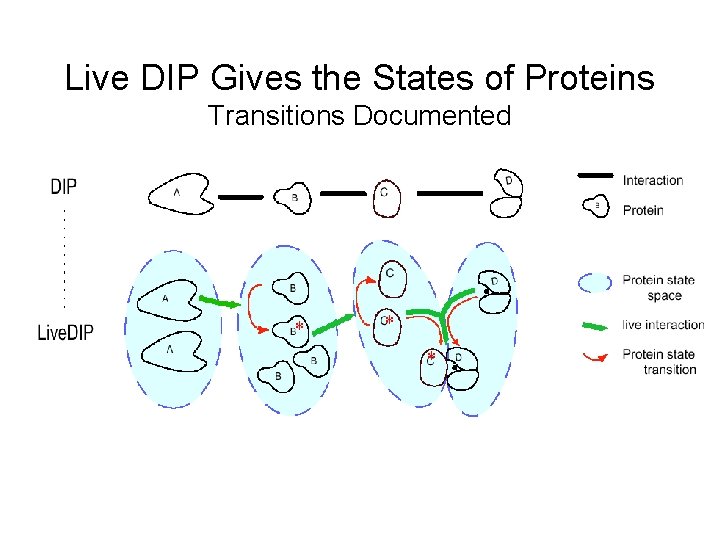
Live DIP Gives the States of Proteins Transitions Documented * * *

Pro. Links Database and the Protein Navigator • Contains some 10, 000 inferred functional linkages from 83 genomes • Available at www. doe-mbi. ucla. edu • Soon to be expanded to 250 fully sequenced genomes • Eventually to be reconciled with DIP

Summary Many functional linkages are revealed from genomic and microarray data (high coverage) Validity of functional linkages can be assessed by comparison to known complexes, and to expression data, and by keyword recovery Clustered genome-wide functional maps can reveal and organize information on complexes and pathways Functional linkages can reveal protein complexes suitable for structural studies B C A protein’s function is defined by Y A X the cellular context of its linkages V Z

Protein Interactions Analysis of M. tb. Genome Michael Strong Whole Genome Interaction Maps Michael Strong & Tom Graeber Methods of Inferring Interactions Edward Marcotte, Matteo Pellegrini, Todd Yeates Michael Thompson, Richard Llwellyn Database of Interacting Proteins Lukasz Salwinski, Joyce Duan, Ioannis Xenarios, Robert Riley, Christopher Miller Parallel pathways Huiying Li