Linkage Mapping Part 1 PBG 430 Linkage Linkage

Linkage Mapping – Part 1 PBG 430

Linkage § Linkage is the non-random association of two or more phenotypes or sequences in a population (genes/sequences are (almost) always inherited together) § Biological explanation for linkage: genes or sequences are closely located in the same chromosome § Loci in the same chromosome are said to be “members of the same linkage group” § The number of linkage groups corresponds to n Plant Formula Number of Linkage Groups Fragaria vesca 2 n = 2 x = 14 7 Hordeum vulgare 2 n = 2 X = 14 7 Triticum aestivum 2 n = 6 x = 42 21 Population Yellow/16 petals or Pink/8 petals Pleiotropy or linkage? 2 genes flower color is linked with petal number
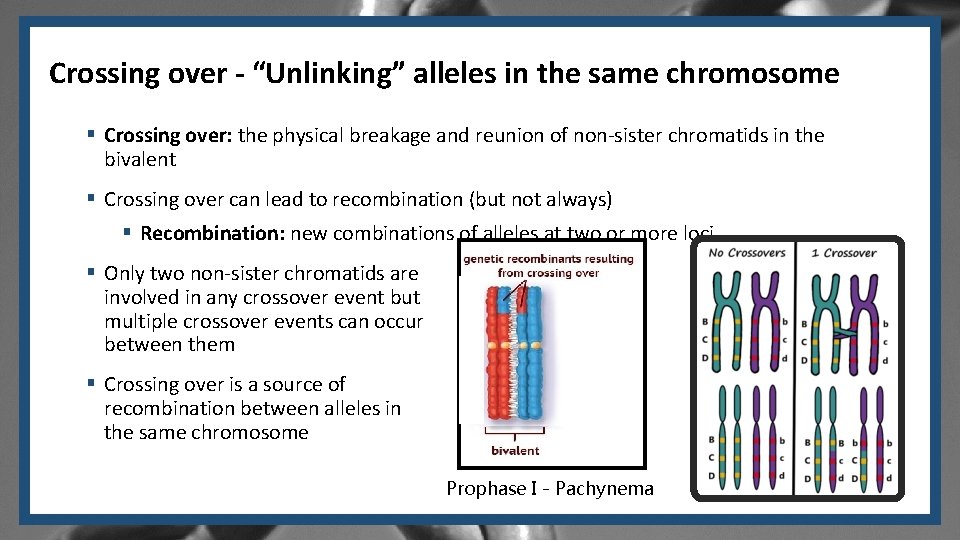
Crossing over - “Unlinking” alleles in the same chromosome § Crossing over: the physical breakage and reunion of non-sister chromatids in the bivalent § Crossing over can lead to recombination (but not always) § Recombination: new combinations of alleles at two or more loci § Only two non-sister chromatids are involved in any crossover event but multiple crossover events can occur between them § Crossing over is a source of recombination between alleles in the same chromosome Prophase I - Pachynema
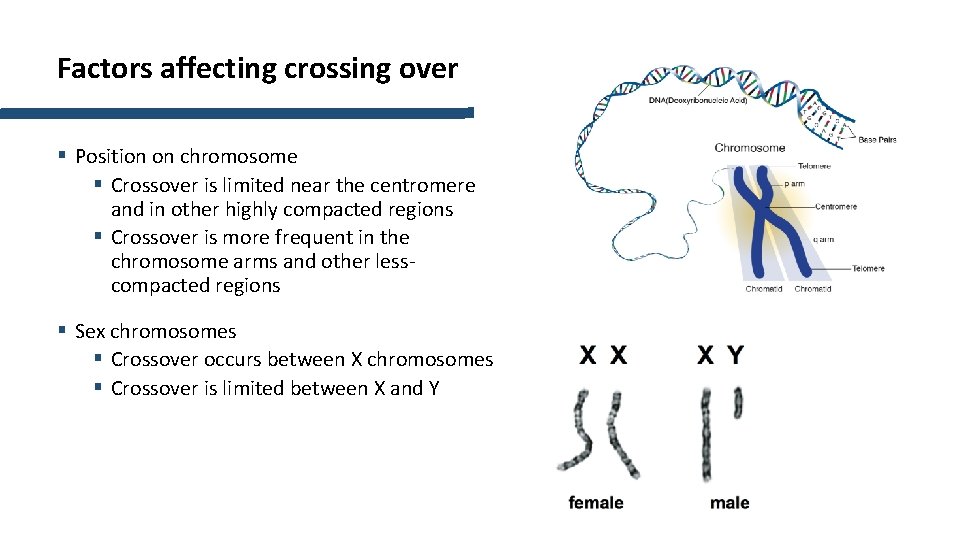
Factors affecting crossing over § Position on chromosome § Crossover is limited near the centromere and in other highly compacted regions § Crossover is more frequent in the chromosome arms and other lesscompacted regions § Sex chromosomes § Crossover occurs between X chromosomes § Crossover is limited between X and Y

Linkage is an exception to independent assortment § Mendel’s Law of Independent Assortment: Alleles of two or more different genes are sorted into gametes independently of one another. § True: if the genes are in different chromosomes § Exception: if genes are linked (same chromosome, few or no crossovers) 2 loci on non-homologous chromosomes 2 loci on same chromosome No crossover Gametes: RY RY ry ry Gametes: RY Ry r. Y ry

Assessing linkage § Recombination frequency Example: AABB x aabb AABB Aa. Bb aabb 25 25 Parental types: AABB & aabb Recombinant types: Aa. Bb RF = 50% § RF = 50% Independent assortment (max. RF possible) loci in different chromosomes or far apart in the same chromosome § RF = 0 complete linkage (no crossover) loci very close in the same chromosome § 0 < RF < 50 partial linkage (crossover is possible) loci in the same chromosome § Considering 2 loci in the same chromosome, the closer they are, the tighter the linkage (low chance of crossover between them) A B § Allelic arrangement at the linked loci: a b § Coupling (dominant/favorable alleles are in the same chromosome) § Repulsion (dominant/favorable alleles are in different chromosomes) A b a B Coupling phase Repulsion phase
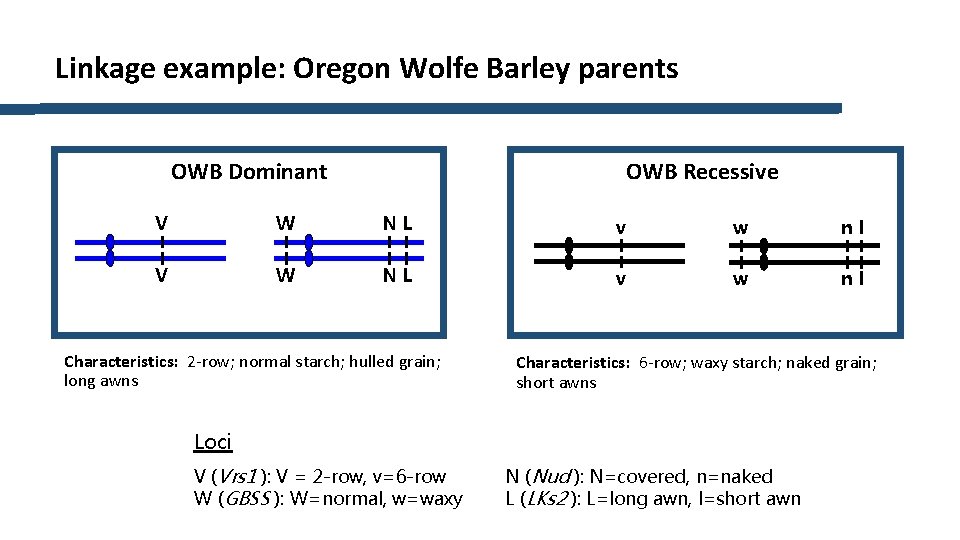
Linkage example: Oregon Wolfe Barley parents OWB Recessive OWB Dominant V W NL v w nl Characteristics: 2 -row; normal starch; hulled grain; long awns Characteristics: 6 -row; waxy starch; naked grain; short awns Loci V (Vrs 1 ): V = 2 -row, v=6 -row W (GBSS ): W=normal, w=waxy N (Nud ): N=covered, n=naked L (LKs 2 ): L=long awn, l=short awn
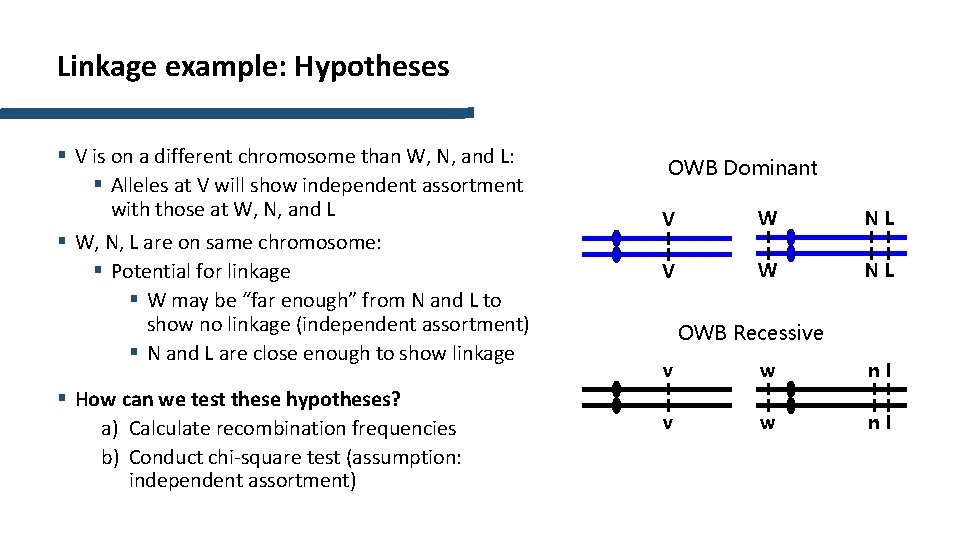
Linkage example: Hypotheses § V is on a different chromosome than W, N, and L: § Alleles at V will show independent assortment with those at W, N, and L § W, N, L are on same chromosome: § Potential for linkage § W may be “far enough” from N and L to show no linkage (independent assortment) § N and L are close enough to show linkage § How can we test these hypotheses? a) Calculate recombination frequencies b) Conduct chi-square test (assumption: independent assortment) OWB Dominant V W NL OWB Recessive v w nl

Linkage example: Dihybrid Ratio for Doubled Haploids (OWB Dominant) VVWW x vvww (OWB Recessive) Characteristic Progeny A Progeny B Progeny C Progeny D Gametes VW Vw v. W vw DH VVWW VVww vv. WW vvww Class Non-recombinant (Parental) Recombinant (Non-parental) Non-recombinant (Parental) Ratio 1 1 Note: The “recombinant“ types in this case (between the V and W loci) are not due to crossovers. They are due to random alignment of paired homologous chromosomes at Metaphase I.

Linkage example: Linkage test between V and W Characteristic Progeny A Progeny B Progeny C Progeny D Gametes VW Vw v. W vw DH VVWW VVww vv. WW vvww Class Non-recombinant Recombinant Non-Recombinant Ratio 1 1 Expected 20. 5 Observed 20 15 25 22 (O-E)2/E 0. 012 1. 476 0. 988 0. 110 = 2. 586; P=0. 460; Fail to reject the null hypothesis of no linkage There is Independent Assortment : Loci are on different chromosomes Random alignment of chromosomes at Metaphase 1 (or loci far apart on same chromosome) 2 [3]

Linkage example: Linkage test between N and L Characteristic Progeny A Progeny B Progeny C Progeny D Gametes NL Nl n. L nl DH NNLL NNll nn. LL nnll Class Non-recombinant Recombinant Non-Recombinant Ratio 1 1 Expected 20. 5 Observed 37 3 9 33 (O-E)2/E 13. 280 14. 939 6. 451 7. 622 = 42. 293; P=<0. 001; Reject the null hypothesis of no linkage Not Independent Assortment = linkage Few crossovers between loci close together on same chromosome 2 [3]
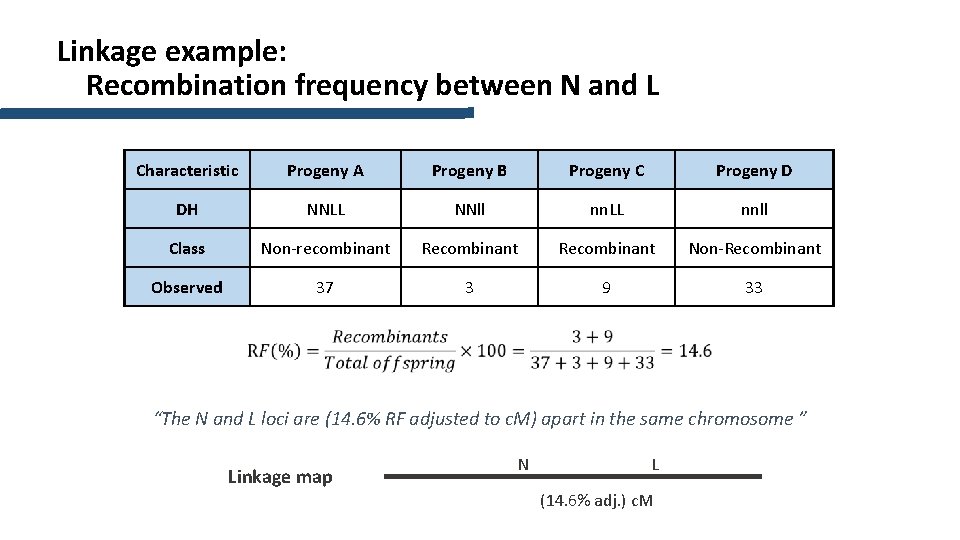
Linkage example: Recombination frequency between N and L Characteristic Progeny A Progeny B Progeny C Progeny D DH NNLL NNll nn. LL nnll Class Non-recombinant Recombinant Non-Recombinant Observed 37 3 9 33 “The N and L loci are (14. 6% RF adjusted to c. M) apart in the same chromosome ” Linkage map N L (14. 6% adj. ) c. M

Why convert recombination frequency to centi. Morgan? § Double crossovers can give “false” parental combinations of alleles recombination frequencies are underestimated (especially at RF>10%) § Analyzing all loci: § Parental types: ABC and abc § RF in gametes= 50% § Analyzing only loci A and C: § “False” parental types: AC and ac § RF in gametes= 0% (underestimated) § Recombination frequencies are not additive 9. 5% 9% 17%

Centi. Morgans (c. M) Formulas Distance (c. M) Haldane =-0. 5*LN(1 -2*r) Distance (c. M) Kosambi=0. 25*LN((1+2*r)/(1 -2*r)) % Recombination c. M Formula 1 1 Haldane 1 Kosambi 11. 2 Haldane 10. 1 Kosambi 25. 5 Haldane 21. 2 Kosambi 45. 8 Haldane 34. 7 Kosambi 80. 5 Haldane 54. 9 Kosambi 310. 7 Haldane 172. 7 Kosambi NA Haldane NA Kosambi 10 20 30 40 c. M (Kosambi) Note: the ratio of c. M to base pairs can vary enormously across the genome no direct conversion from centi. Morgan to base pairs! 49. 9 50

Three-Point Linkage Test § The recombination frequency over the interval W – L (r. WL) is greater than either r. WN or r. NL § Therefore, W and L are the two loci furthest apart and N is likely to lie between them. § N is likely to be closer to L as r. WN > r. NL. r. WN = 0. 39 r. NL = 0. 14 N W r. WL = 0. 48 L

Making Linkage Maps/ Linkage Analysis § Linkage analysis is now an "automated" procedure! § Data used to create dense maps (tens of thousands of loci) § Different software packages offer different algorithms (originally based on chi-square and three-point tests) that can automatically test for linkage and calculate loci order and distance. § Likelihood odds (LOD) score: tests the null-hypothesis that there is no linkage § LOD of 3. 00 p-value ~. 001 § LOD > 3 two loci are linked
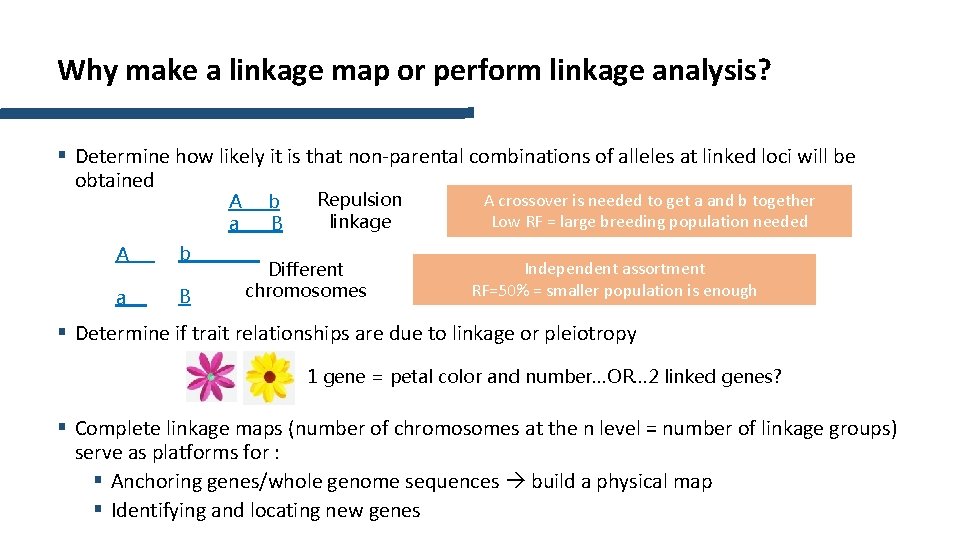
Why make a linkage map or perform linkage analysis? § Determine how likely it is that non-parental combinations of alleles at linked loci will be obtained A a A b a B b B Repulsion linkage Different chromosomes A crossover is needed to get a and b together Low RF = large breeding population needed Independent assortment RF=50% = smaller population is enough § Determine if trait relationships are due to linkage or pleiotropy 1 gene = petal color and number…OR… 2 linked genes? § Complete linkage maps (number of chromosomes at the n level = number of linkage groups) serve as platforms for : § Anchoring genes/whole genome sequences build a physical map § Identifying and locating new genes
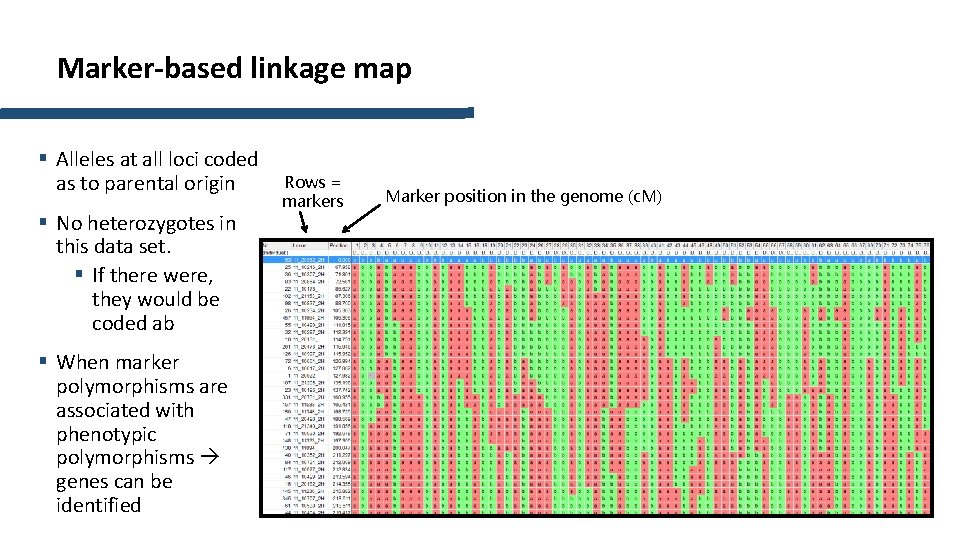
Marker-based linkage map § Alleles at all loci coded as to parental origin § No heterozygotes in this data set. § If there were, they would be coded ab § When marker polymorphisms are associated with phenotypic polymorphisms genes can be identified Rows = markers Marker position in the genome (c. M)
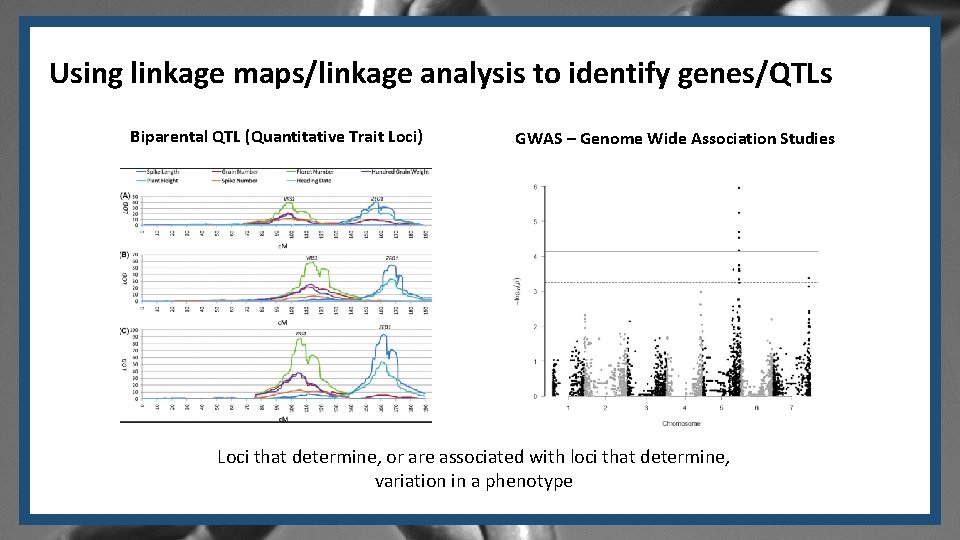
Using linkage maps/linkage analysis to identify genes/QTLs Biparental QTL (Quantitative Trait Loci) GWAS – Genome Wide Association Studies Loci that determine, or are associated with loci that determine, variation in a phenotype
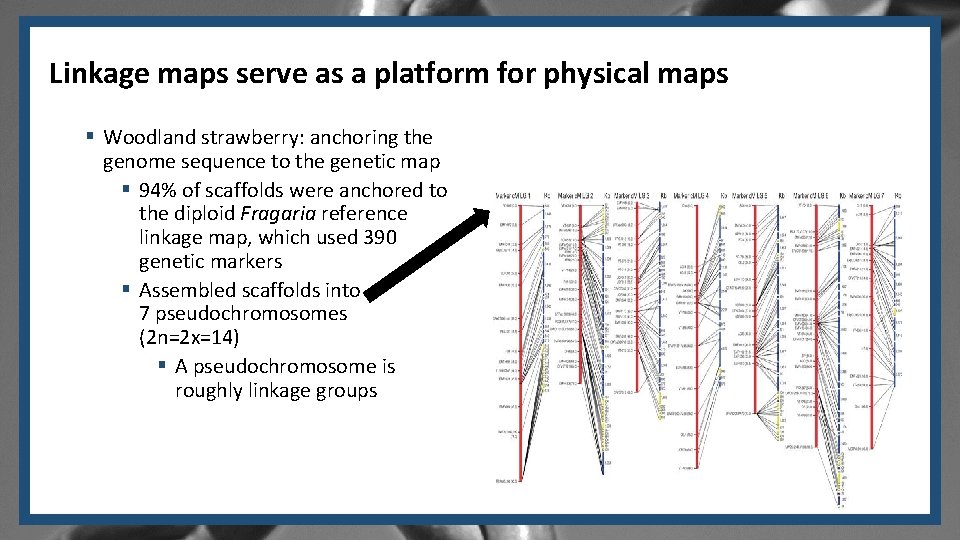
Linkage maps serve as a platform for physical maps § Woodland strawberry: anchoring the genome sequence to the genetic map § 94% of scaffolds were anchored to the diploid Fragaria reference linkage map, which used 390 genetic markers § Assembled scaffolds into 7 pseudochromosomes (2 n=2 x=14) § A pseudochromosome is roughly linkage groups

By now, you should be able to… § Define linkage and give the biological explanation for this phenomena. § Remember: The number of linkage groups in a species = n. § Describe a crossover event and explain how crossing over is a source of recombination. § Remember: Crossovers rarely occur between X and Y chromosomes and in centromeric or other highly compacted regions in the chromosome. § Calculate recombination frequency in a population, given the genotype/phenotype of the parents and offspring. § What can you conclude from this calculated value? Does it suggest that the 2 loci are in the same/different chromosomes, far/close in the same chromosome? § Given the genotype of the parents and the generation studied, test the hypothesis of independent assortment using a chi-square test. § Explain why recombination frequencies are converted to centi. Morgans (c. M). § Remember: There is no direct conversion from c. M to base pairs because recombination patterns are uneven throughout the chromosome length. § Given 2 -locus recombination frequencies for three loci, determine the order and relative distances between the three loci. § Describe how a linkage map can be useful for geneticists and breeders. § How are linkage maps used for finding genes/QTLs?
- Slides: 21