SIMULATIONS OF PEPTIDE FOLDING and DYNAMICS Krzysztof Kuczera




![Folding: transition state w(n) = -k. Tln[P(n)] w(n) = -k. Tln[P(n)/C(n)] C(n) = 17!/n!(17 Folding: transition state w(n) = -k. Tln[P(n)] w(n) = -k. Tln[P(n)/C(n)] C(n) = 17!/n!(17](https://slidetodoc.com/presentation_image_h/9f5e33636deb2d48cbe9cd277e98be0e/image-17.jpg)



- Slides: 27

SIMULATIONS OF PEPTIDE FOLDING and DYNAMICS Krzysztof Kuczera Departments of Chemistry and Molecular Biosciences University of Kansas IMA Workshop Jan 14 -18, 2008

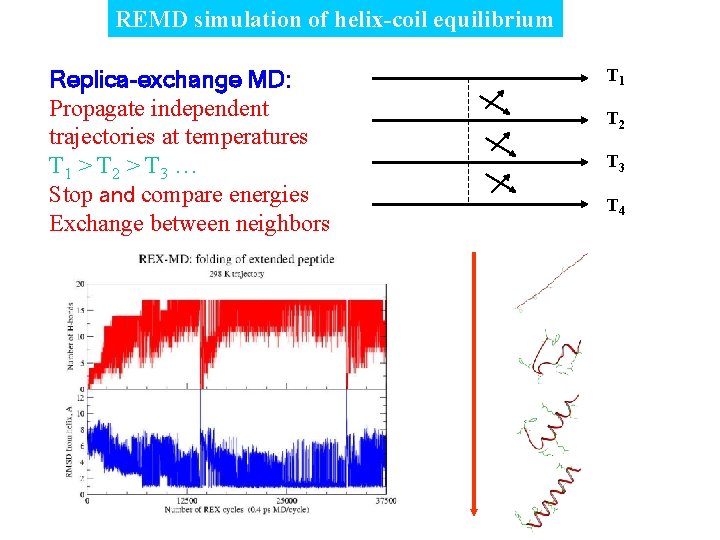
Replica-exchange molecular dynamics T 1 T 2 T 3 Propagate independent trajectories at temperatures T 1 > T 2 > T 3 … Stop and compare energies Exchange between neighbors T 4 Advantages: + accelerated sampling @ low T + Boltzmann distributions @ all T + Minimal process communication D 0 D>0

21 -residue helix-forming peptide Blocked peptide: Ac-WAAAH+-(AAARA)3 -A-NH 2 21 residues Simulation 1: equilibrium distribution start: helix trajectory: 30 ns Simulation 2: folding start: extended trajectory: 15 ns Method: CHARMM with CMAP electrostatics: GBMV nonpolar: DGnp=- g DA with g = 4 cal/(mol Å2) REMD at 8 temperatures, 280 -450 K MMTSB tool kit (C. L. Brooks, Scripps)





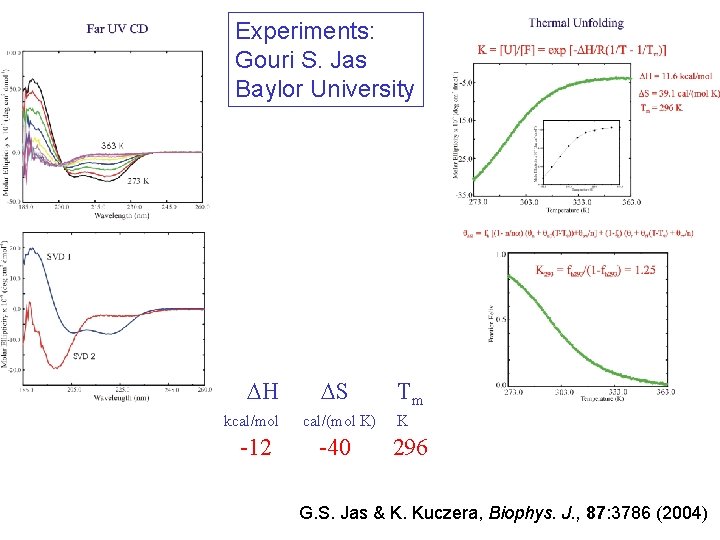
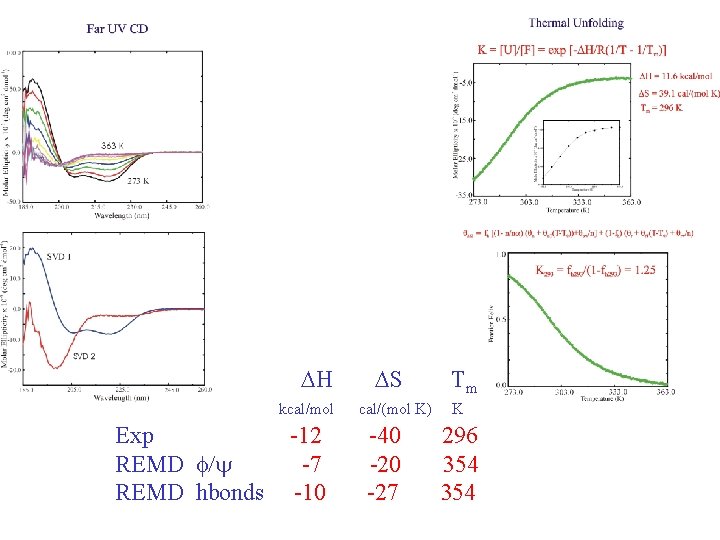
Experiments: Gouri S. Jas Baylor University DH kcal/mol -12 DS cal/(mol K) -40 Tm K 296 G. S. Jas & K. Kuczera, Biophys. J. , 87: 3786 (2004)

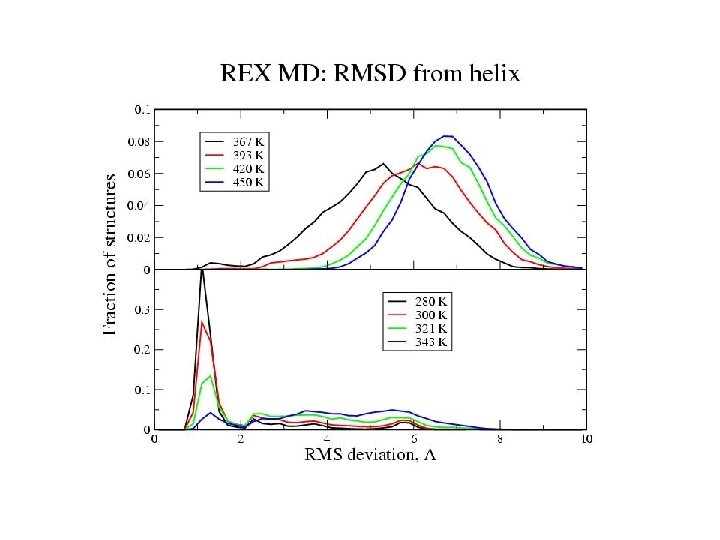
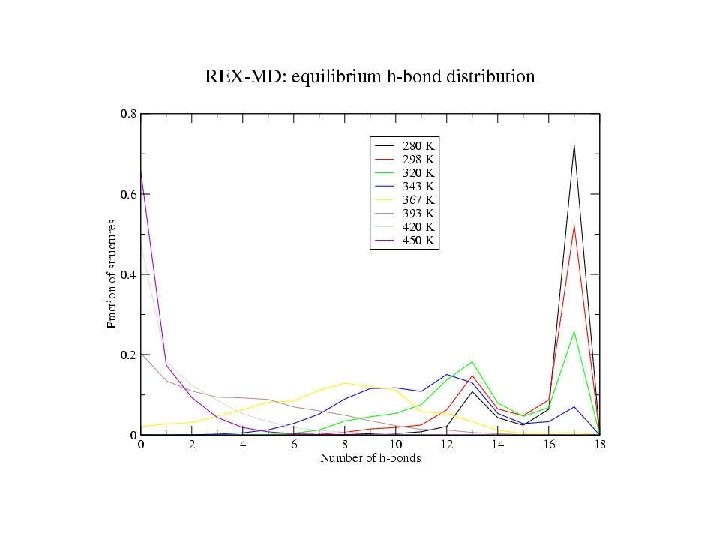
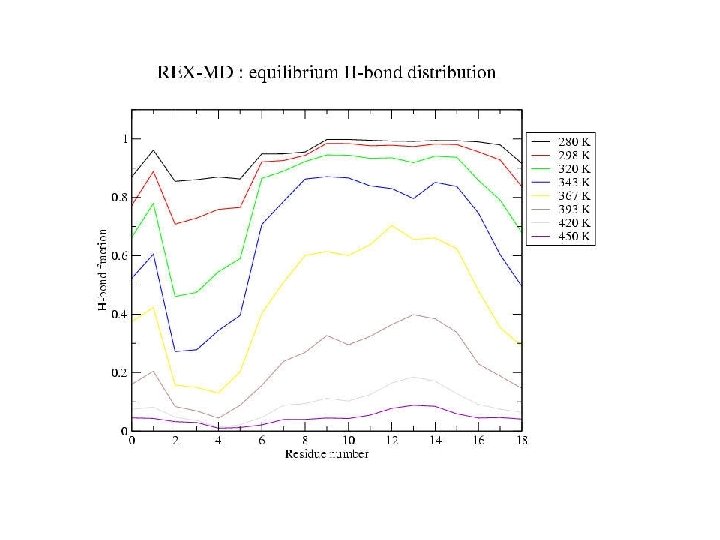
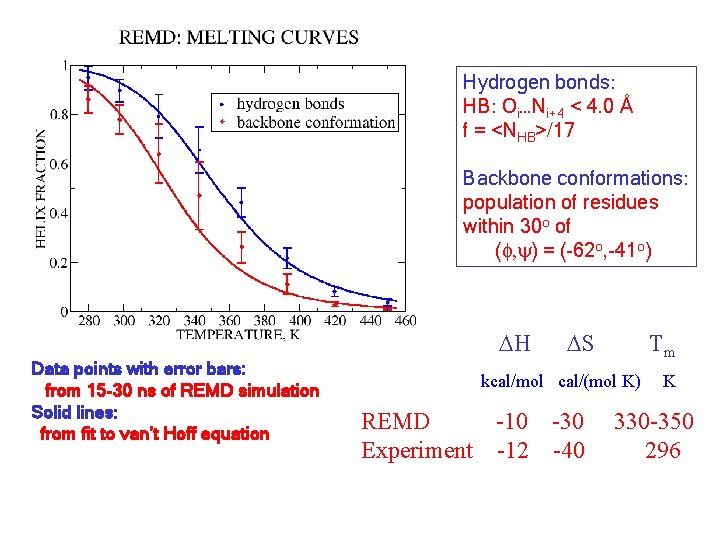
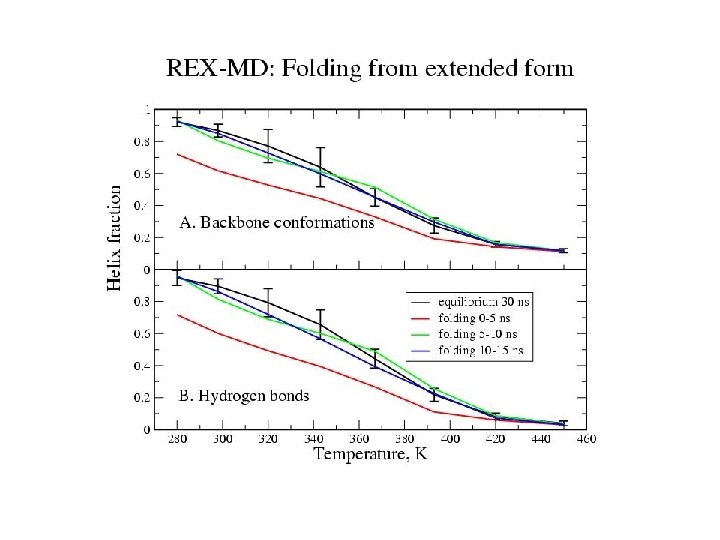
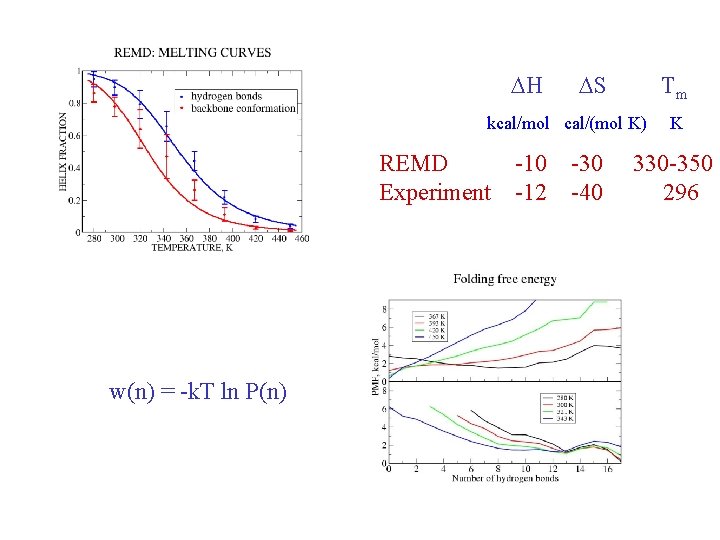
Hydrogen bonds: HB: Oi…Ni+4 < 4. 0 Å f = <NHB>/17 Backbone conformations: population of residues within 30 o of (f, y) = (-62 o, -41 o) DH Data points with error bars: from 15 -30 ns of REMD simulation Solid lines: from fit to van’t Hoff equation DS Tm kcal/mol cal/(mol K) REMD -10 -30 Experiment -12 -40 K 330 -350 296

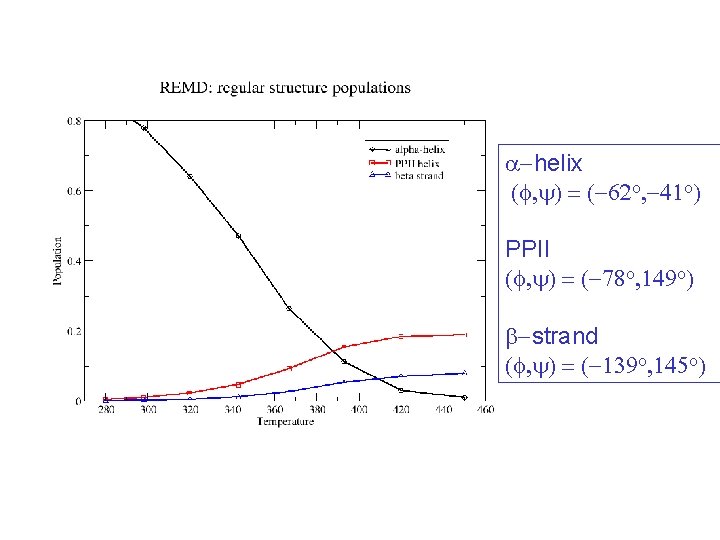
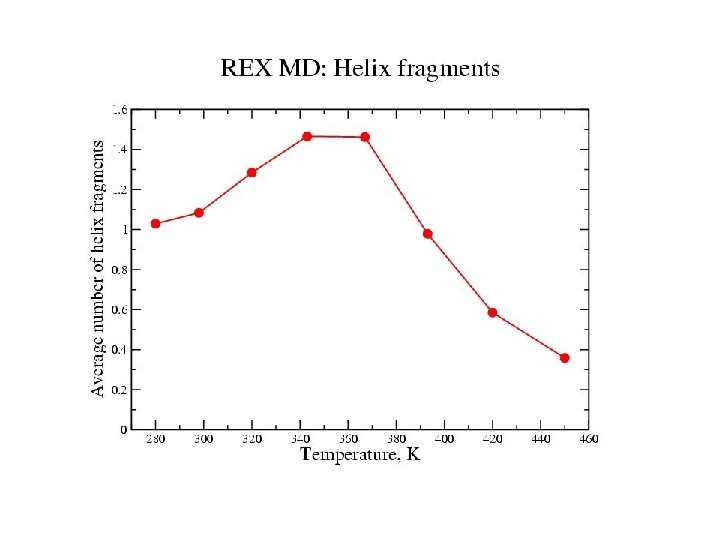
Zimm-Bragg Parameters Statistical weights partition function coil : 1 helix with h-bond: w=s helix without h-bond: v = s 1/2 REMD results at 300 K: helix fraction = 0. 78 number of helix fragments = 1. 08 w = 1. 86 v = 0. 11 Previous Ala-based peptide simulations: w = 1. 12 – 2. 12 v = 0. 06 – 0. 30



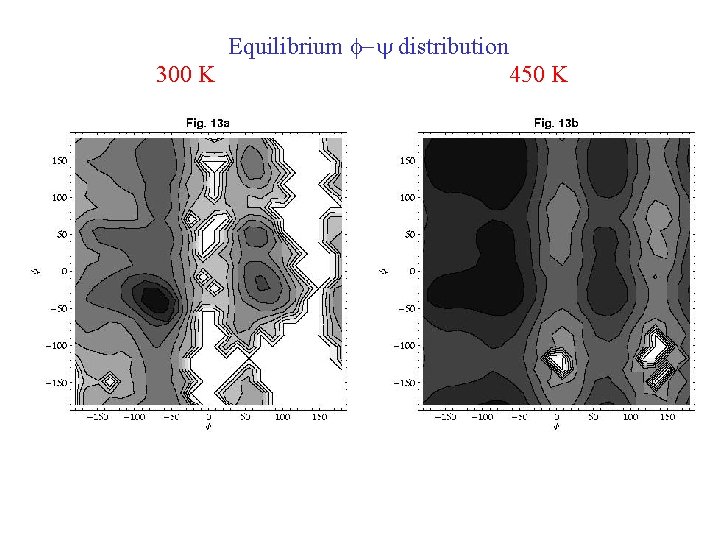
Equilibrium f-y distribution 300 K 450 K

a-helix (f, y) = (-62 o, -41 o) PPII (f, y) = (-78 o, 149 o) b-strand (f, y) = (-139 o, 145 o)


![Folding transition state wn k TlnPn wn k TlnPnCn Cn 17n17 Folding: transition state w(n) = -k. Tln[P(n)] w(n) = -k. Tln[P(n)/C(n)] C(n) = 17!/n!(17](https://slidetodoc.com/presentation_image_h/9f5e33636deb2d48cbe9cd277e98be0e/image-17.jpg)
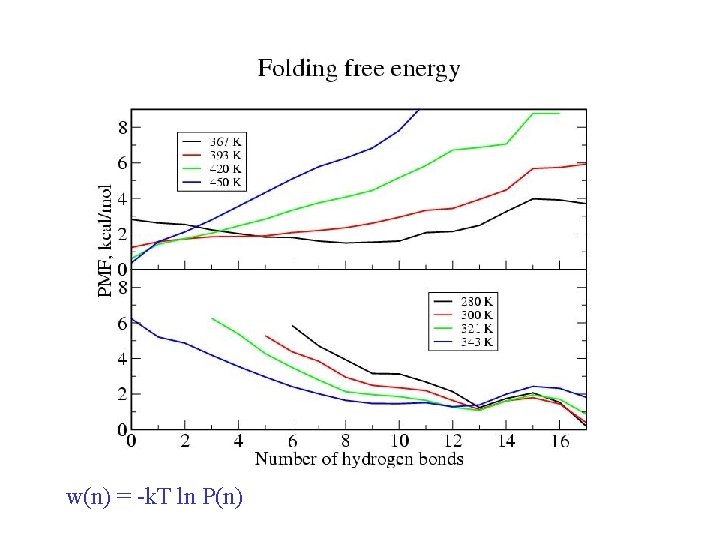
Folding: transition state w(n) = -k. Tln[P(n)] w(n) = -k. Tln[P(n)/C(n)] C(n) = 17!/n!(17 -n)!

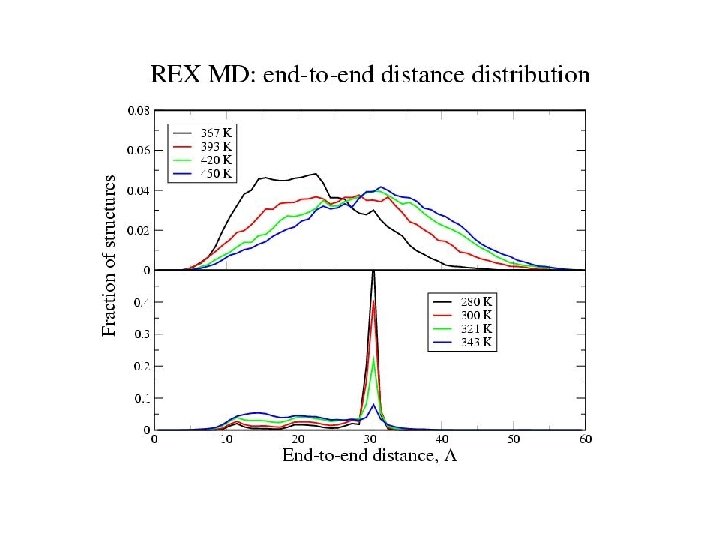
Equilibrium REMD: conclusions • We have calculated an equilibrium melting curve for a helix-forming peptide with GB/SA model • Thermodynamics qualitatively correct Melting temperature exaggerated • Global free energy minimum = a-helix at low T = coil at high T • Microscopic information about helix unfolding - unfolding initiated at termini transition state ½ helix - formation of compact structures at low T - b and PPII structures at high T

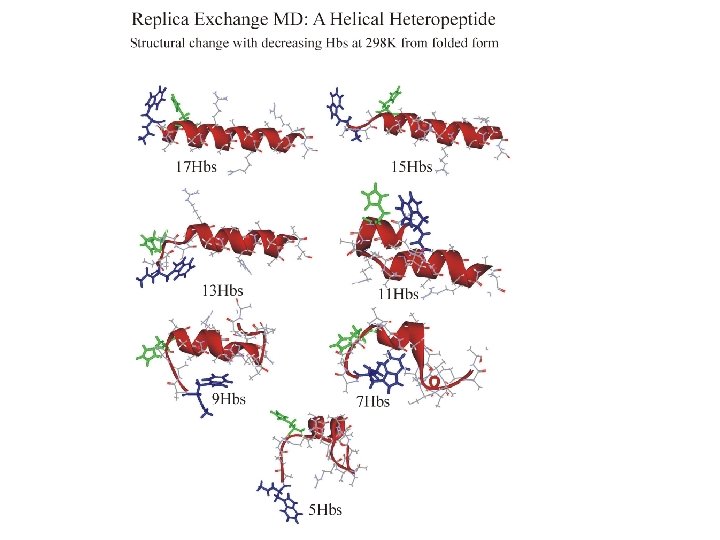
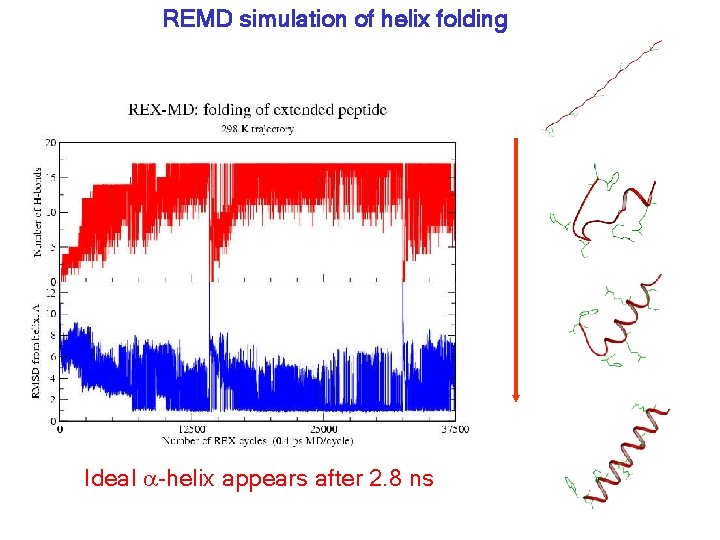
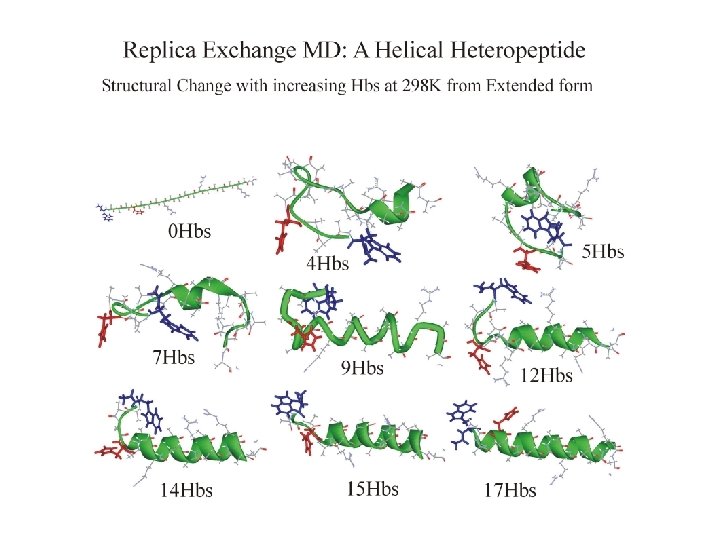
REMD simulation of helix folding Ideal a-helix appears after 2. 8 ns



Folding REMD: conclusions • a-helix structure found after ca. 3 ns simulation time • confirm that helix is global FE minimum, not a memory effect • folding in room T replica is sequential • structures and properties sampled are similar to trajectory starting from helix • conformational sampling acceleration of REX-MD over direct MD is ca. 100 overall (ca. 12 per CPU) [3 ns folding in REMD vs. 300 ns exp time scale]

Acknowledgments • Gerald Lushington, University of Kansas Molecular Modeling and Graphics Laboratory – Athlon cluster • Michael Feig, John Caranicolas and Charles L. Brooks III MMTSB Tool Set (2001), The Scripps Research Institute • Gouri S. Jas, Baylor University - experiments • ACS PRF $$$

DH DS Tm kcal/mol cal/(mol K) REMD -10 -30 Experiment -12 -40 w(n) = -k. T ln P(n) K 330 -350 296

REMD simulation of helix-coil equilibrium Replica-exchange MD: Propagate independent trajectories at temperatures T 1 > T 2 > T 3 … Stop and compare energies Exchange between neighbors T 1 T 2 T 3 T 4

DH kcal/mol Exp REMD f/y REMD hbonds -12 -7 -10 DS cal/(mol K) -40 -27 Tm K 296 354

w(n) = -k. T ln P(n)