An Introduction to Multiple Sequence Alignments Cdric Notredame













































- Slides: 96

An Introduction to Multiple Sequence Alignments Cédric Notredame

An Introduction to Multiple Sequence Alignments Cédric Notredame

chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. :

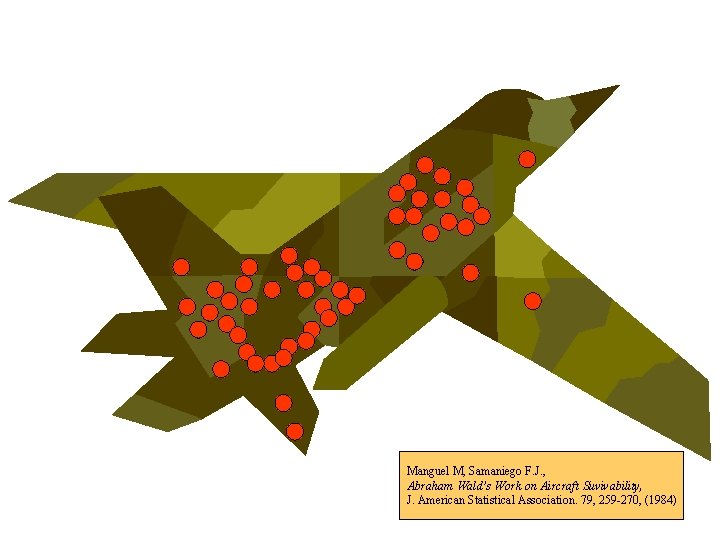
Manguel M, Samaniego F. J. , Abraham Wald’s Work on Aircraft Suvivability, J. American Statistical Association. 79, 259 -270, (1984)

Our Scope How Can I Use My Alignment? How Does The Computer Align The Sequences? How Can I Assemble a Mult. Aln? What are the Difficulties?

Outline -Why Do We Need Multiple Sequence Alignment ? -The progressive Alignment Algorithm -A possible Strategy… -Potential Difficulties

Pre-requisite -How Do Sequences Evolve? -How can We COMPARE Sequences ? -How can We ALIGN Sequences ?

Why Do We Need Multiple Sequence Alignment ?

Sometimes Two Sequences Are Not Enough… The man with TWO watches NEVER knows the time

What is A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Structural Criteria: Residues are arranged so that those playing a similar role end up in the same column. Evolution Criteria: Residues are arranged so that those having the same ancestor end up in the same column.

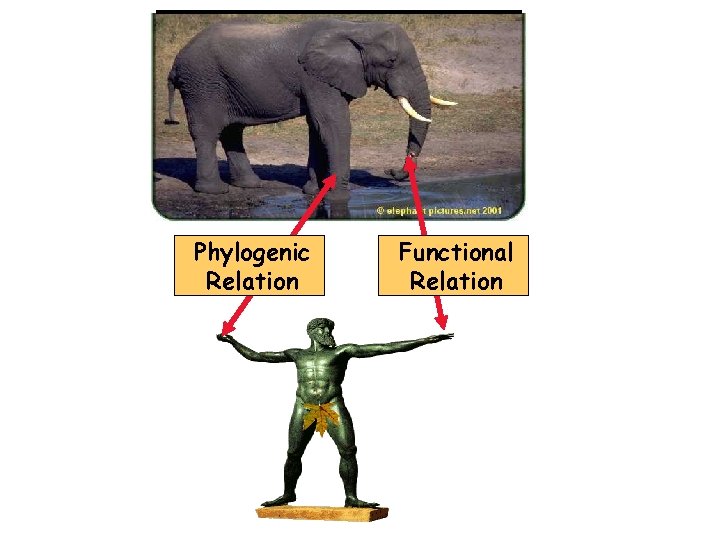
Phylogenic Relation Functional Relation


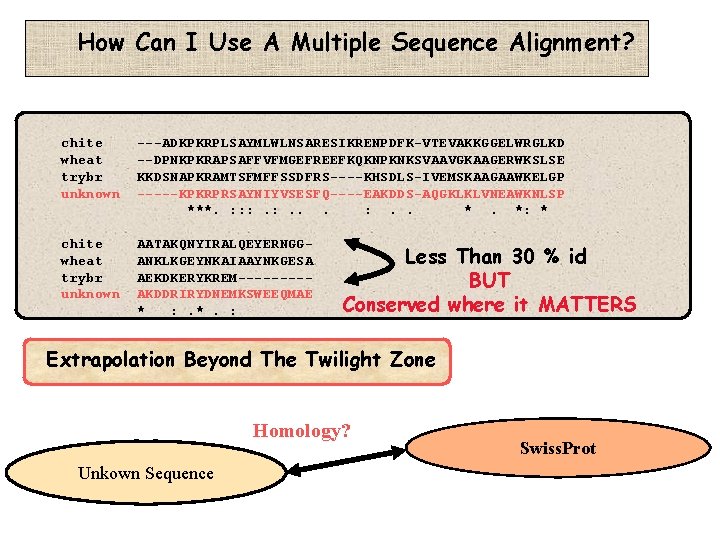
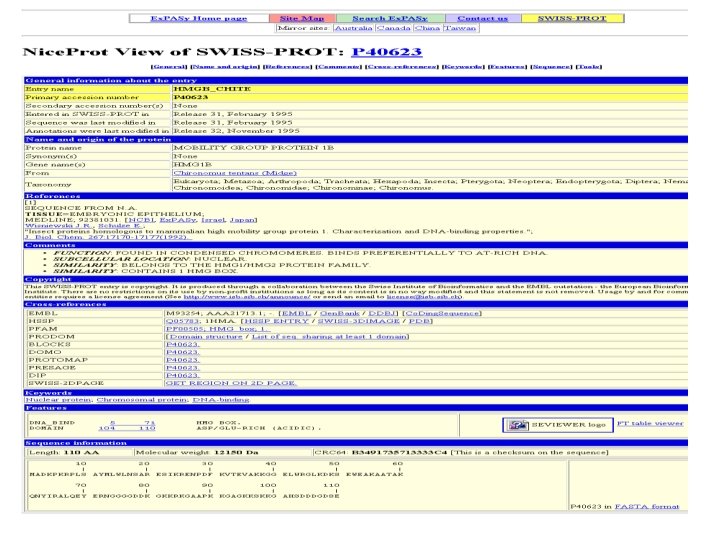
How Can I Use A Multiple Sequence Alignment? chite wheat trybr unknown ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr unknown AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Less Than 30 % id BUT Conserved where it MATTERS Extrapolation Beyond The Twilight Zone Homology? Unkown Sequence Swiss. Prot


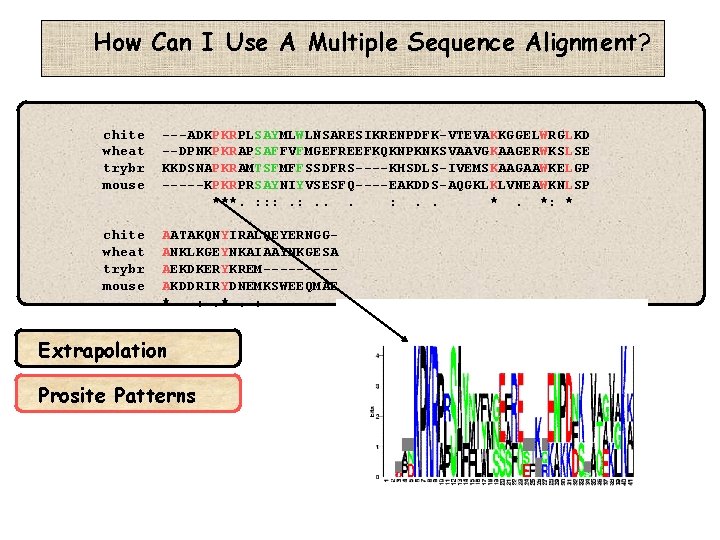
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Prosite Patterns

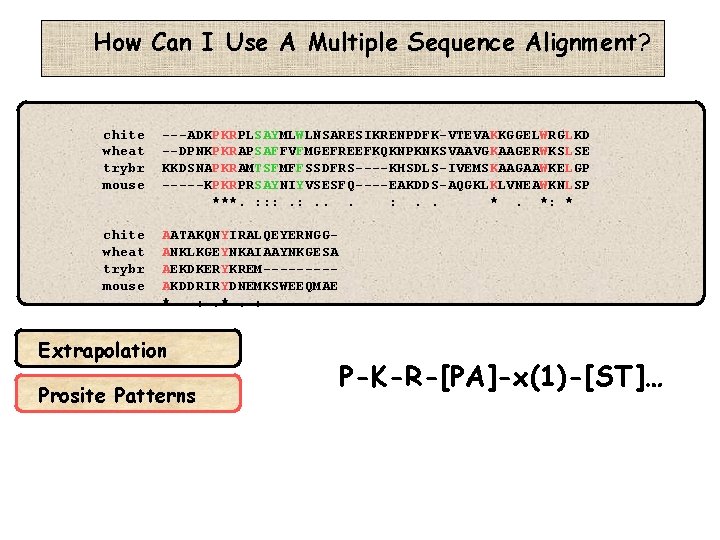
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Prosite Patterns P-K-R-[PA]-x(1)-[ST]…

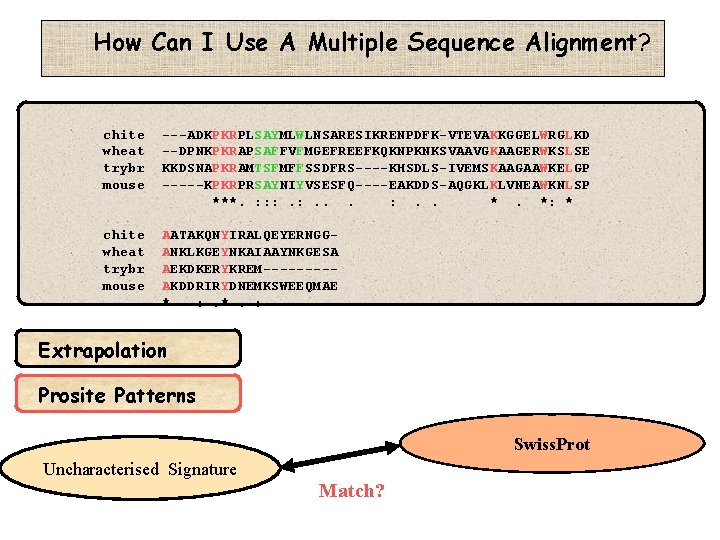
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Prosite Patterns Swiss. Prot Uncharacterised Signature Match?

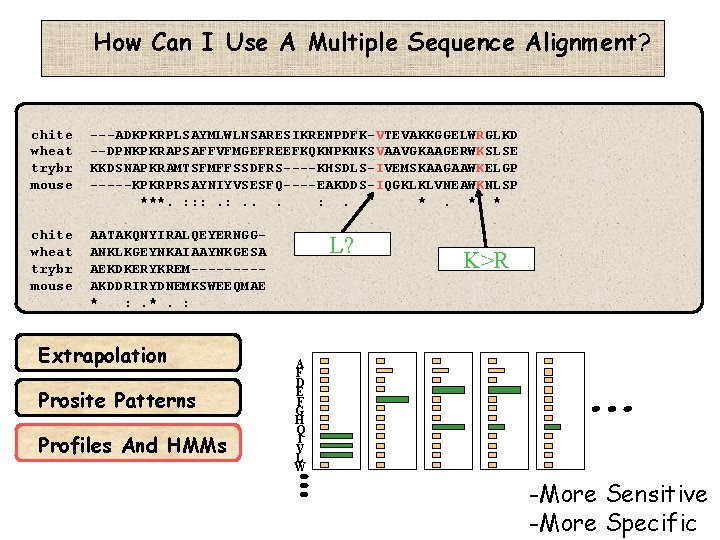
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-IQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Prosite Patterns Profiles And HMMs L? K>R A F D E F G H Q I V L W -More Sensitive -More Specific

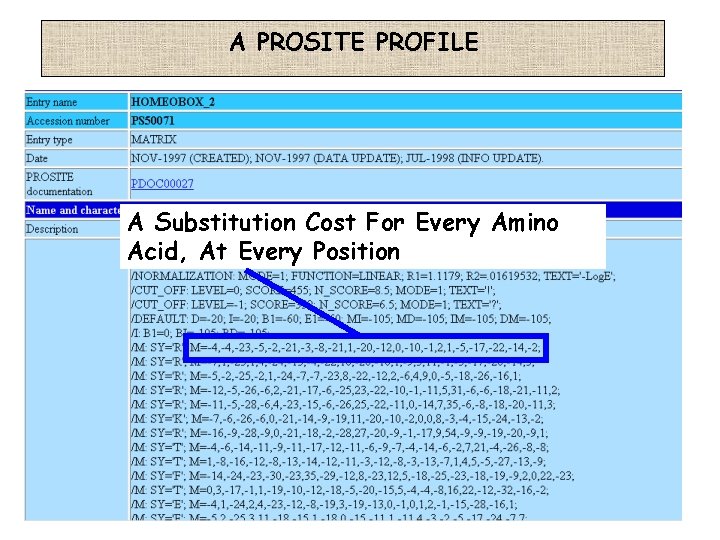
A PROSITE PROFILE A Substitution Cost For Every Amino Acid, At Every Position

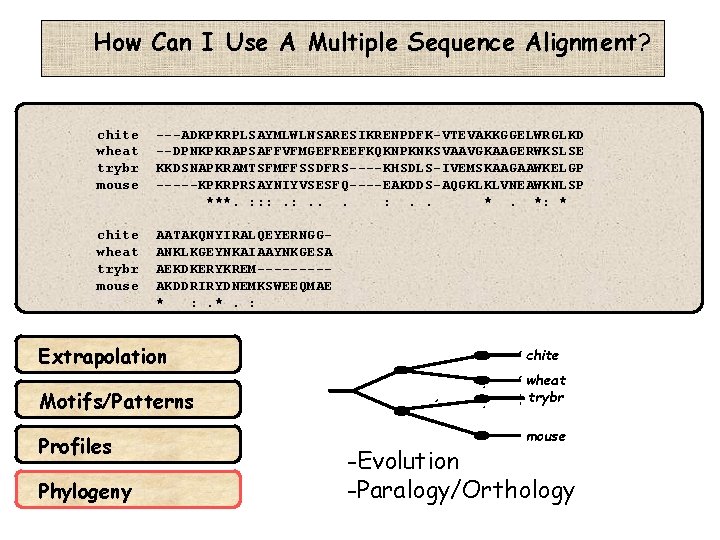
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Motifs/Patterns Profiles Phylogeny chite wheat trybr mouse -Evolution -Paralogy/Orthology

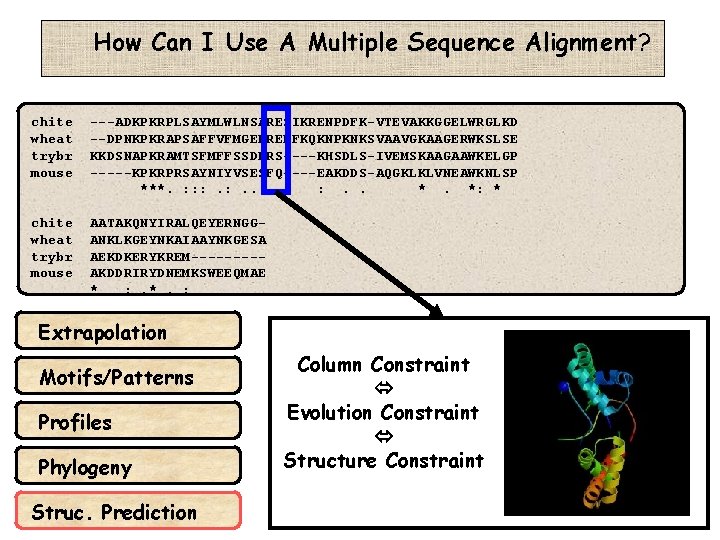
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Motifs/Patterns Profiles Phylogeny Struc. Prediction Column Constraint Evolution Constraint Structure Constraint

How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Extrapolation Motifs/Patterns Profiles Phylogeny Struc. Prediction Psi. Pred OR Ph. D For secondary Structure Prediction: 75% Accurate. Threading: is improving but is not yet as good.

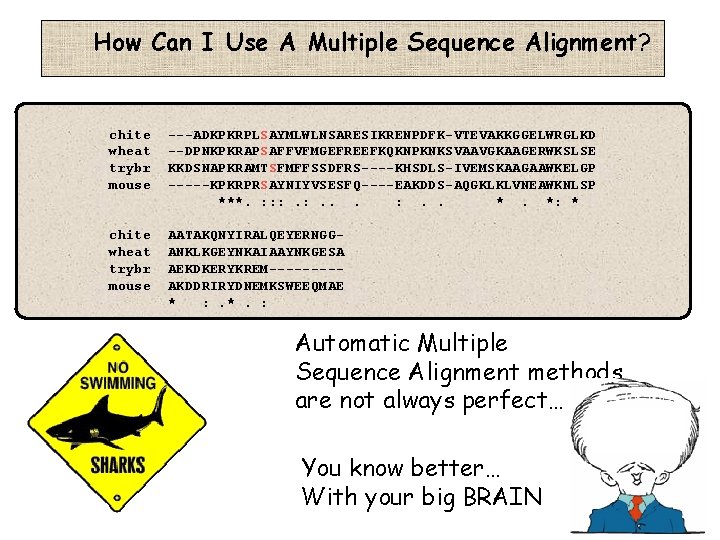
How Can I Use A Multiple Sequence Alignment? chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : Automatic Multiple Sequence Alignment methods are not always perfect… You know better… With your big BRAIN


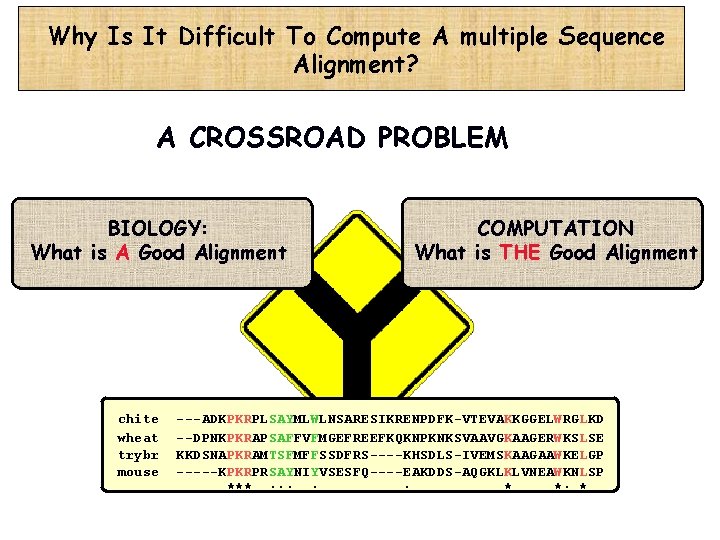
Why Is It Difficult To Compute A multiple Sequence Alignment? A CROSSROAD PROBLEM BIOLOGY: What is A Good Alignment chite wheat trybr mouse COMPUTATION What is THE Good Alignment ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: *

The Biological Problem. Same as Pair. Wise Alignment Problem We do NOT know how Sequences Evolve. We do NOT understand the Relation Between Structures and Sequences. We would NOT recognize the Correct Alignment if we had it IN FRONT of our eyes…

The Biological Problem. The Charlie Chaplin Paradox

The Biological Problem. How to Evaluate an Alignment -A nice set of Sequences -Substitution Matrix (Blosum) -Gap Penalties. -An Evaluation Function A A A C C A A A C Sums of Pairs: Cost=6 C Over-estimation of the Substitutions Easy to compute

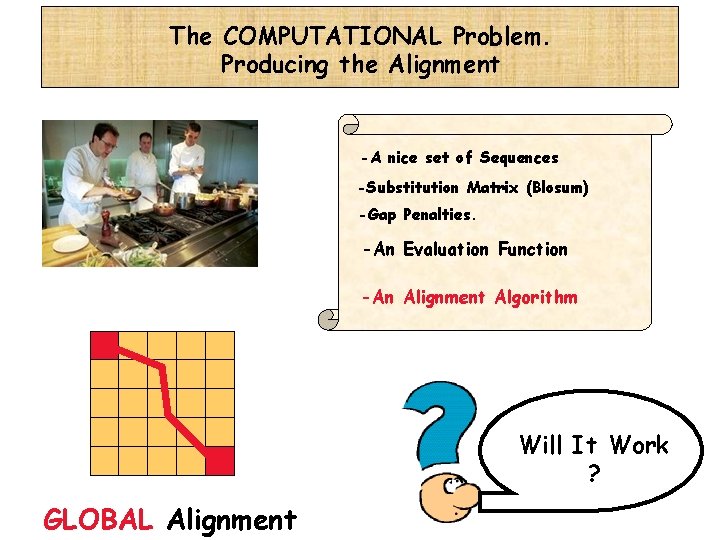
The COMPUTATIONAL Problem. Producing the Alignment -A nice set of Sequences -Substitution Matrix (Blosum) -Gap Penalties. -An Evaluation Function -An Alignment Algorithm Will It Work ? GLOBAL Alignment

HOW CAN I ALIGN MANY SEQUENCES 2 Globins =>1 Min

HOW CAN I ALIGN MANY SEQUENCES 3 Globins =>2 hours

HOW CAN I ALIGN MANY SEQUENCES 4 Globins => 10 days

HOW CAN I ALIGN MANY SEQUENCES 5 Globins => 3 years

HOW CAN I ALIGN MANY SEQUENCES ! DHEA Loaded 6 Globins =>300 years

HOW CAN I ALIGN MANY SEQUENCES 7 Globins =>30. 000 years Solidified Fossil, Old stuff

HOW CAN I ALIGN MANY SEQUENCES 8 Globins =>3 Million years

The Progressive Multiple Alignment Algorithm (Clustal W)


Making An Alignment Any Exact Method would be TOO SLOW We will use a Heuristic Algorithm. Progressive Alignment Algorithm is the most Popular -Clustal. W -Greedy Heuristic (No Guarranty). -Fast

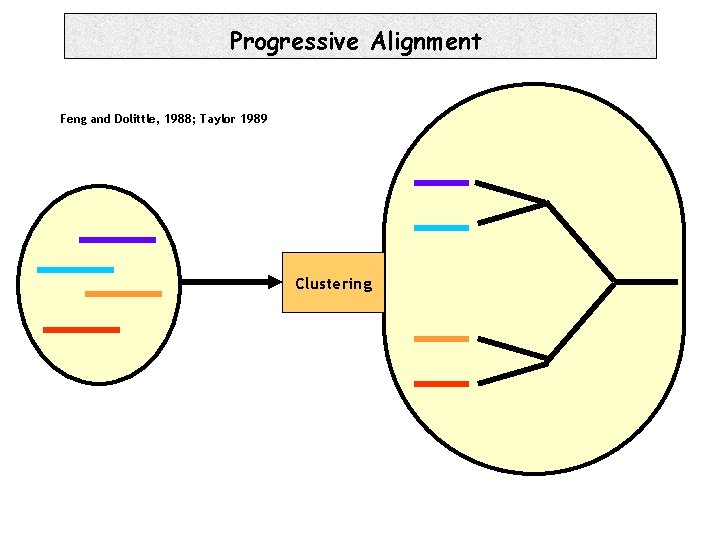
Progressive Alignment Feng and Dolittle, 1988; Taylor 1989 Clustering

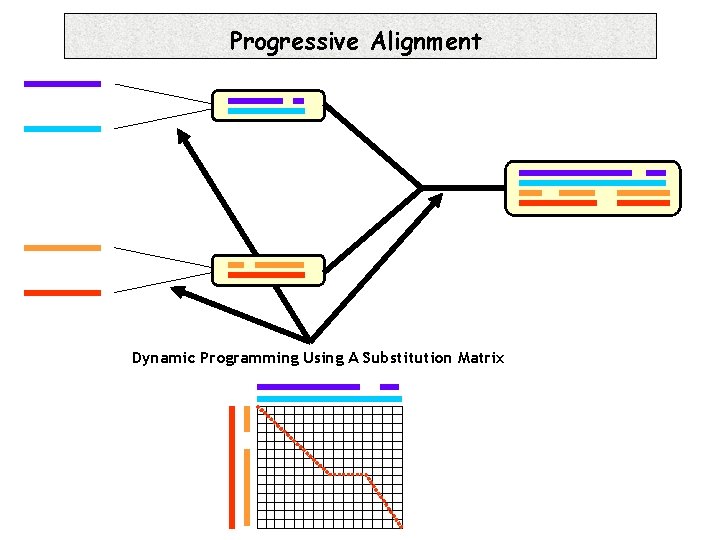
Progressive Alignment Dynamic Programming Using A Substitution Matrix

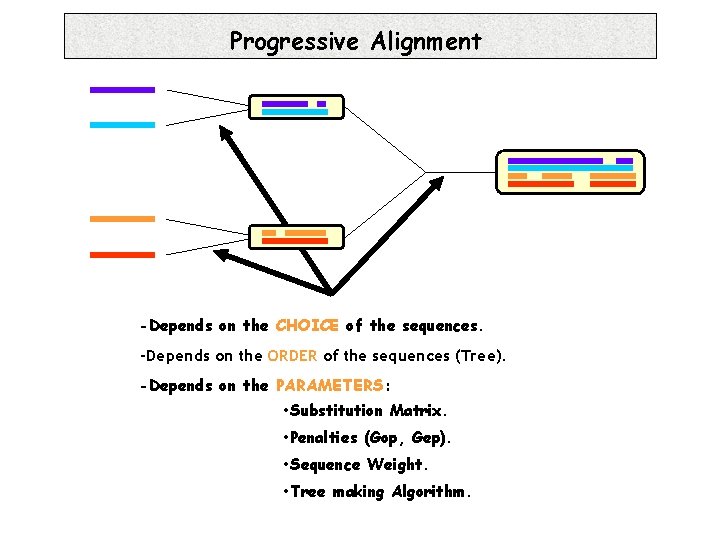
Progressive Alignment -Depends on the CHOICE of the sequences. -Depends on the ORDER of the sequences (Tree). -Depends on the PARAMETERS: • Substitution Matrix. • Penalties (Gop, Gep). • Sequence Weight. • Tree making Algorithm.

Progressive Alignment When Does It Works Well When Phylogeny is Dense No outlayer Sequence. Image: River Crossing

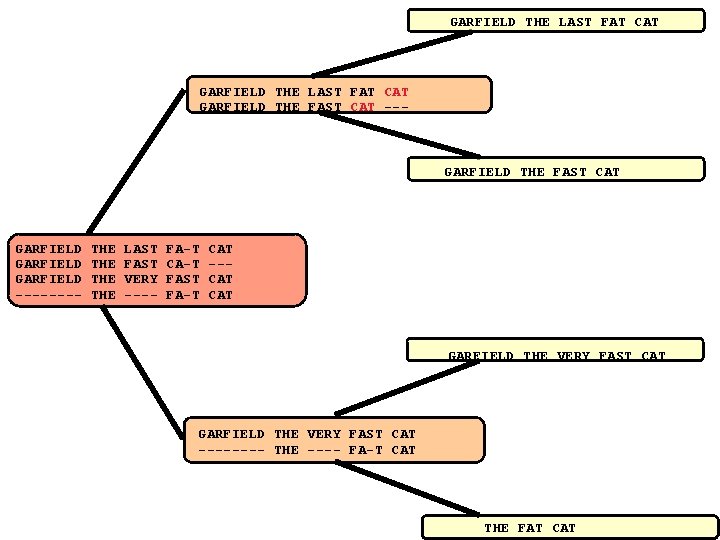
Progressive Alignment When Doesn’t It Work CLUSTALW (Score=20, Gop=-1, Gep=0, M=1) Seq. A Seq. B Seq. C Seq. D GARFIELD ---- THE THE LAST FAST VERY ---- FA-T CA-T FAST FA-T CAT --CAT LAST FAST VERY ---- FA-T ---FAST FA-T CAT CAT CORRECT (Score=24) Seq. A Seq. B Seq. C Seq. D GARFIELD ---- THE THE

GARFIELD THE LAST FAT CAT GARFIELD THE FAST CAT --- GARFIELD THE FAST CAT GARFIELD ---- THE THE LAST FAST VERY ---- FA-T CA-T FAST FA-T CAT --CAT GARFIELD THE VERY FAST CAT ---- THE ---- FA-T CAT THE FAT CAT

Building the Right Multiple Sequence Alignment.

Recognizing The Right Sequences When you Meet Them…

Gathering Sequences: BLAST

Common Mistake: Sequences Too Closely Related PRVA_MACFU PRVA_HUMAN PRVA_GERSP PRVA_MOUSE PRVA_RAT PRVA_RABIT SMTDLLNAEDIKKAVGAFSAIDSFDHKKFFQMVGLKKKSADDVKKVFHILDKDKSGFIEE SMTDLLNAEDIKKAVGAFSATDSFDHKKFFQMVGLKKKSADDVKKVFHMLDKDKSGFIEE SMTDLLSAEDIKKAIGAFAAADSFDHKKFFQMVGLKKKTPDDVKKVFHILDKDKSGFIEE SMTDVLSAEDIKKAIGAFAAADSFDHKKFFQMVGLKKKNPDEVKKVFHILDKDKSGFIEE SMTDLLSAEDIKKAIGAFTAADSFDHKKFFQMVGLKKKSADDVKKVFHILDKDKSGFIEE AMTELLNAEDIKKAIGAFAAAESFDHKKFFQMVGLKKKSTEDVKKVFHILDKDKSGFIEE : **: : *. *******: * : ********. . : : *********** PRVA_MACFU PRVA_HUMAN PRVA_GERSP PRVA_MOUSE PRVA_RAT PRVA_RABIT DELGFILKGFSPDARDLSAKETKTLMAAGDKDGDGKIGVDEFSTLVAES DELGFILKGFSPDARDLSAKETKMLMAAGDKDGDGKIGVDEFSTLVAES DELGFILKGFSSDARDLSAKETKTLLAAGDKDGDGKIGVEEFSTLVSES DELGSILKGFSSDARDLSAKETKTLLAAGDKDGDGKIGVEEFSTLVAES DELGSILKGFSSDARDLSAKETKTLMAAGDKDGDGKIGVEEFSTLVAES EELGFILKGFSPDARDLSVKETKTLMAAGDKDGDGKIGADEFSTLVSES : ****** *: ******: ** -IDENTICAL SEQUENCES BRING NO INFORMATION FOR THE MULTIPLE SEQUENCE ALIGNMENT -MULTIPLE SEQUENCE ALIGNMENTS THRIVE ON DIVERSITY…

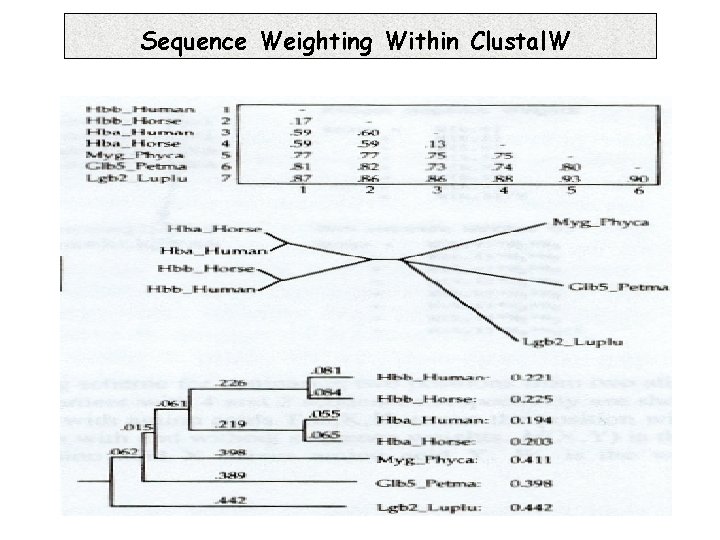
Sequence Weighting Within Clustal. W

Selecting Diverse Sequences (Opus II)

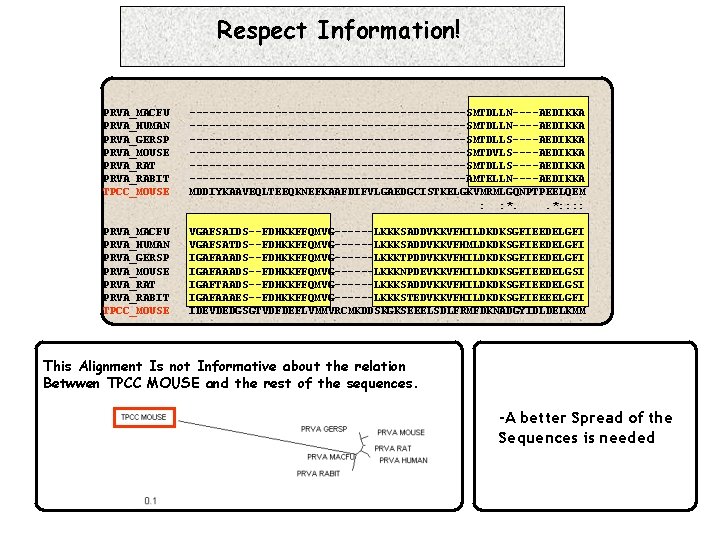
Respect Information! PRVA_MACFU PRVA_HUMAN PRVA_GERSP PRVA_MOUSE PRVA_RAT PRVA_RABIT TPCC_MOUSE ------------------------------------------SMTDLLN----AEDIKKA ---------------------SMTDLLS----AEDIKKA ---------------------SMTDVLS----AEDIKKA ---------------------SMTDLLS----AEDIKKA ---------------------AMTELLN----AEDIKKA MDDIYKAAVEQLTEEQKNEFKAAFDIFVLGAEDGCISTKELGKVMRMLGQNPTPEELQEM : : *. . *: : PRVA_MACFU PRVA_HUMAN PRVA_GERSP PRVA_MOUSE PRVA_RAT PRVA_RABIT TPCC_MOUSE VGAFSAIDS--FDHKKFFQMVG------LKKKSADDVKKVFHILDKDKSGFIEEDELGFI VGAFSATDS--FDHKKFFQMVG------LKKKSADDVKKVFHMLDKDKSGFIEEDELGFI IGAFAAADS--FDHKKFFQMVG------LKKKTPDDVKKVFHILDKDKSGFIEEDELGFI IGAFAAADS--FDHKKFFQMVG------LKKKNPDEVKKVFHILDKDKSGFIEEDELGSI IGAFTAADS--FDHKKFFQMVG------LKKKSADDVKKVFHILDKDKSGFIEEDELGSI IGAFAAAES--FDHKKFFQMVG------LKKKSTEDVKKVFHILDKDKSGFIEEEELGFI IDEVDEDGSGTVDFDEFLVMMVRCMKDDSKGKSEEELSDLFRMFDKNADGYIDLDELKMM This Alignment Is not Informative about the relation Betwwen TPCC MOUSE and the rest of the sequences. -A better Spread of the Sequences is needed

Selecting Diverse Sequences (Opus II)

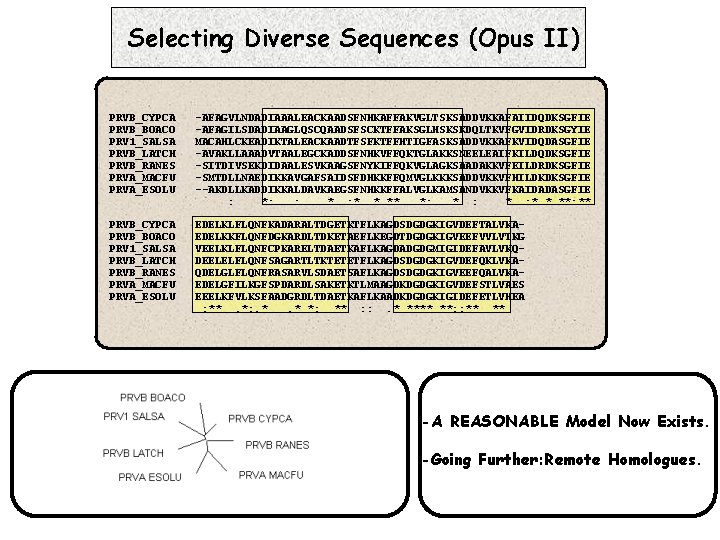
Selecting Diverse Sequences (Opus II) PRVB_CYPCA PRVB_BOACO PRV 1_SALSA PRVB_LATCH PRVB_RANES PRVA_MACFU PRVA_ESOLU -AFAGVLNDADIAAALEACKAADSFNHKAFFAKVGLTSKSADDVKKAFAIIDQDKSGFIE -AFAGILSDADIAAGLQSCQAADSFSCKTFFAKSGLHSKSKDQLTKVFGVIDRDKSGYIE MACAHLCKEADIKTALEACKAADTFSFKTFFHTIGFASKSADDVKKAFKVIDQDASGFIE -AVAKLLAAADVTAALEGCKADDSFNHKVFFQKTGLAKKSNEELEAIFKILDQDKSGFIE -SITDIVSEKDIDAALESVKAAGSFNYKIFFQKVGLAGKSAADAKKVFEILDRDKSGFIE -SMTDLLNAEDIKKAVGAFSAIDSFDHKKFFQMVGLKKKSADDVKKVFHILDKDKSGFIE --AKDLLKADDIKKALDAVKAEGSFNHKKFFALVGLKAMSANDVKKVFKAIDADASGFIE : *: . . *. : *. * ** *: * : * * **: ** PRVB_CYPCA PRVB_BOACO PRV 1_SALSA PRVB_LATCH PRVB_RANES PRVA_MACFU PRVA_ESOLU EDELKLFLQNFKADARALTDGETKTFLKAGDSDGDGKIGVDEFTALVKAEDELKKFLQNFDGKARDLTDKETAEFLKEGDTDGDGKIGVEEFVVLVTKG VEELKLFLQNFCPKARELTDAETKAFLKAGDADGDGMIGIDEFAVLVKQDEELELFLQNFSAGARTLTKTETETFLKAGDSDGDGKIGVDEFQKLVKAQDELGLFLQNFRASARVLSDAETSAFLKAGDSDGDGKIGVEEFQALVKAEDELGFILKGFSPDARDLSAKETKTLMAAGDKDGDGKIGVDEFSTLVAES EEELKFVLKSFAADGRDLTDAETKAFLKAADKDGDGKIGIDEFETLVHEA : **. *: . *. * *: ** : : . * **** **: : ** ** -A REASONABLE Model Now Exists. -Going Further: Remote Homologues.

Aligning Remote Homologues PRVA_MACFU PRVA_ESOLU PRVB_CYPCA PRVB_BOACO PRV 1_SALSA PRVB_LATCH PRVB_RANES TPCS_RABIT TPCS_PIG TPCC_MOUSE ---------------------SMTDLLNA----EDIKKA ----------------------AKDLLKA----DDIKKA ---------------------AFAGVLND----ADIAAA ---------------------AFAGILSD----ADIAAG ---------------------MACAHLCKE----ADIKTA ---------------------AVAKLLAA----ADVTAA ---------------------SITDIVSE----KDIDAA -TDQQAEARSYLSEEMIAEFKAAFDMFDADGG-GDISVKELGTVMRMLGQTPTKEELDAI MDDIYKAAVEQLTEEQKNEFKAAFDIFVLGAEDGCISTKELGKVMRMLGQNPTPEELQEM : : : PRVA_MACFU PRVA_ESOLU PRVB_CYPCA PRVB_BOACO PRV 1_SALSA PRVB_LATCH PRVB_RANES TPCS_RABIT TPCS_PIG TPCC_MOUSE VGAFSAIDS--FDHKKFFQMVG------LKKKSADDVKKVFHILDKDKSGFIEEDELGFI LDAVKAEGS--FNHKKFFALVG------LKAMSANDVKKVFKAIDADASGFIEEEELKFV LEACKAADS--FNHKAFFAKVG------LTSKSADDVKKAFAIIDQDKSGFIEEDELKLF LQSCQAADS--FSCKTFFAKSG------LHSKSKDQLTKVFGVIDRDKSGYIEEDELKKF LEACKAADT--FSFKTFFHTIG------FASKSADDVKKAFKVIDQDASGFIEVEELKLF LEGCKADDS--FNHKVFFQKTG------LAKKSNEELEAIFKILDQDKSGFIEDEELELF LESVKAAGS--FNYKIFFQKVG------LAGKSAADAKKVFEILDRDKSGFIEQDELGLF IEEVDEDGSGTIDFEEFLVMMVRQMKEDAKGKSEEELAECFRIFDRNADGYIDAEELAEI IEEVDEDGSGTIDFEEFLVMMVRQMKEDAKGKSEEELAECFRIFDRNMDGYIDAEELAEI IDEVDEDGSGTVDFDEFLVMMVRCMKDDSKGKSEEELSDLFRMFDKNADGYIDLDELKMM : . . . *: * : * : . *: *: : **. PRVA_MACFU PRVA_ESOLU PRVB_CYPCA PRVB_BOACO PRV 1_SALSA PRVB_LATCH PRVB_RANES TPCS_RABIT TPCS_PIG TPCC_MOUSE LKGFSPDARDLSAKETKTLMAAGDKDGDGKIGVDEFSTLVAESLKSFAADGRDLTDAETKAFLKAADKDGDGKIGIDEFETLVHEALQNFKADARALTDGETKTFLKAGDSDGDGKIGVDEFTALVKA-LQNFDGKARDLTDKETAEFLKEGDTDGDGKIGVEEFVVLVTKGLQNFCPKARELTDAETKAFLKAGDADGDGMIGIDEFAVLVKQ-LQNFSAGARTLTKTETETFLKAGDSDGDGKIGVDEFQKLVKA-LQNFRASARVLSDAETSAFLKAGDSDGDGKIGVEEFQALVKA-FR---ASGEHVTDEEIESLMKDGDKNNDGRIDFDEFLKMMEGVQ FR---ASGEHVTDEEIESIMKDGDKNNDGRIDFDEFLKMMEGVQ LQ---ATGETITEDDIEELMKDGDKNNDGRIDYDEFLEFMKGVE : : . . : : : . ** *. : ** : :

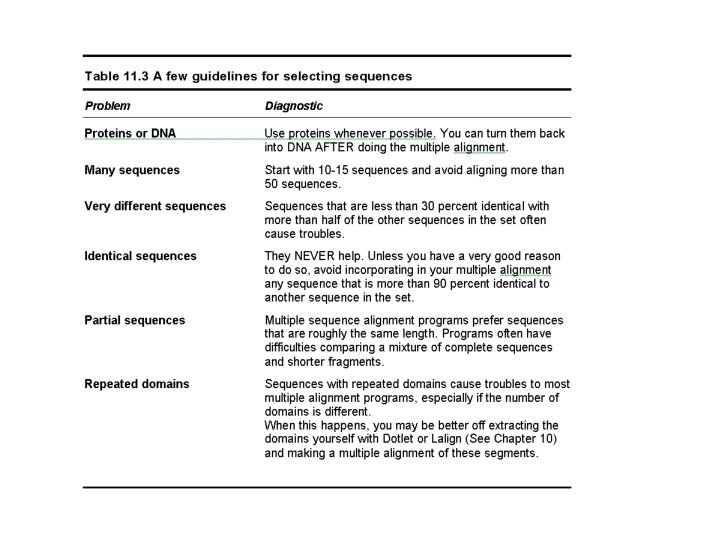
Some Guidelines …

Do Not Use Two Many Sequences…

Reading Your Alignment


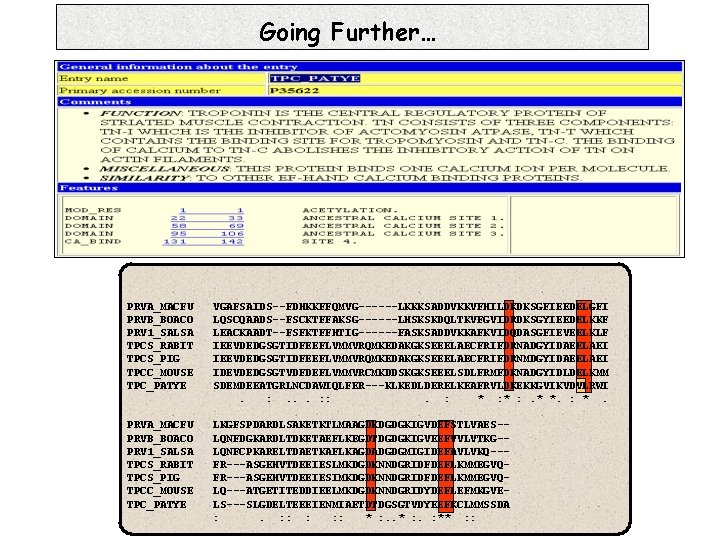
Going Further… PRVA_MACFU PRVB_BOACO PRV 1_SALSA TPCS_RABIT TPCS_PIG TPCC_MOUSE TPC_PATYE VGAFSAIDS--FDHKKFFQMVG------LKKKSADDVKKVFHILDKDKSGFIEEDELGFI LQSCQAADS--FSCKTFFAKSG------LHSKSKDQLTKVFGVIDRDKSGYIEEDELKKF LEACKAADT--FSFKTFFHTIG------FASKSADDVKKAFKVIDQDASGFIEVEELKLF IEEVDEDGSGTIDFEEFLVMMVRQMKEDAKGKSEEELAECFRIFDRNADGYIDAEELAEI IEEVDEDGSGTIDFEEFLVMMVRQMKEDAKGKSEEELAECFRIFDRNMDGYIDAEELAEI IDEVDEDGSGTVDFDEFLVMMVRCMKDDSKGKSEEELSDLFRMFDKNADGYIDLDELKMM SDEMDEEATGRLNCDAWIQLFER---KLKEDLDERELKEAFRVLDKEKKGVIKVDVLRWI. : . . . : : . : * : . * *. : *. PRVA_MACFU PRVB_BOACO PRV 1_SALSA TPCS_RABIT TPCS_PIG TPCC_MOUSE TPC_PATYE LKGFSPDARDLSAKETKTLMAAGDKDGDGKIGVDEFSTLVAES-LQNFDGKARDLTDKETAEFLKEGDTDGDGKIGVEEFVVLVTKG-LQNFCPKARELTDAETKAFLKAGDADGDGMIGIDEFAVLVKQ--FR---ASGEHVTDEEIESLMKDGDKNNDGRIDFDEFLKMMEGVQFR---ASGEHVTDEEIESIMKDGDKNNDGRIDFDEFLKMMEGVQLQ---ATGETITEDDIEELMKDGDKNNDGRIDYDEFLEFMKGVELS---SLGDELTEEEIENMIAETDTDGSGTVDYEEFKCLMMSSDA : . : : : * : . : ** : :

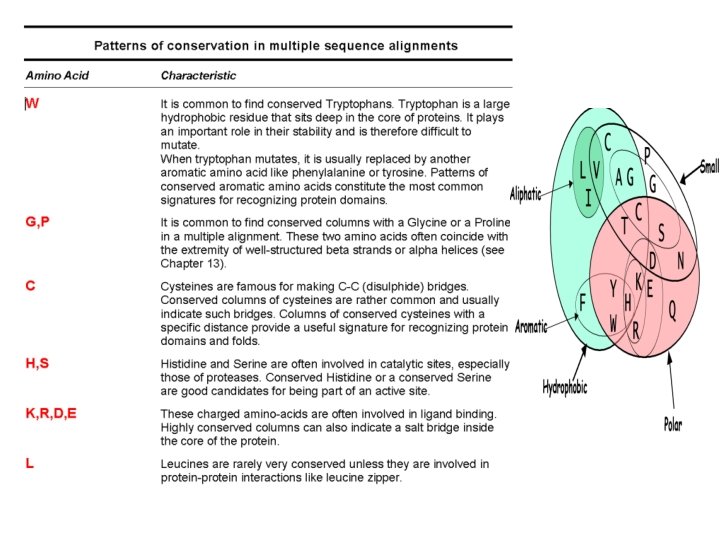
WHAT MAKES A GOOD ALIGNMENT… -THE MORE DIVERGEANT THE SEQUENCES, THE BETTER -THE FEWER INDELS, THE BETTER -NICE UNGAPPED BLOCKS SEPARATED WITH INDELS -DIFFERENT CLASSES OF RESIDUES WITHIN A BLOCK: • Completely Conserved • Conserved For Size and Hydropathy • Conserved For Size or Hydropathy -THE ULTIMATE EVALUATION IS A MATTER OF PERSONNAL JUDGEMENT AND KNOWLEDGE.


Potential Difficulties

DO NOT OVERTUNE!!! chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. : DO NOT PLAY WITH PARAMETERS IF YOU KNOW THE ALIGNMENT YOU WANT: MAKE IT YOURSELF! chite wheat trybr mouse ---ADKPKRPL-SAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAP-SAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPR-SAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : *: . . *. *: * chite wheat trybr mouse AATAKQNYIRALQEYERNGGANKLKGEYNKAIAAYNKGESA AEKDKERYKREM----AKDDRIRYDNEMKSWEEQMAE * : . *. :

TUNING or NOT TUNING!!! -PARAMETERS TO TUNE USUALLY INCLUDE: • GOP/ GEP • MATRIX • SENSITIVITY Vs SPEED Substitution Matrices (Etzold and al. 1993) GOP Gonnet Blosum 50 Pam 250 61. 7 % 59. 2 % GEP -MOST METHODS ARE TUNED FOR WORKING WELL ON AVERAGE -PARAMETERS BEHAVIOUR DO NOT NECESSARILY FOLLOW THEORY (i. e. Substitution Matrices). -A GOOD ALIGNMENT IS USUALLY ROBUST(i. e. Changes little). -TUNE IF YOU WANT TO CONVINCE YOURSELF.


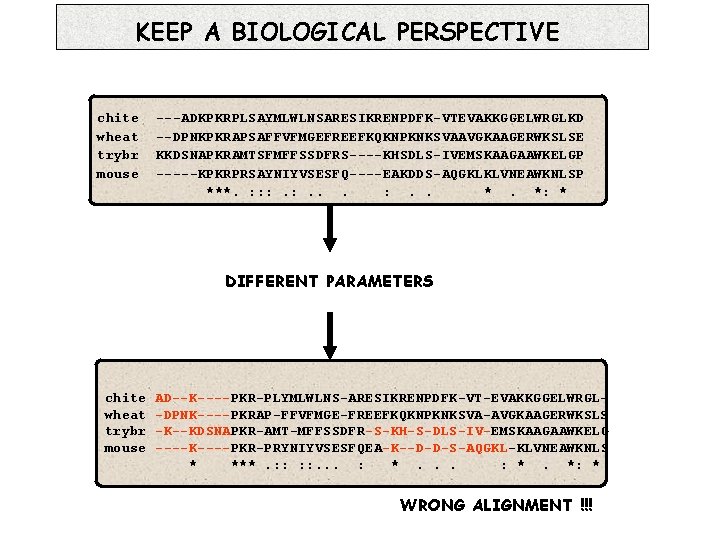
KEEP A BIOLOGICAL PERSPECTIVE chite wheat trybr mouse ---ADKPKRPLSAYMLWLNSARESIKRENPDFK-VTEVAKKGGELWRGLKD --DPNKPKRAPSAFFVFMGEFREEFKQKNPKNKSVAAVGKAAGERWKSLSE KKDSNAPKRAMTSFMFFSSDFRS----KHSDLS-IVEMSKAAGAAWKELGP -----KPKRPRSAYNIYVSESFQ----EAKDDS-AQGKLKLVNEAWKNLSP ***. : : : . . *. *: * DIFFERENT PARAMETERS chite wheat trybr mouse AD--K----PKR-PLYMLWLNS-ARESIKRENPDFK-VT-EVAKKGGELWRGL-DPNK----PKRAP-FFVFMGE-FREEFKQKNPKNKSVA-AVGKAAGERWKSLS -K--KDSNAPKR-AMT-MFFSSDFR-S-KH-S-DLS-IV-EMSKAAGAAWKELG ----K----PKR-PRYNIYVSESFQEA-K--D-D-S-AQGKL-KLVNEAWKNLS * ***. : : . . . : *. *: * WRONG ALIGNMENT !!!

REPEATS THERE IS A PROBLEM WHEN TWO SEQUENCES DO NOT CONTAIN THE SAME NUMBER OF REPEATS IT IS THEN BETTER TO MANUALLY EXTRACT THE REPEATS AND TO ALIGN THEM. INDIVIDUAL REPEATS CAN BE RECOGNIZED USING DOTTER


Naming Your Sequences The Right Way

What Are The Available Methods ? ? ?

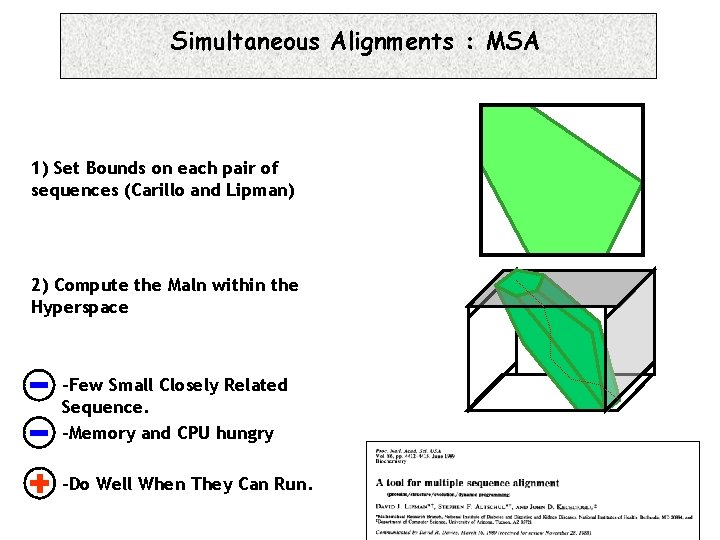
Simultaneous Alignments : MSA 1) Set Bounds on each pair of sequences (Carillo and Lipman) 2) Compute the Maln within the Hyperspace -Few Small Closely Related Sequence. -Memory and CPU hungry -Do Well When They Can Run.

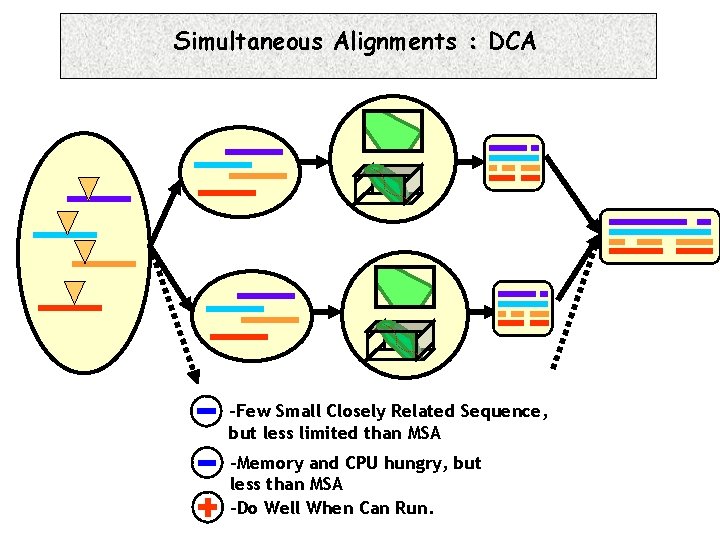
Simultaneous Alignments : DCA -Few Small Closely Related Sequence, but less limited than MSA -Memory and CPU hungry, but less than MSA -Do Well When Can Run.

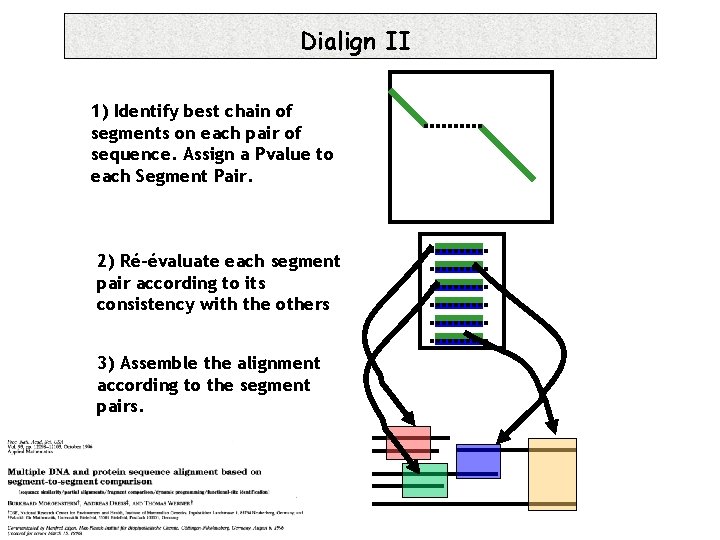
Dialign II 1) Identify best chain of segments on each pair of sequence. Assign a Pvalue to each Segment Pair. 2) Ré-évaluate each segment pair according to its consistency with the others 3) Assemble the alignment according to the segment pairs.

Muscle

Iterative Methods 7. 16. 1 Progressive -HMMs, HMMER, SAM, MUSCLE -Slow, Sometimes Inaccurate -Good Profile Generators

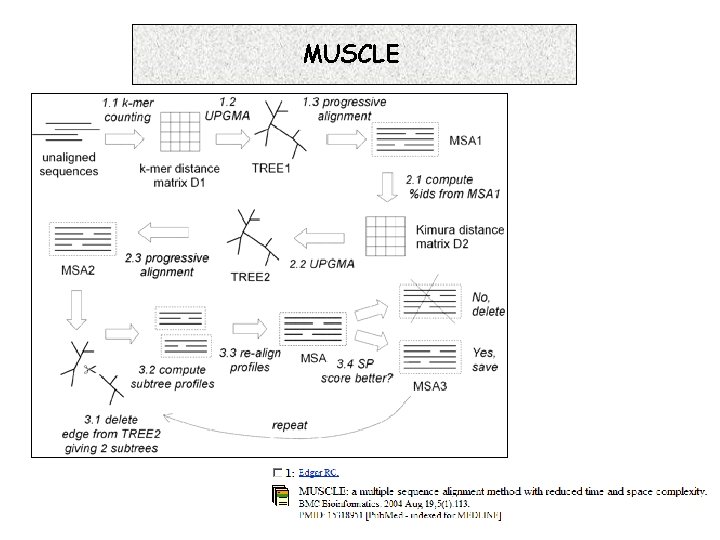
MUSCLE 7. 16. 1 Progressive

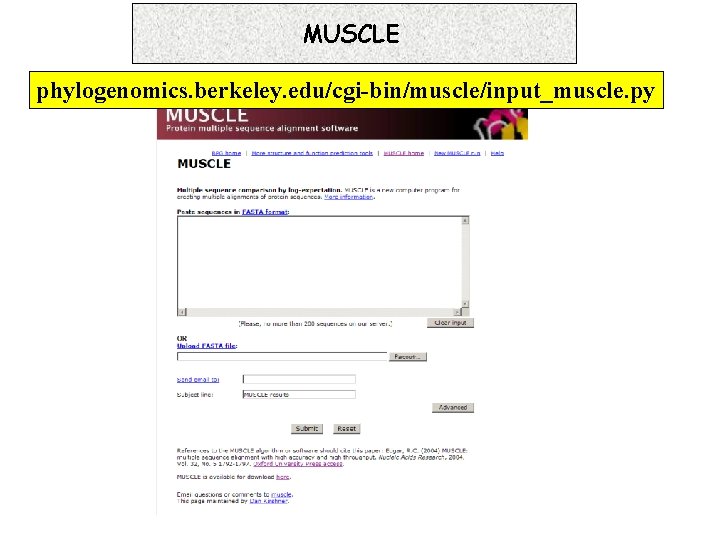
MUSCLE phylogenomics. berkeley. edu/cgi-bin/muscle/input_muscle. py 7. 16. 1 Progressive

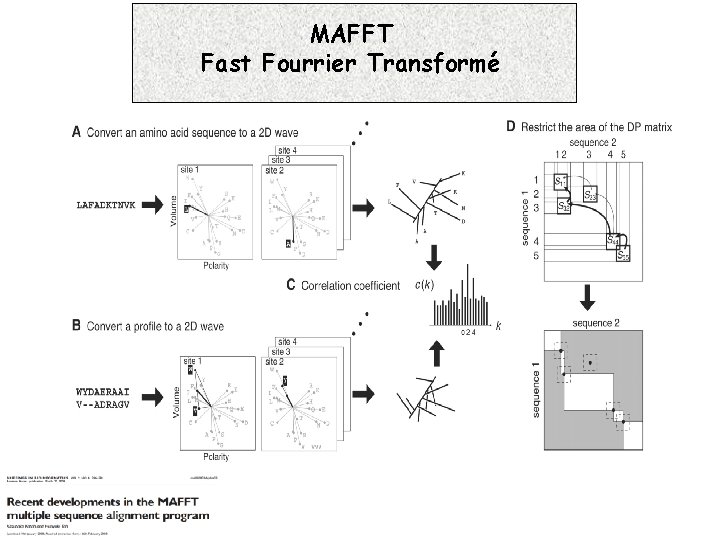
MAFFT Fast Fourrier Transformé

Prank

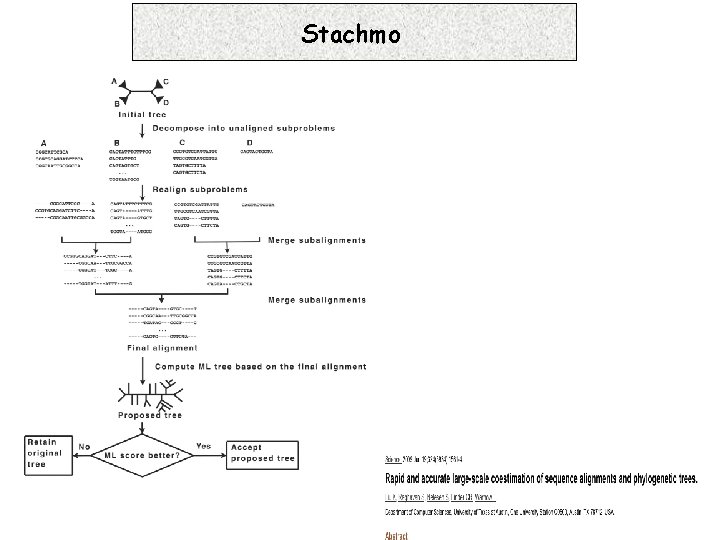
Stachmo

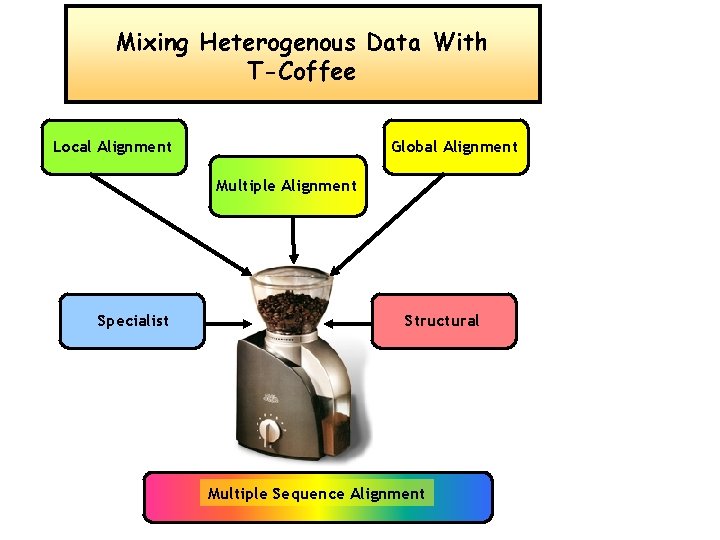
Mixing Heterogenous Data With T-Coffee Local Alignment Global Alignment Multiple Alignment Specialist Structural Multiple Sequence Alignment

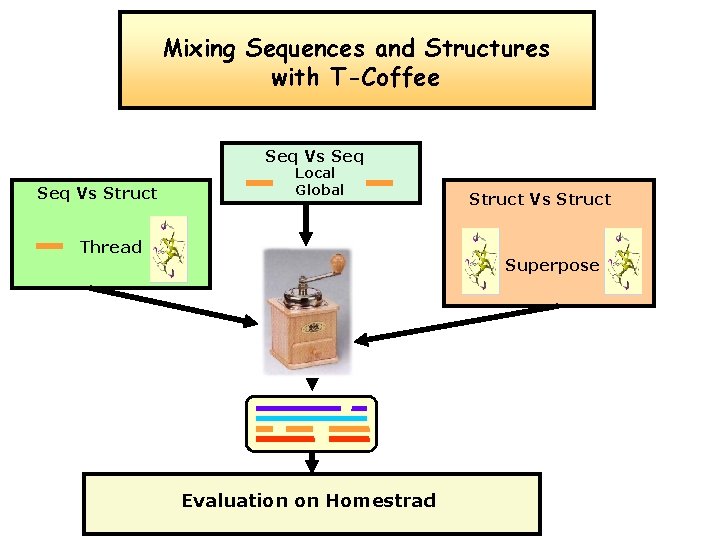
Mixing Sequences and Structures with T-Coffee Seq Vs Struct Local Global Thread Struct Vs Struct Superpose Evaluation on Homestrad

www. tcoffee. org

What is The Best Method ?


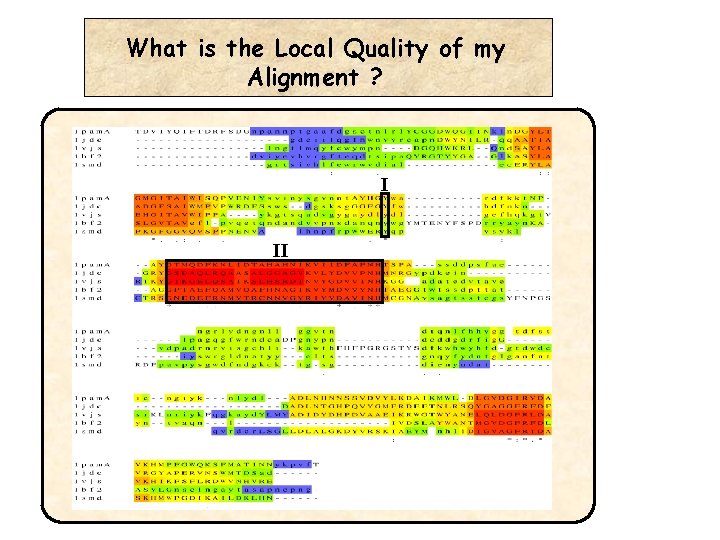
A better Question… • What is the Best Alignment ? • What is the best bit of my alignment ?

What is the Local Quality of my Alignment ? I II

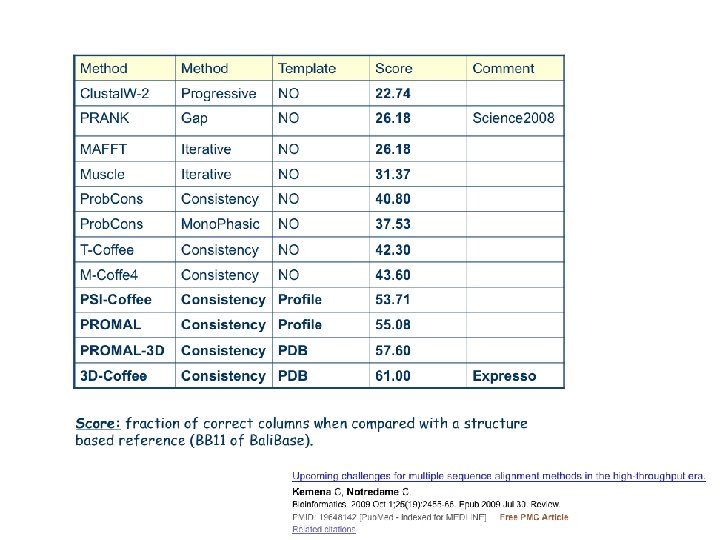
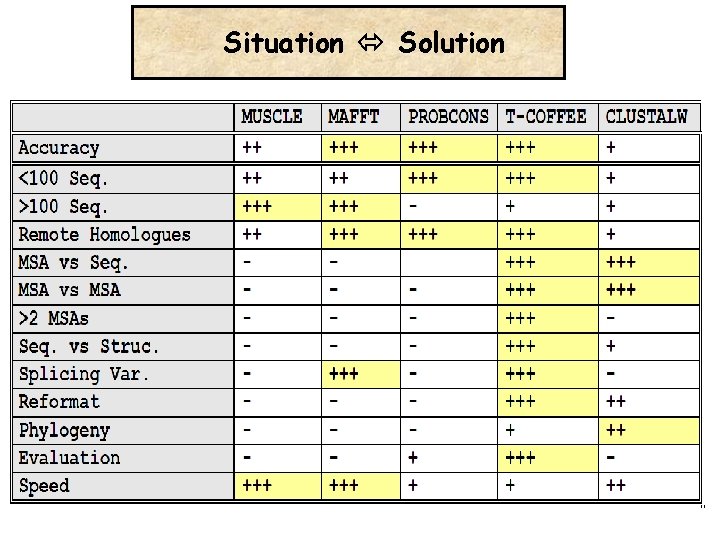
Choosing the right method

Situation Solution

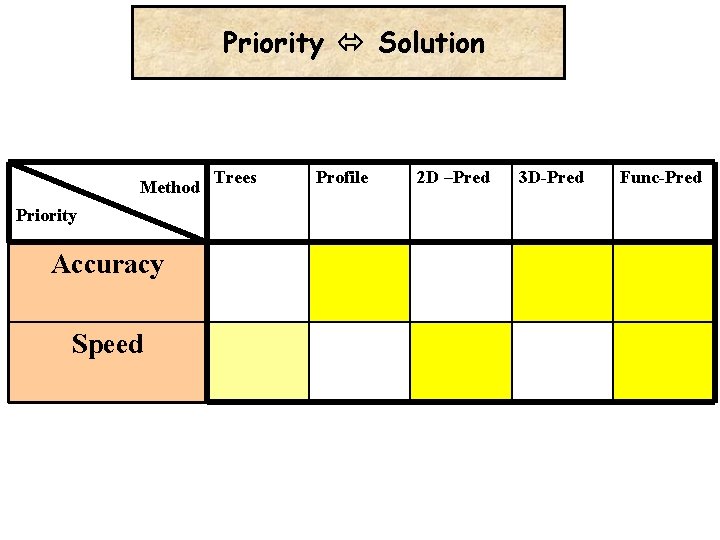
Priority Solution Method Priority Accuracy Speed Trees Profile 2 D –Pred 3 D-Pred Func-Pred

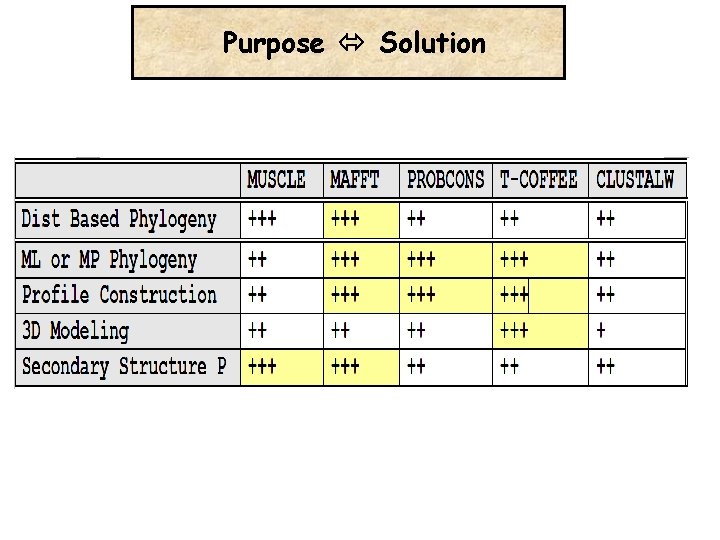
Purpose Solution

Conclusion

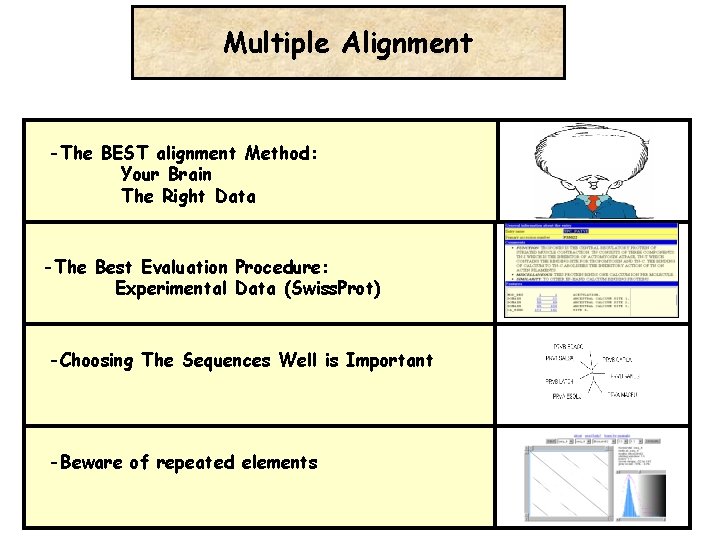
Multiple Alignment -The BEST alignment Method: Your Brain The Right Data -The Best Evaluation Procedure: Experimental Data (Swiss. Prot) -Choosing The Sequences Well is Important -Beware of repeated elements

Multiple Alignment Know Your Problem: What do you want to do with your MSA

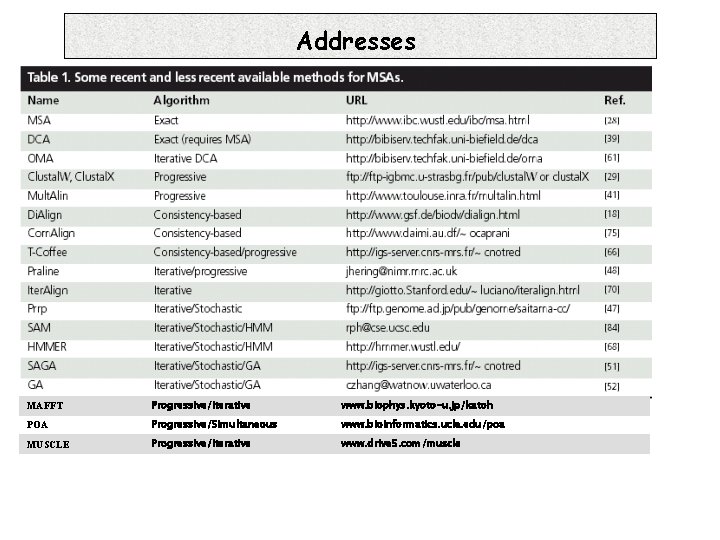
Addresses MAFFT Progressive/iterative www. biophys. kyoto-u. jp/katoh POA Progressive/Simultaneous www. bioinformatics. ucla. edu/poa MUSCLE Progressive/Iterative www. drive 5. com/muscle