Protein Folding BL 4010 10 06 Protein folding

Protein Folding BL 4010 10. 06
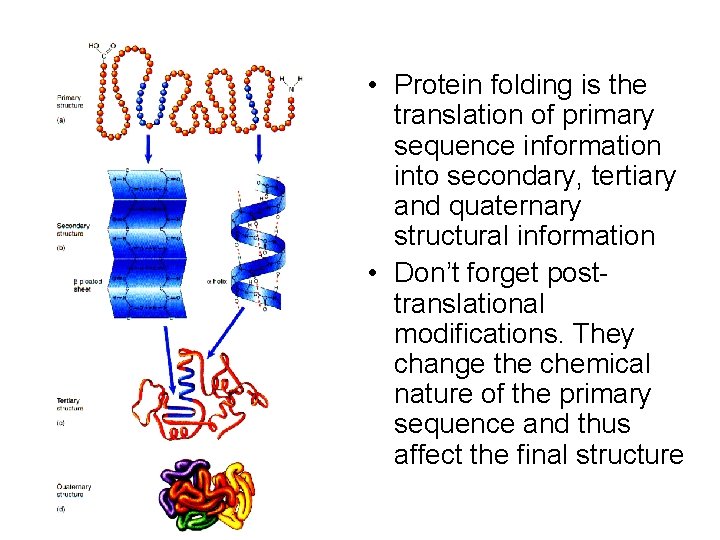
• Protein folding is the translation of primary sequence information into secondary, tertiary and quaternary structural information • Don’t forget posttranslational modifications. They change the chemical nature of the primary sequence and thus affect the final structure
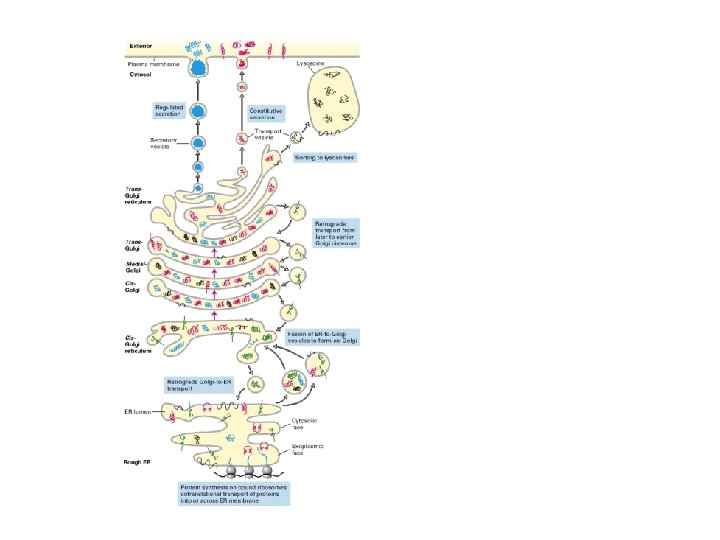
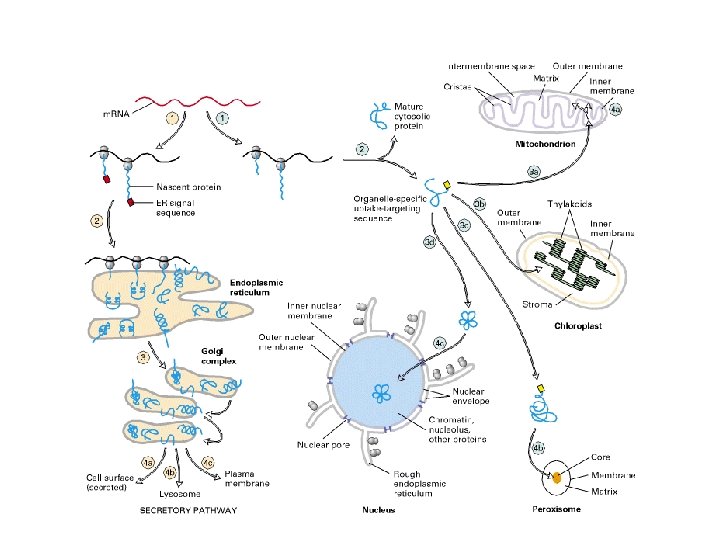
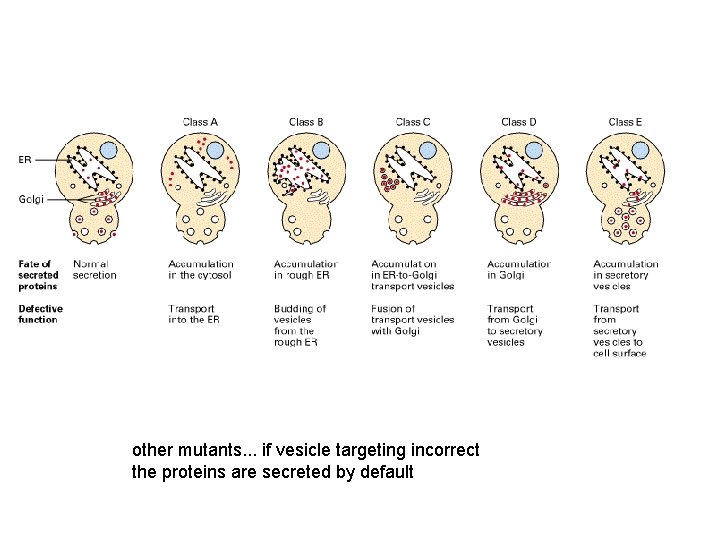
other mutants. . . if vesicle targeting incorrect the proteins are secreted by default

Michel Van Lun http: //xray. bmc. uu. se/~michiel/research_files/ext 1_10 ns. mpg Paul Berg (protein synthesis represented by dance) http: //video. google. com/videoplay? docid=-2657697036715872139

Protein Folding Models • Hydrophobic collapse (oil drop model) – entropy driven, temperature dependent – size? • Nucleation theory – alpha helical and reverse turn • Viscosity/collision • Energy landscape
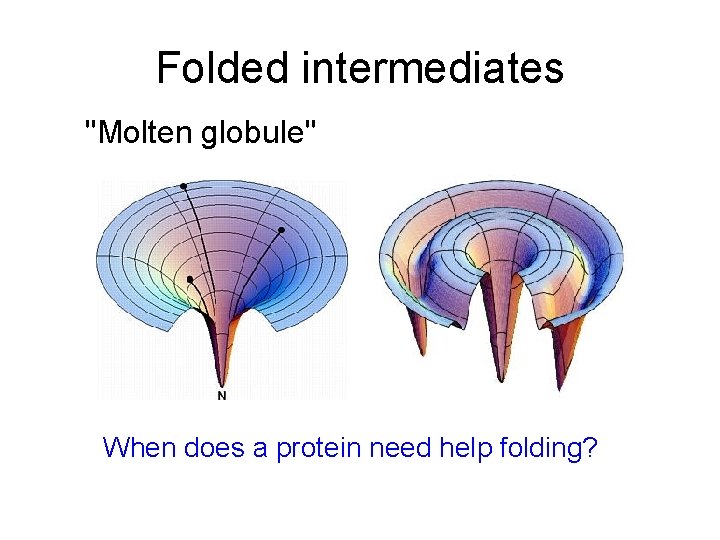
Folded intermediates "Molten globule" When does a protein need help folding?

Protein Folding Machinery • When does protein folding need help? – During synthesis – Upon stress (esp. heat shock) • Players – – – Chaperone and chaperonins (heat shock factors) Peptide binding proteins ATP Disulfide isomerase Petidyl prolyl cis-trans isomerase

What happens if proteins don't fold properly?
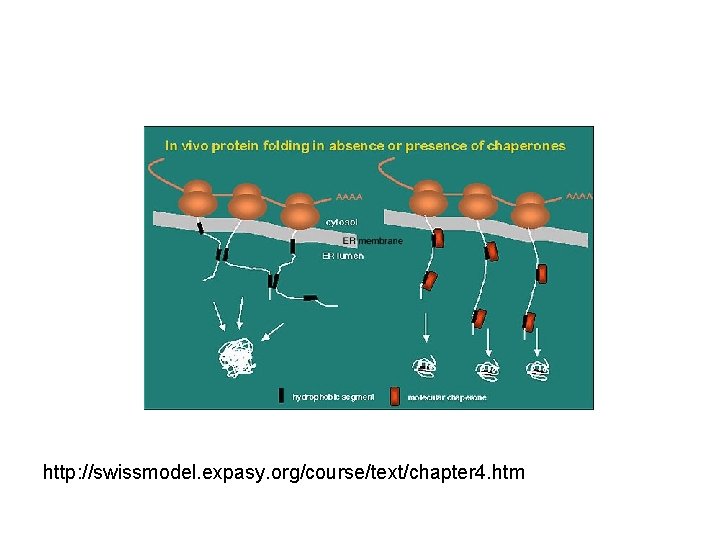
http: //swissmodel. expasy. org/course/text/chapter 4. htm

Improperly folded proteins • Non-functional – improper interactions – nonproductive interactions • Aggregation • Targeted for proteolysis/degradation • Resource drain (commitment of material and energy)
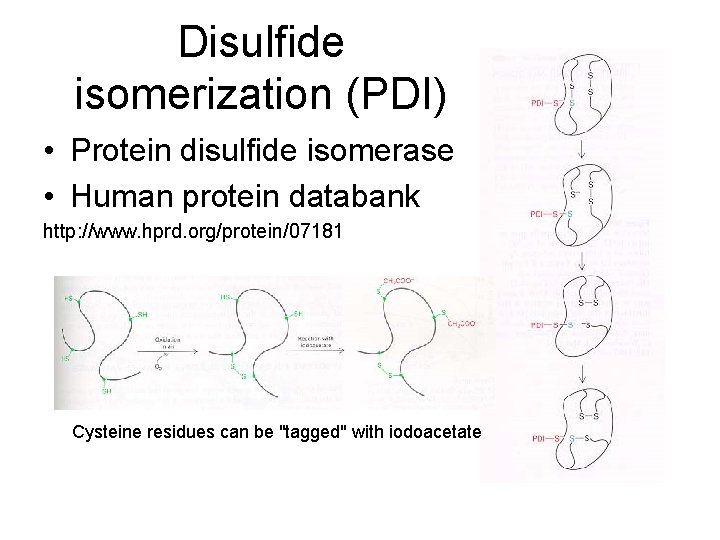
Disulfide isomerization (PDI) • Protein disulfide isomerase • Human protein databank http: //www. hprd. org/protein/07181 Cysteine residues can be "tagged" with iodoacetate
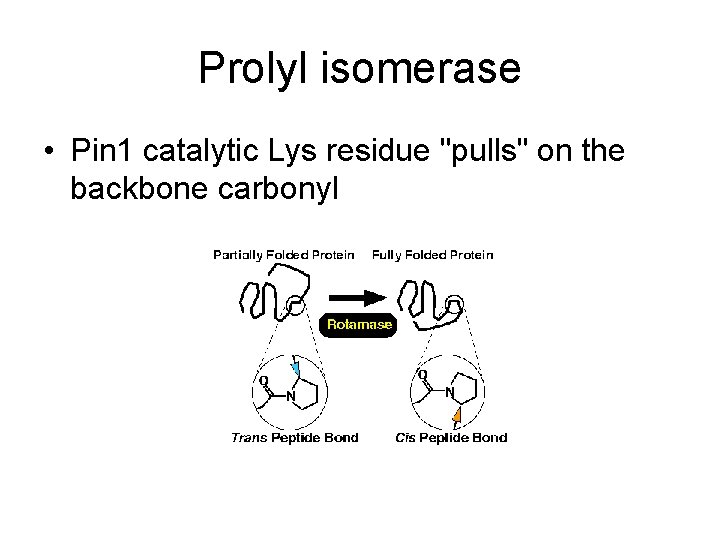
Prolyl isomerase • Pin 1 catalytic Lys residue "pulls" on the backbone carbonyl

Pin 1 an important PPI • Breast cancer: – Pin 1 isomerizes phosphorylated Ser/Thr-Pro – Regulates various cellular processes. – Overexpressed in human breast cancer. – Overexpression causes upregulation of cyclin D 1 and transformation of breast epithelial cells. • Alzheimer's Disease – Pin 1 becomes depleted from the nucleus of diseased neurons – Redirected to the large amounts of abnormally phosphorylated tau proteins that will aggregate into filaments. – This depletion from the nucleus may ultimately contribute to neuronal cell death by reactivating the cell cycle.

What property do misfolded proteins share? • We have seen that protein/protein interactions can be highly specific • Chaperones and Chaperonins must bind a wide variety of different proteins • What is the mechanism of recognition that protein is misfolded?

Chaperones: Variations on a theme

Dna. J (Hsp 40) • Binds folding and misfolded proteins • Contains a 30 aa glycine-rich region ('G' domain'), a central Zn binding domain containing 4 repeats of a CXXCXGXG ('CRR' domain) and a C-terminal region of 120 to 170 residues • Associates with Dna. K • Discovered as requirement for viral replication
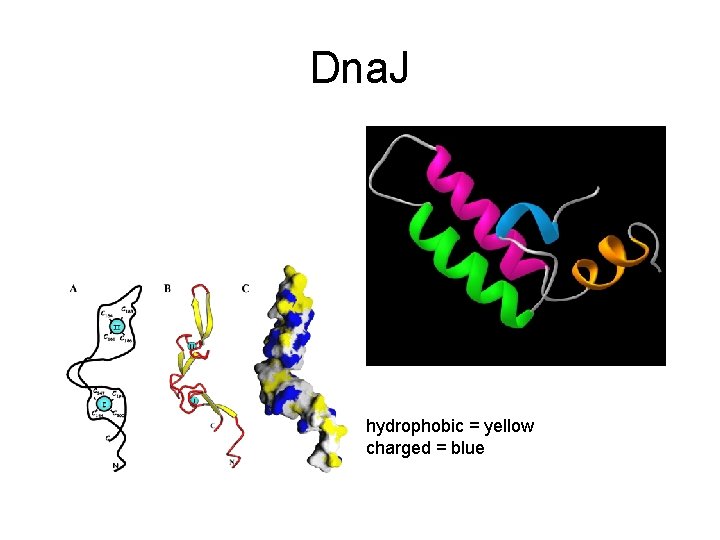
Dna. J hydrophobic = yellow charged = blue

Dna. K (Hsp 70) • Binds folding and misfolded proteins • Associates with Dna. J • ATP binding and hydrolysis • ER homologue Grp 78 • The interaction of Dna. KATP to substrate proteins is transient and characterized by high on/off rates • 2 domains – alpha helix – beta sheet: peptide binding
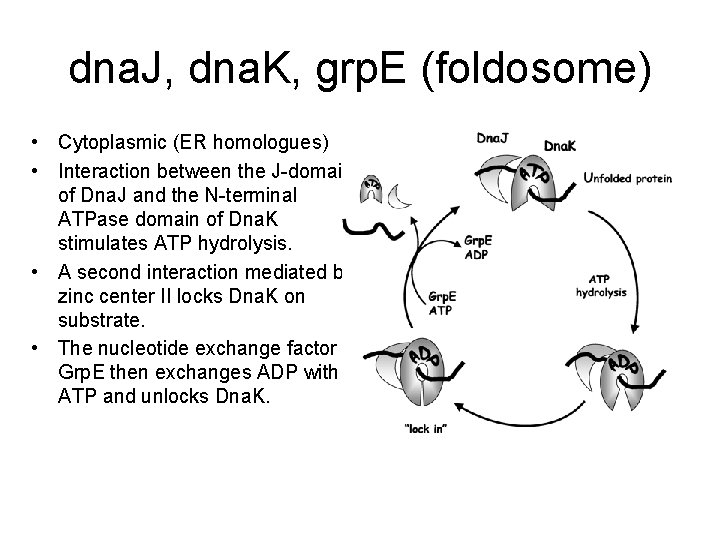
dna. J, dna. K, grp. E (foldosome) • Cytoplasmic (ER homologues) • Interaction between the J-domain of Dna. J and the N-terminal ATPase domain of Dna. K stimulates ATP hydrolysis. • A second interaction mediated by zinc center II locks Dna. K on substrate. • The nucleotide exchange factor Grp. E then exchanges ADP with ATP and unlocks Dna. K.

Chaperones

Chaperonin complex Gro. ES/Gro. EL Journal of Biochemistry 2005 137(5): 543 -549
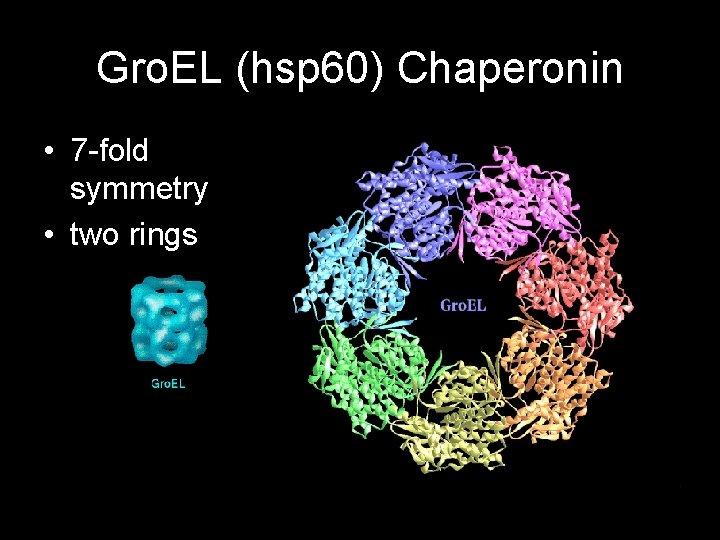
Gro. EL (hsp 60) Chaperonin • 7 -fold symmetry • two rings
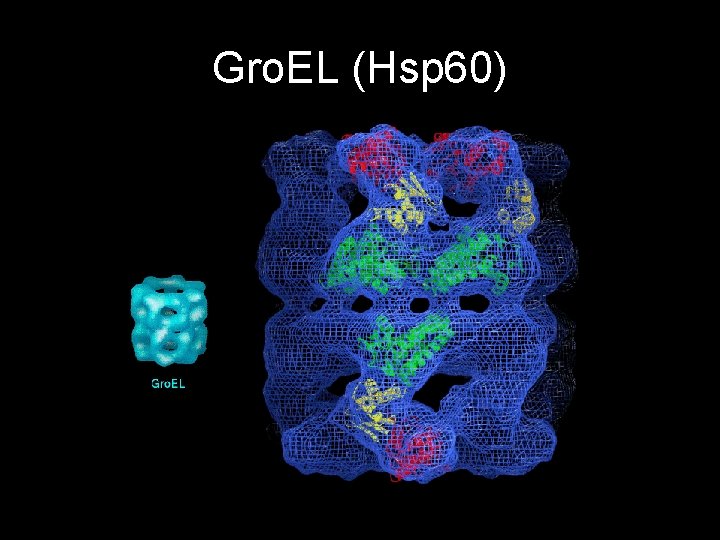
Gro. EL (Hsp 60)
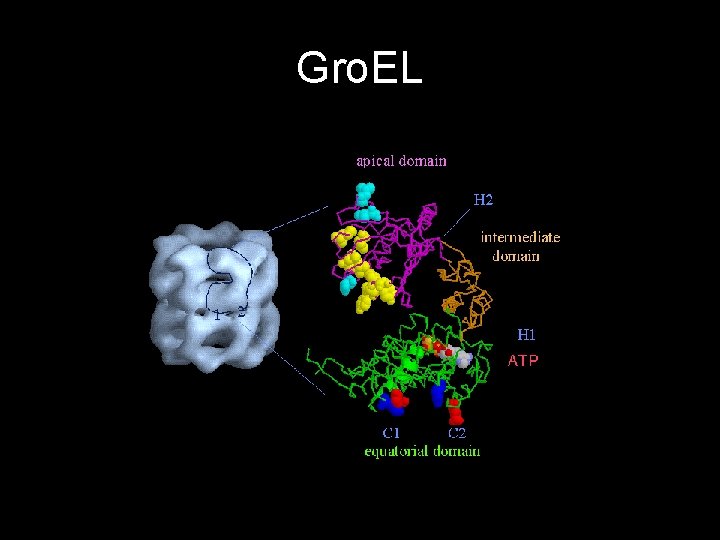
Gro. EL
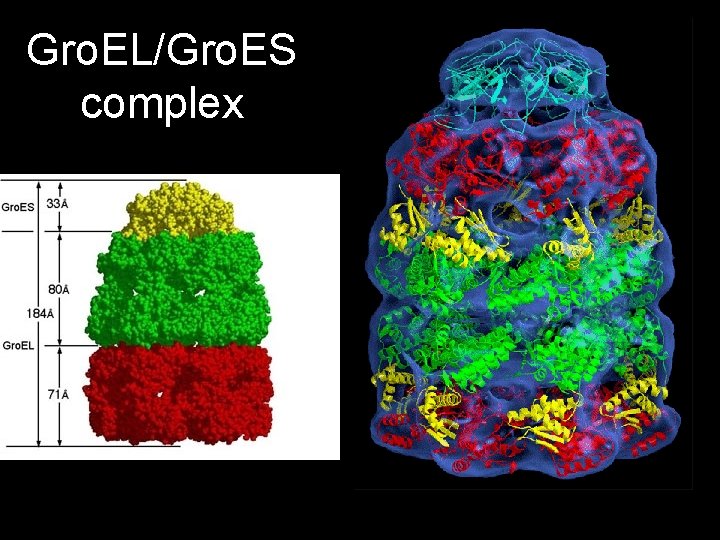
Gro. EL/Gro. ES complex

Gro. El/Gro. ES complex • Free Gro. EL binds denatured proteins very tightly. • Gro. EL undergoes an allosteric transition from its tightbinding state to a weaker binding state on the cooperative binding of nucleotides (ATP/ADP) and Gro. ES. • Gro. ES modulates binding of Gro. EL • 7 ATP molecules bind to Gro. EL • ATP hydrolysis drives transition back to tight binding state

Folding cage/Iterative annealing • Denatured proteins bind to the resting Gro. EL·Gro. ES·nucleotide complex. • Fast-folding proteins are ejected as native structures before ATP hydrolysis. • Slow-folding proteins enter chaperoning cycles of annealing and folding after the initial ATP hydrolysis. • Hydrolysis causes transient release of Gro. ES and formation of the Gro. EL·denatured-protein complexes with higher annealing potential. • The intermediately fast-folding complex is formed on subsequent rebinding of Gro. ES. • The ATPase activity of Gro. EL·Gro. ES is thus the gatekeeper that selects for initial entry of slow-folding proteins to the chaperone action and then pumps successive transitions from the faster-folding R-states to the tighter-binding/stronger annealing T-states.

Percolator model What happens when an Anfinsen-cage chaperone pulls its target? • The initial volume of the cis cavity (assumed to be 85000 A 3) • Water excluded from chamber on protein target • When ATP binds to the cis ring and induces a large conformational change increasing the cavity volume to 175000 A 3 • Water moves from the outside (organized structure) to the inside of chaperone and inside of target protein • Influx of water - resolvation of protein • 10 to 15 kcal/mol - provided by the conformational changes driven by ATP-hydrolysis?
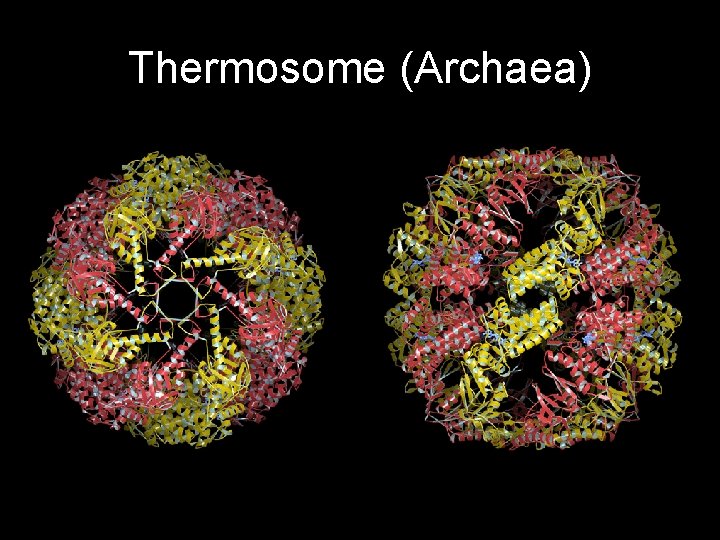
Thermosome (Archaea)
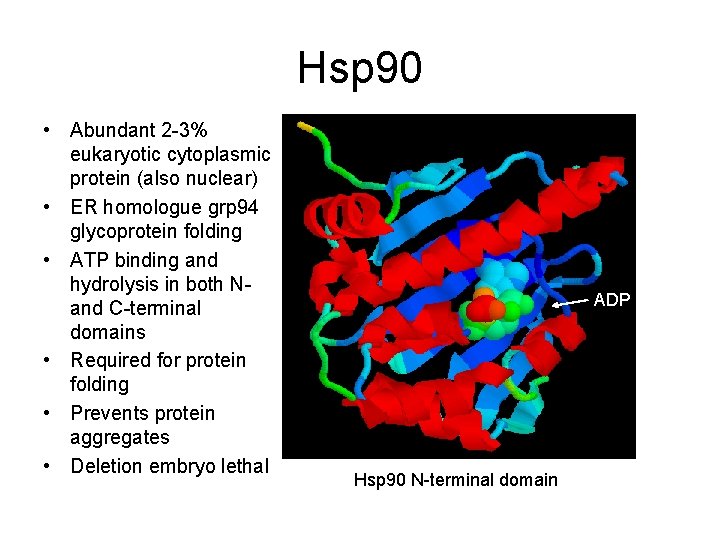
Hsp 90 • Abundant 2 -3% eukaryotic cytoplasmic protein (also nuclear) • ER homologue grp 94 glycoprotein folding • ATP binding and hydrolysis in both Nand C-terminal domains • Required for protein folding • Prevents protein aggregates • Deletion embryo lethal ADP Hsp 90 N-terminal domain
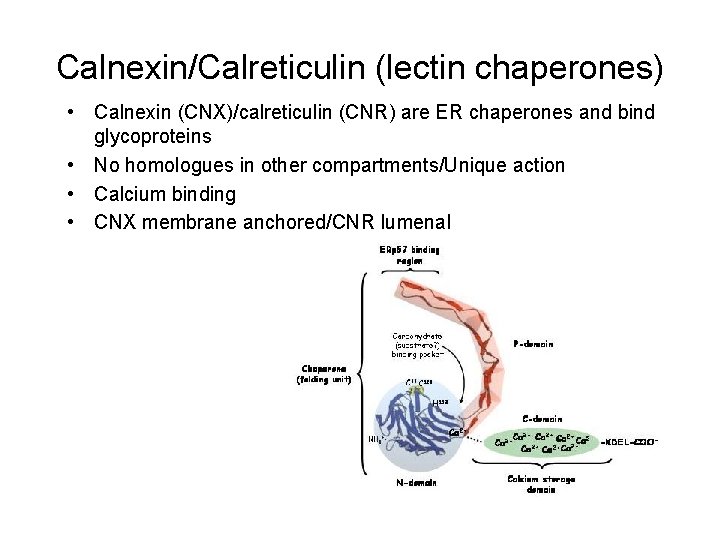
Calnexin/Calreticulin (lectin chaperones) • Calnexin (CNX)/calreticulin (CNR) are ER chaperones and bind glycoproteins • No homologues in other compartments/Unique action • Calcium binding • CNX membrane anchored/CNR lumenal
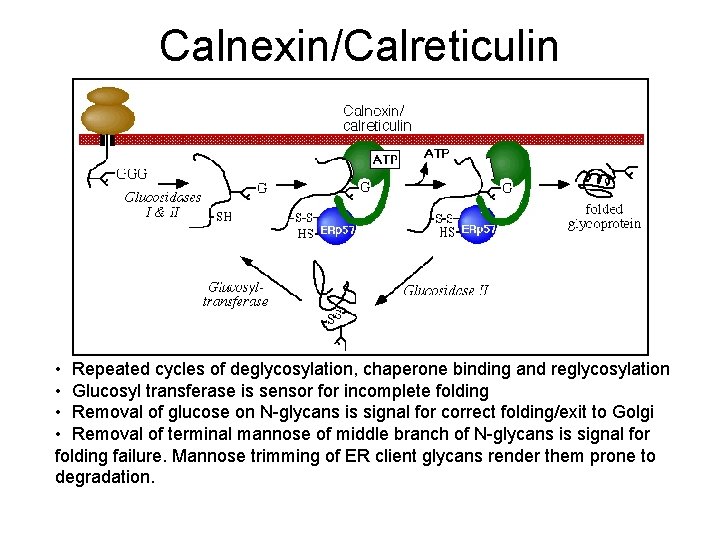
Calnexin/Calreticulin • Repeated cycles of deglycosylation, chaperone binding and reglycosylation • Glucosyl transferase is sensor for incomplete folding • Removal of glucose on N-glycans is signal for correct folding/exit to Golgi • Removal of terminal mannose of middle branch of N-glycans is signal for folding failure. Mannose trimming of ER client glycans render them prone to degradation.
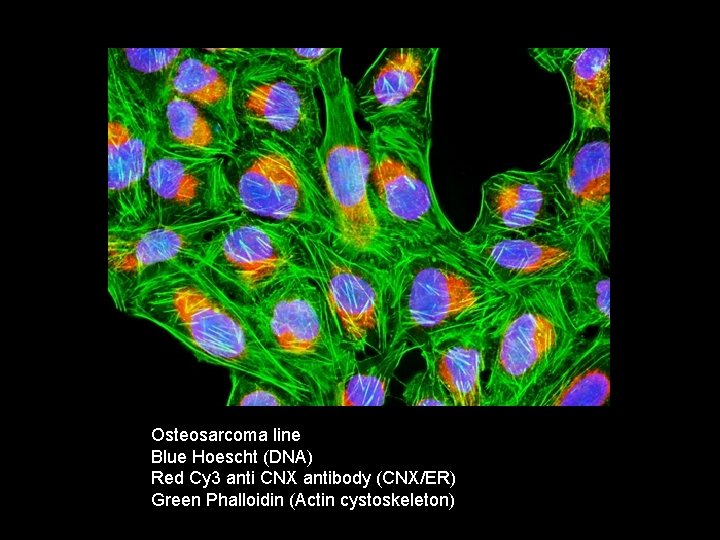
Osteosarcoma line Blue Hoescht (DNA) Red Cy 3 anti CNX antibody (CNX/ER) Green Phalloidin (Actin cystoskeleton)
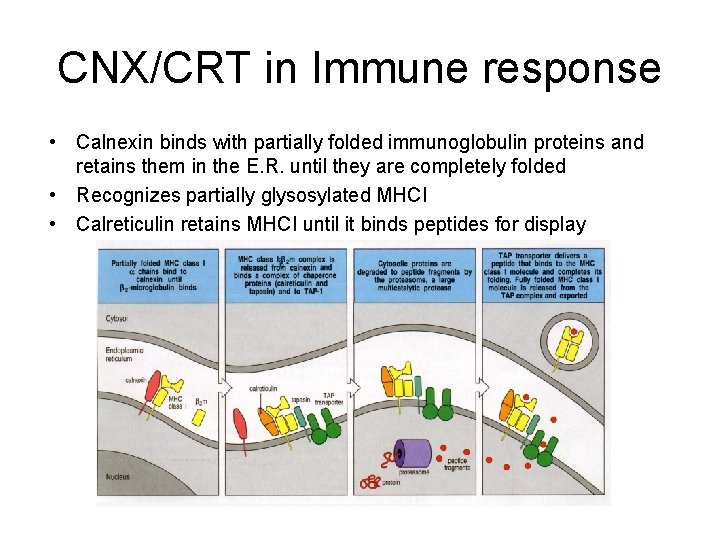
CNX/CRT in Immune response • Calnexin binds with partially folded immunoglobulin proteins and retains them in the E. R. until they are completely folded • Recognizes partially glysosylated MHCI • Calreticulin retains MHCI until it binds peptides for display
- Slides: 36