Statistical Genomics Lecture 24 g BLUP Zhiwu Zhang

Statistical Genomics Lecture 24: g. BLUP Zhiwu Zhang Washington State University

Highlight • MAS by GAPIT • Method development • g. BLUP • Prediction of individuals without phenotypes

MAS by GAPIT • Setup GAPIT • Import data • Simulate phenotype • Validation
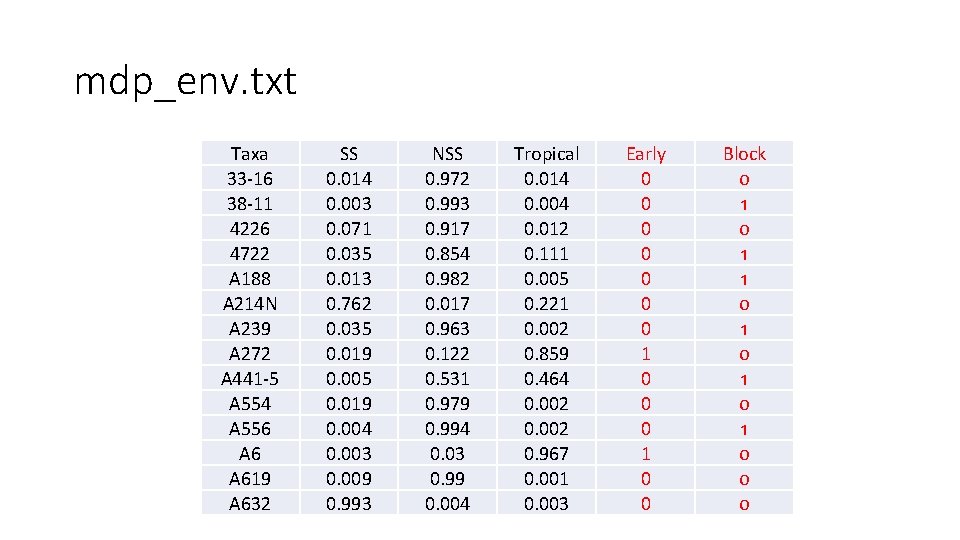
mdp_env. txt Taxa 33 -16 38 -11 4226 4722 A 188 A 214 N A 239 A 272 A 441 -5 A 554 A 556 A 619 A 632 SS 0. 014 0. 003 0. 071 0. 035 0. 013 0. 762 0. 035 0. 019 0. 004 0. 003 0. 009 0. 993 NSS 0. 972 0. 993 0. 917 0. 854 0. 982 0. 017 0. 963 0. 122 0. 531 0. 979 0. 994 0. 03 0. 99 0. 004 Tropical 0. 014 0. 004 0. 012 0. 111 0. 005 0. 221 0. 002 0. 859 0. 464 0. 002 0. 967 0. 001 0. 003 Early 0 0 0 0 1 0 0 Block 0 1 1 0 1 0 0 0

Import data and simulate phenotype library(compiler) #required for cmpfun source("http: //www. zzlab. net/GAPIT/gapit_functions. txt") my. GD=read. table(file="http: //zzlab. net/GAPIT/data/mdp_numeric. txt", head=T) my. GM=read. table(file="http: //zzlab. net/GAPIT/data/mdp_SNP_information. txt", head=T) my. CV=read. table(file="http: //zzlab. net/GAPIT/data/mdp_env. txt", head=T) #Simultate 10 QTN on the first half chromosomes X=my. GD[, -1] index 1 to 5=my. GM[, 2]<6 X 1 to 5 = X[, index 1 to 5] taxa=my. GD[, 1] GD. candidate=cbind(taxa, X 1 to 5) nqtn=50 set. seed(99164) setwd("~/Desktop/temp") my. Sim=GAPIT. Phenotype. Simulation(GD=GD. candidate, GM=my. GM[index 1 to 5, ], h 2=. 5, NQTN=nqtn, effectunit =. 95, QTNDist="normal", CV=my. CV, cveff=c(. 01, . 01))
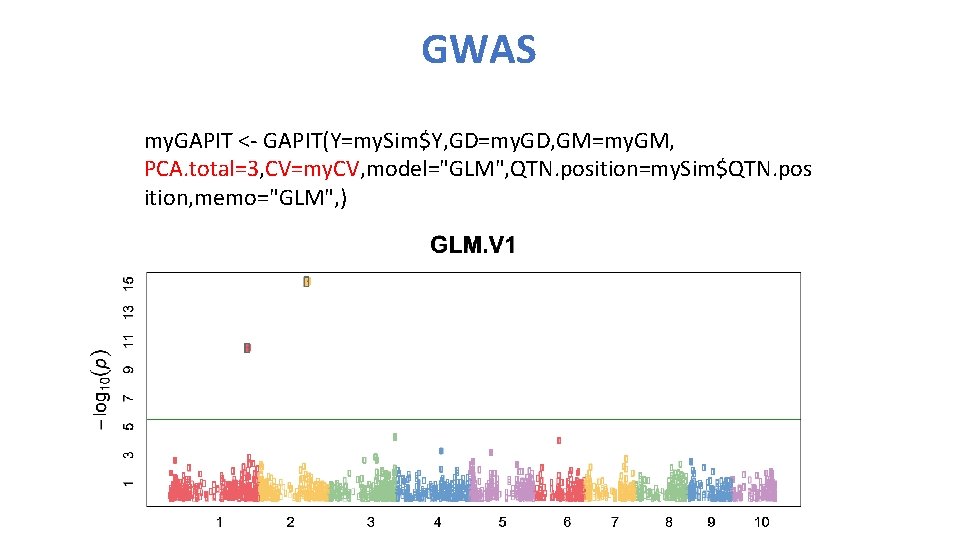
GWAS my. GAPIT <- GAPIT(Y=my. Sim$Y, GD=my. GD, GM=my. GM, PCA. total=3, CV=my. CV, model="GLM", QTN. position=my. Sim$QTN. pos ition, memo="GLM", )
![Prediction with PC and ENV order=match(my. Sim$Y[, 1], my. GAPIT$Pred[, 1]) my. Pred=my. GAPIT$Pred[order, Prediction with PC and ENV order=match(my. Sim$Y[, 1], my. GAPIT$Pred[, 1]) my. Pred=my. GAPIT$Pred[order,](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-7.jpg)
Prediction with PC and ENV order=match(my. Sim$Y[, 1], my. GAPIT$Pred[, 1]) my. Pred=my. GAPIT$Pred[order, ] ry 2=cor(my. Pred[, 8], my. Sim$Y[, 2])^2 ru 2=cor(my. Pred[, 8], my. Sim$u)^2 par(mfrow=c(2, 1), mar = c(3, 4, 1, 1)) plot(my. Pred[, 8], my. Sim$Y[, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[, 8], my. Sim$u) mtext(paste("R square=", ru 2, sep=""), side = 3)
![Choose top five SNPs ntop= nqtn*2 index=order(my. GAPIT$GWAS[, 4]) top=index[1: ntop] my. QTN=cbind(my. GAPIT$PCA[, Choose top five SNPs ntop= nqtn*2 index=order(my. GAPIT$GWAS[, 4]) top=index[1: ntop] my. QTN=cbind(my. GAPIT$PCA[,](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-8.jpg)
Choose top five SNPs ntop= nqtn*2 index=order(my. GAPIT$GWAS[, 4]) top=index[1: ntop] my. QTN=cbind(my. GAPIT$PCA[, 1: 4], my. CV[, 2: 3], my. GD[, c(top+1)]) #4

Estimate effects and prediction my. GAPIT 2 <- GAPIT( Y=my. Sim$Y, GD=my. GD, GM=my. GM, CV=my. QTN, model="GLM", QTN. position=my. Sim$QTN. position, SNP. test=FALSE, memo="GLM+QTN", ) #8 order=match(my. Sim$Y[, 1], my. GAPIT 2$Pred[, 1]) my. Pred=my. GAPIT 2$Pred[order, ] ry 2=cor(my. Pred[, 8], my. Sim$Y[, 2])^2 ru 2=cor(my. Pred[, 8], my. Sim$u)^2 par(mfrow=c(2, 1), mar = c(3, 4, 1, 1)) plot(my. Pred[, 8], my. Sim$Y[, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[, 8], my. Sim$u) mtext(paste("R square=", ru 2, sep=""), side = 3)

Validation by Dividing into training and testing #Cross validation set. seed(99164) n=nrow(my. Sim$Y) testing=sample(n, round(n/5), replace=F) training=-testing #GWAS on training to find QTNs my. GAPIT 3 <- GAPIT( Y=my. Sim$Y[training, ], GD=my. GD, GM=my. GM, CV=my. CV, PCA. total=3, model="GLM", memo="GWAS", )
![Estimate QTN effects in training ntop= nqtn*2 index=order(my. GAPIT 3$GWAS[, 4]) top=index[1: ntop] my. Estimate QTN effects in training ntop= nqtn*2 index=order(my. GAPIT 3$GWAS[, 4]) top=index[1: ntop] my.](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-11.jpg)
Estimate QTN effects in training ntop= nqtn*2 index=order(my. GAPIT 3$GWAS[, 4]) top=index[1: ntop] my. QTN=cbind(my. GAPIT$PCA[, 1: 4], my. CV[, 2: 3], my. GD[, c(top+1)]) PC #Estimate effect and prediction my. GAPIT 4 <- GAPIT( Y=my. Sim$Y[training, ], GD=my. GD, GM=my. GM, CV=my. QTN, model="GLM", SNP. test=FALSE, memo="GLM+QTN", ) ENV QTN
![Model fit in training order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT Model fit in training order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-12.jpg)
Model fit in training order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT 4$Pred[order, ] ry 2=cor(my. Pred[training, 8], my. Sim$Y[training, 2])^2 ru 2=cor(my. Pred[training, 8], my. Sim$u[training])^2 par(mfrow=c(2, 1), mar = c(3, 4, 1, 1)) plot(my. Pred[training, 8], my. Sim$Y[training, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[training, 8], my. Sim$u[training]) mtext(paste("R square=", ru 2, sep=""), side = 3)
![Accuracy in testing order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT 4$Pred[order, Accuracy in testing order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT 4$Pred[order,](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-13.jpg)
Accuracy in testing order=match(my. Sim$Y[, 1], my. GAPIT 4$Pred[, 1]) my. Pred=my. GAPIT 4$Pred[order, ] ry 2=cor(my. Pred[testing, 8], my. Sim$Y[testing, 2])^2 ru 2=cor(my. Pred[testing, 8], my. Sim$u[testing])^2 par(mfrow=c(2, 1), mar = c(3, 4, 1, 1)) plot(my. Pred[testing, 8], my. Sim$Y[testing, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[testing, 8], my. Sim$u[testing]) mtext(paste("R square=", ru 2, sep=""), side = 3) Why so good? Only 2 QTNs
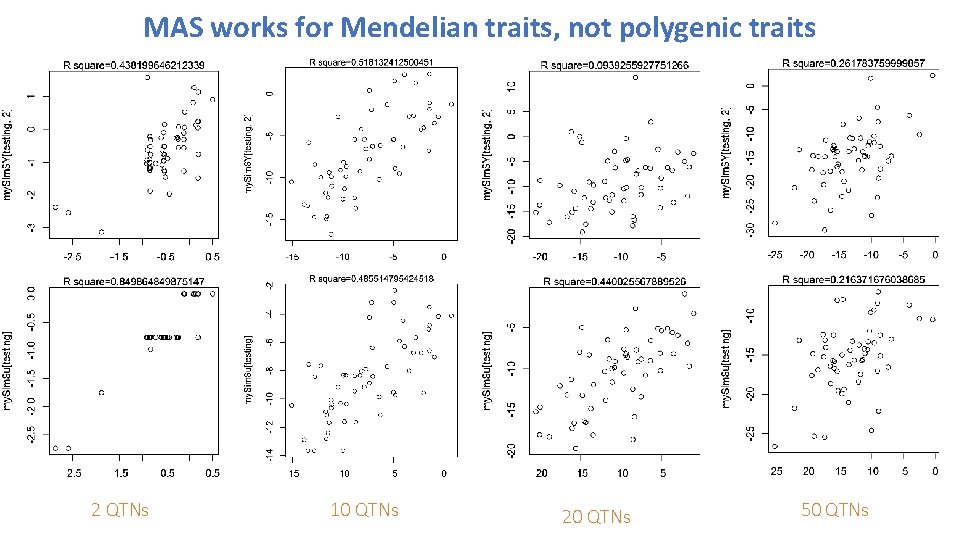
MAS works for Mendelian traits, not polygenic traits 2 QTNs 10 QTNs 20 QTNs 50 QTNs
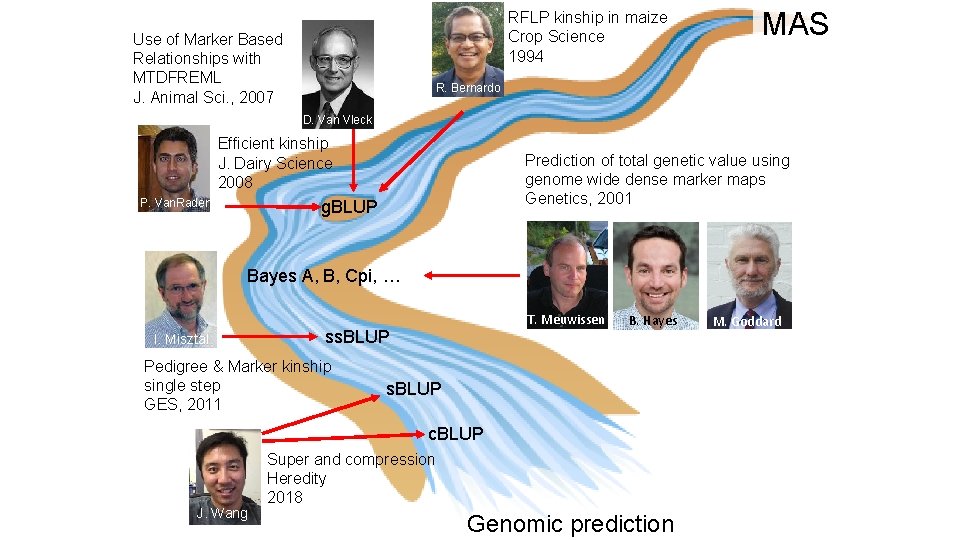
RFLP kinship in maize Crop Science 1994 Use of Marker Based Relationships with MTDFREML J. Animal Sci. , 2007 MAS R. Bernardo D. Van Vleck Efficient kinship J. Dairy Science 2008 P. Van. Raden Prediction of total genetic value using genome wide dense marker maps Genetics, 2001 g. BLUP Bayes A, B, Cpi, … I. Misztal T. Meuwissen ss. BLUP Pedigree & Marker kinship single step GES, 2011 B. Hayes s. BLUP c. BLUP J. Wang Super and compression Heredity 2018 Genomic prediction M. Goddard

g. BLUP Prediction of maize single-cross performance using RFLPs and information related hybrids Rex Bernardo Crop Science, 1994 YM = C V− 1 Yp y. M = m × 1 vector of predicted yields of missingle crosses; C = m × n matrix of genetic covariances between the missing and predictor hybrids; V = n × n matrix of phenotypic variances and covariances among predictor hybrids; y. P = nn × 1 vector of predictor hybrid yields corrected for trial effects.

g. BLUP reinvent

Multiple Trait Derivative Free REML (MTDFREML) Welcome to the Multiple Trait Derivative Free REML (MTDFREML) home page. The programs were developed by Keith Boldman and Dale Van Vleck. Evolutionary development and debugging support have also been provided by by Lisa Kriese and Curt Van Tassell. Please contact Curt Van Tassell (e-mail curtvt@aipl. arsusda. gov) or Dale Van Vleck. (e-mail lvanvleck@unlnotes. unl. edu) with any problems with the programs or discovered bugs. • Obtaining the MTDFREML programs • Get the manual • Sample analyses • Enter user information using web browser that handles forms • FTP the userinfo. txt file to enter user information (then mail completed form) • Get the Microsoft Powerstation fix for Windows 95 (compressed)
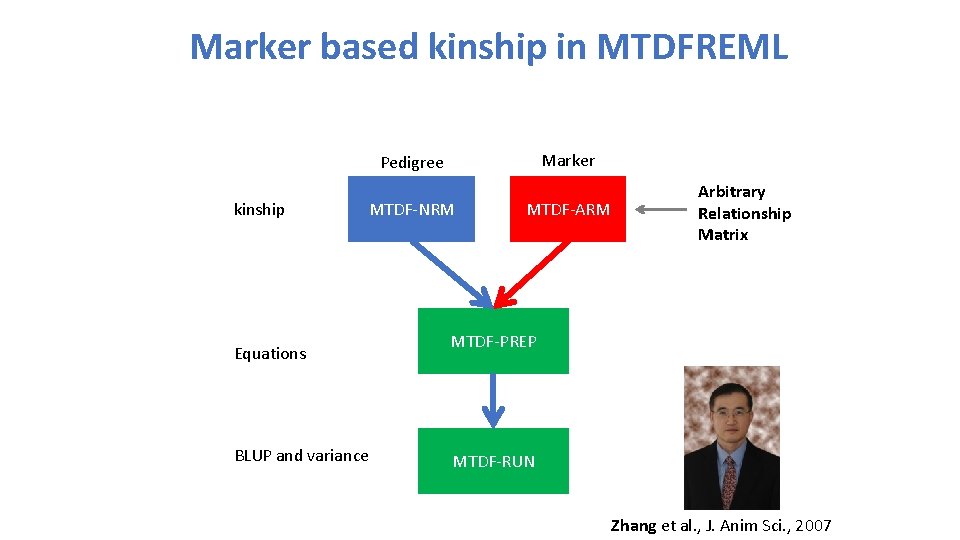
Marker based kinship in MTDFREML Marker Pedigree kinship MTDF-NRM Equations BLUP and variance MTDF-ARM Arbitrary Relationship Matrix MTDF-PREP MTDF-RUN Zhang et al. , J. Anim Sci. , 2007

Mixed Linear Model (MLM)

Z matrix observation mean b= [ b 0 y [1 PC 2 b 1 x 1 SNP b 2 ] u= x 2 ] =X y = Xb + Zu +e Ind 1 [ u 1 1 0 Ind 2 u 2 0 1 … … 0 0 … … Z Ind 9 Ind 10 u 9 u 10 ] 0 0 1

Generic Z matrix u= Ind 1 [ u 1 1 0 Ind 2 u 2 0 1 … … 0 0 … … ZR Ind 9 Ind 10 u 9 u 10 ] 0 0 1 Ind 12 u 11 u 12 0 0 0 0 … … … ZI Ind 19 Ind 20 u 19 u 20 ] 0 0 0 0

Efficient kinship algorithm • M: n individual by m SNPs • M: -1, 0 and 1 • Pi: frequency of 2 nd allele for SNP i • P: Column of i is 2(pi-. 5) • Z=M-P MMt, Efficient g. BLUP=Ridge Regression J. Dairy Sci. 2008. 91 (11) 4414 -4423. Efficient Methods to Compute Genomic Predictions P. M. Van. Raden Paul Van. Raden: Image Number K 7168 -6

Pedigree + Marker Non genotyped Pedigree kinship Marker kinship Pedigree kinship of non genotyped Arbitrary weight

Henderson's formula
![g. BLUP by GAPIT my. GAPIT 5 <- GAPIT( Y=my. Sim$Y[training, ], GD=my. GD, g. BLUP by GAPIT my. GAPIT 5 <- GAPIT( Y=my. Sim$Y[training, ], GD=my. GD,](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-26.jpg)
g. BLUP by GAPIT my. GAPIT 5 <- GAPIT( Y=my. Sim$Y[training, ], GD=my. GD, GM=my. GM, PCA. total=3, CV=my. CV, model="g. BLUP", SNP. test=FALSE, memo="g. BLUP", )
![Training True BV predicted phenotype ry 2=cor(my. Pred[training, 8], my. Sim$Y[training, 2])^2 ru 2=cor(my. Training True BV predicted phenotype ry 2=cor(my. Pred[training, 8], my. Sim$Y[training, 2])^2 ru 2=cor(my.](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-27.jpg)
Training True BV predicted phenotype ry 2=cor(my. Pred[training, 8], my. Sim$Y[training, 2])^2 ru 2=cor(my. Pred[training, 8], my. Sim$u[training])^2 ry 2. blup=cor(my. Pred[training, 5], my. Sim$Y[training, 2])^2 ru 2. blup=cor(my. Pred[training, 5], my. Sim$u[training])^2 par(mfrow=c(2, 2), mar = c(3, 4, 1, 1)) plot(my. Pred[training, 8], my. Sim$Y[training, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[training, 8], my. Sim$u[training]) mtext(paste("R square=", ru 2, sep=""), side = 3) plot(my. Pred[training, 5], my. Sim$Y[training, 2]) mtext(paste("R square=", ry 2. blup, sep=""), side = 3) plot(my. Pred[training, 5], my. Sim$u[training]) mtext(paste("R square=", ru 2. blup, sep=""), side = 3) phenotype predicted BV order=match(my. Sim$Y[, 1], my. GAPIT 5$Pred[, 1]) my. Pred=my. GAPIT 5$Pred[order, ]
![Testing 50 QTNs: g. BLUP=MAS True BV predicted phenotype ry 2=cor(my. Pred[testing, 8], my. Testing 50 QTNs: g. BLUP=MAS True BV predicted phenotype ry 2=cor(my. Pred[testing, 8], my.](http://slidetodoc.com/presentation_image_h/d9a9c0f7ed4d2749c8c8034011dfbf19/image-28.jpg)
Testing 50 QTNs: g. BLUP=MAS True BV predicted phenotype ry 2=cor(my. Pred[testing, 8], my. Sim$Y[testing, 2])^2 ru 2=cor(my. Pred[testing, 8], my. Sim$u[testing])^2 ry 2. blup=cor(my. Pred[testing, 5], my. Sim$Y[testing, 2])^2 ru 2. blup=cor(my. Pred[testing, 5], my. Sim$u[testing])^2 par(mfrow=c(2, 2), mar = c(3, 4, 1, 1)) plot(my. Pred[testing, 8], my. Sim$Y[testing, 2]) mtext(paste("R square=", ry 2, sep=""), side = 3) plot(my. Pred[testing, 8], my. Sim$u[testing]) mtext(paste("R square=", ru 2, sep=""), side = 3) plot(my. Pred[testing, 5], my. Sim$Y[testing, 2]) mtext(paste("R square=", ry 2. blup, sep=""), side = 3) plot(my. Pred[testing, 5], my. Sim$u[testing]) mtext(paste("R square=", ru 2. blup, sep=""), side = 3) phenotype predicted BV order=match(my. Sim$Y[, 1], my. GAPIT 5$Pred[, 1]) my. Pred=my. GAPIT 5$Pred[order, ]

Highlight • The power of molecular breeding • Method development • g. BLUP • Prediction of individuals without phenotypes
- Slides: 29