PROTEIN FOLDING Basics of proteins Protein folding is

PROTEIN FOLDING

Basics of proteins • Protein folding is the process by which a protein assumes its functional shape or minimum energy conformation. • Conformation generally means structural arrangement.

Conformations of Protein • Primary : Linear structure of Amino acids • Secondary: the twisting of polypeptides to form a helices, b sheets turns. • Tertiary : twisting of a helices, b sheets to form a larger structure due to disulphide bond formation and ionic bond formation.
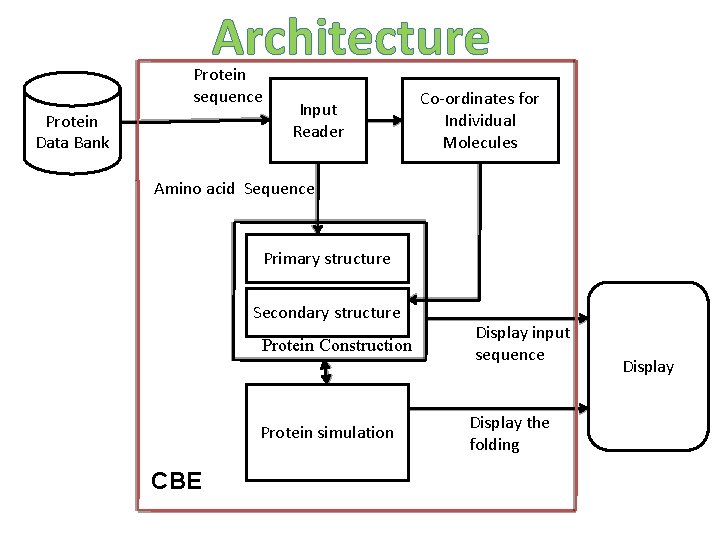
Architecture Protein sequence Protein Data Bank Input Reader Co-ordinates for Individual Molecules Amino acid Sequence Primary structure Secondary structure Protein Construction Protein simulation CBE Display input sequence Display the folding Display
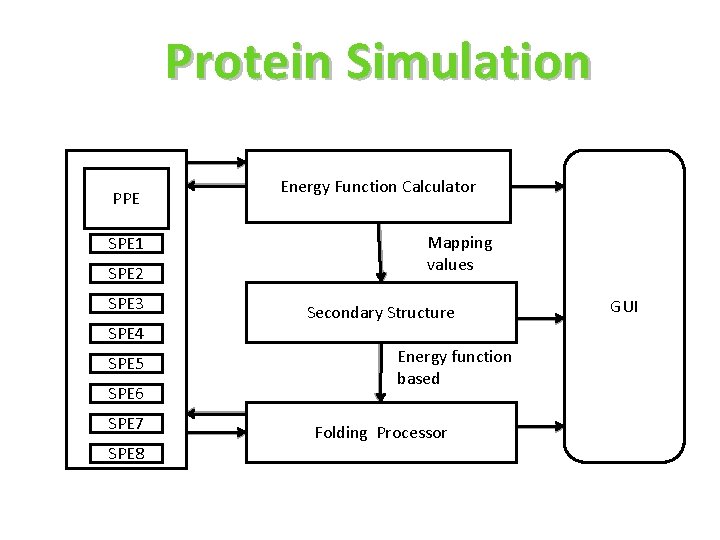
Protein Simulation PPE SPE 1 SPE 2 SPE 3 SPE 4 SPE 5 SPE 6 SPE 7 SPE 8 Energy Function Calculator Mapping values Secondary Structure Energy function based Folding Processor GUI

Current Implementation • A Cube grid was developed to act as a base to hold the amino acid molecules using Open. GL in Linux on x 86 to be represented in future. • The coordinates from the PDB files were extracted. • Designed the amino acid molecule representation in Open. GL. • Placement of these molecules on the grid. References: 1. Open. GL man pages: http: //www. opengl. org/documentation/specs/man_pages/hardcopy/GL/html/glu/ 2. Py. Open. GL man pages: http: //pyopengl. sourceforge. net/documentation/manual/

Challenges • Calculating the energy function. • Mapping the energy values on the secondary structure to show the folding. • Calculating the movements of the protein structure. • Implementation on CBE and parallel execution of instructions. • Display of folded 3 D structure. • Implementation of Activity list. [1] 1. Source: Activity List http: //tinyurl. com/9 aztkq
Input Processor
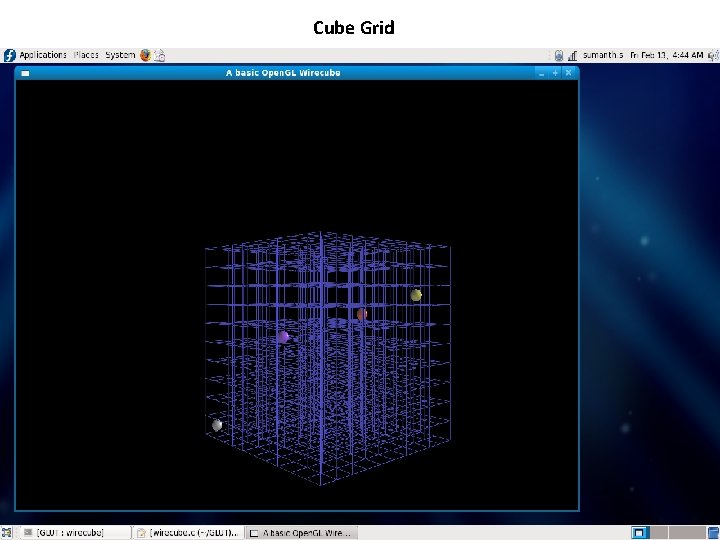
Cube Grid

THANK YOU!. .
- Slides: 10