Hidden Markov Models Slides by Bill Majoros based

Hidden Markov Models Slides by Bill Majoros, based on Methods for Computational Gene Prediction

What is an HMM? An HMM is a stochastic machine M=(Q, , Pt, Pe) consisting of the following: • a finite set of states, Q={q 0, q 1, . . . , qm} • a finite alphabet ={s 0, s 1, . . . , sn} • a transition distribution Pt : Q×Q a¡ • an emission distribution Pe : Q× a¡ i. e. , Pt (qj | qi) i. e. , Pe (sj | qi) An Example 5% M 1=({q 0, q 1, q 2}, {Y, R}, Pt, Pe) Pt={(q 0, q 1, 1), (q 1, 0. 8), (q 1, q 2, 0. 15), (q 1, q 0, 0. 05), (q 2, 0. 7), (q 2, q 1, 0. 3)} q 0 Pe={(q 1, Y, 1), (q 1, R, 0), (q 2, Y, 0), (q 2, R, 1)} 80% 15% Y=0% R = 100% Y = 100% q 1 100% 30% q 2 70%
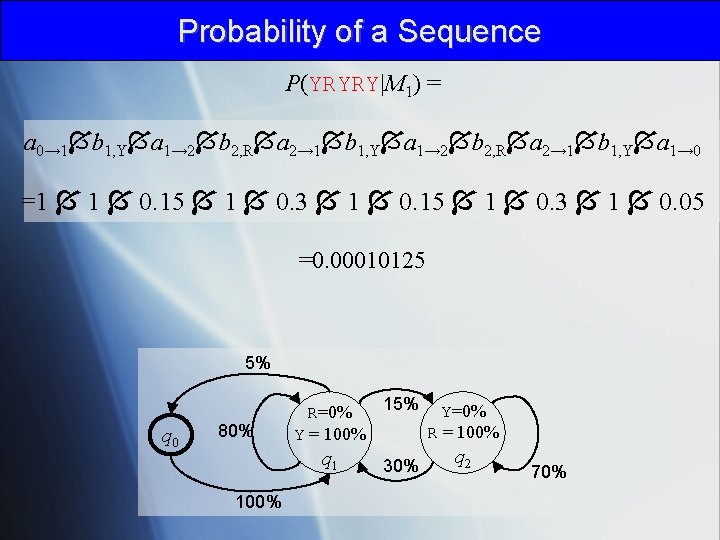
Probability of a Sequence P(YRYRY|M 1) = a 0→ 1 b 1, Y a 1→ 2 b 2, R a 2→ 1 b 1, Y a 1→ 0 =1 1 0. 15 1 0. 3 1 0. 05 =0. 00010125 5% q 0 80% 15% Y=0% R = 100% Y = 100% q 1 100% 30% q 2 70%
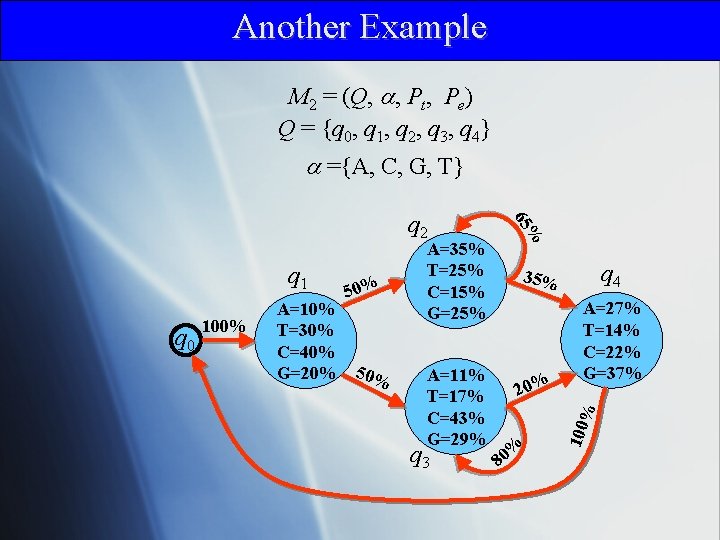
Another Example M 2 = (Q, , Pt, Pe) Q = {q 0, q 1, q 2, q 3, q 4} ={A, C, G, T} A=11% T=17% C=43% G=29% 20% q 3 A=27% T=14% C=22% G=37% % A=10% T=30% C=40% G=20% 50% q 4 35% % 80 100 q 0 100% 50% A=35% T=25% C=15% G=25% % q 1 65 q 2
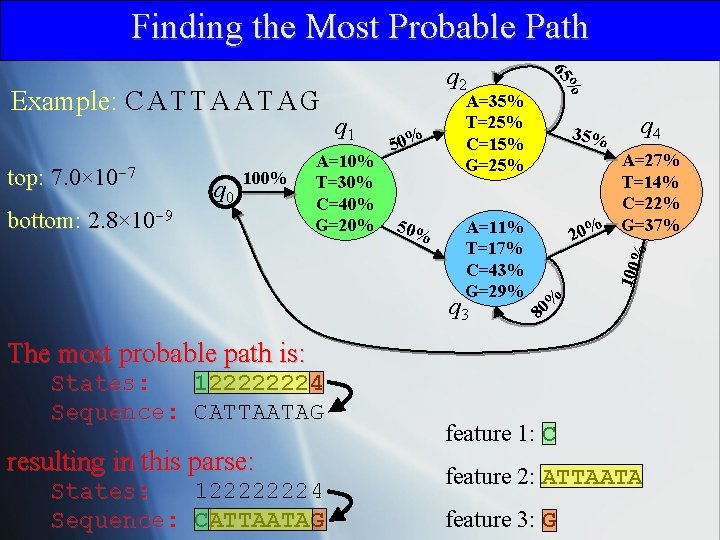
Finding the Most Probable Path 100% bottom: 2. 8× 10 -9 50% A=11% T=17% C=43% G=29% q 3 35% q 4 A=27% T=14% C=22% G=37% 20% % q 0 A=10% T=30% C=40% G=20% 50% 100 top: 7. 0× 10 -7 q 1 A=35% T=25% C=15% G=25% % 65 Example: C A T T A A T A G q 2 % 80 The most probable path is: States: 122222224 Sequence: CATTAATAG resulting in this parse: States: 122222224 Sequence: CATTAATAG feature 1: C feature 2: ATTAATA feature 3: G
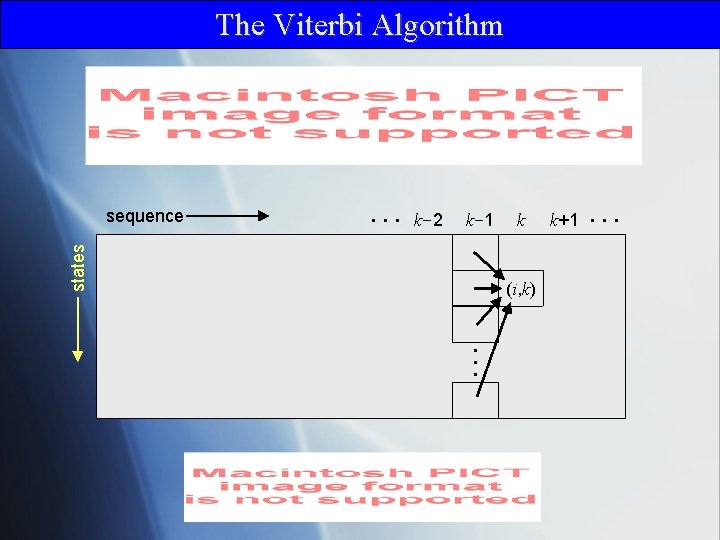
The Viterbi Algorithm . . . k-2 k-1 states sequence k (i, k) k+1 . . .
Viterbi: Traceback T( T( T(. . . T( T(i, L-1), L-2). . . , 2), 1), 0) = 0
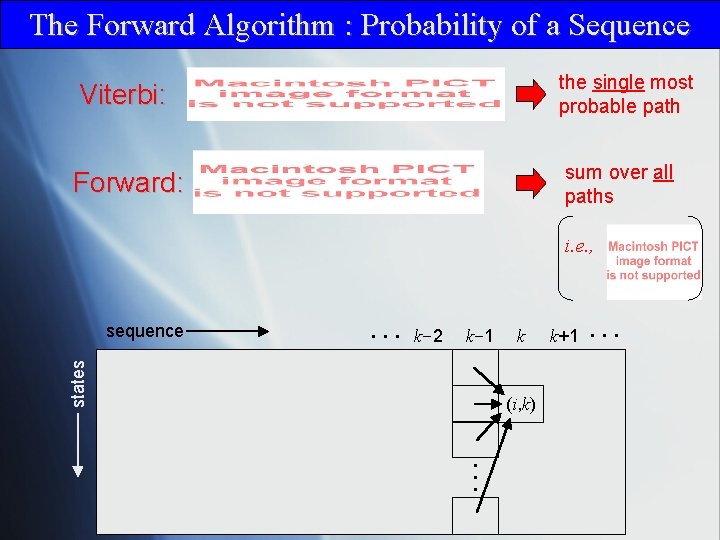
The Forward Algorithm : Probability of a Sequence Viterbi: the single most probable path Forward: sum over all paths i. e. , . . . k-2 k-1 states sequence k (i, k) k+1 . . .

Training an HMM from Labeled Sequences transitions CGATATTCGATTCTACGCGCGTATACTAGCTTATCTGATC 01111111222222211111112222111110 to state 0 from state 1 2 0 0 (0%) 1 (100%) 0 (0%) 1 1 (4%) 21 (84%) 3 (12%) 2 0 (0%) 3 (20%) 12 (80%) emissions symbol A C G T in 1 state 6 (24%) 7 (28%) 5 (20%) 7 (28%) 2 3 (20%) 2 (13%) 7 (47%) From the textbook, Ch. 6. 3
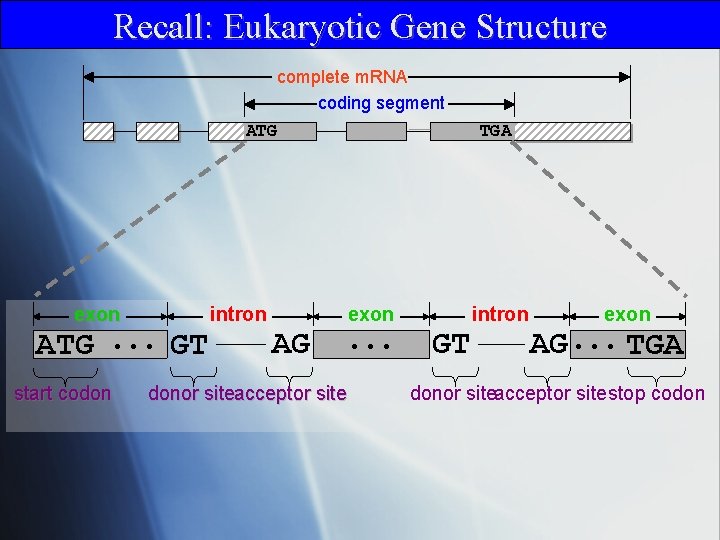
Recall: Eukaryotic Gene Structure complete m. RNA coding segment ATG exon ATG. . . GT start codon intron TGA exon AG donor siteacceptor site . . . intron GT exon AG. . . TGA donor siteacceptor sitestop codon
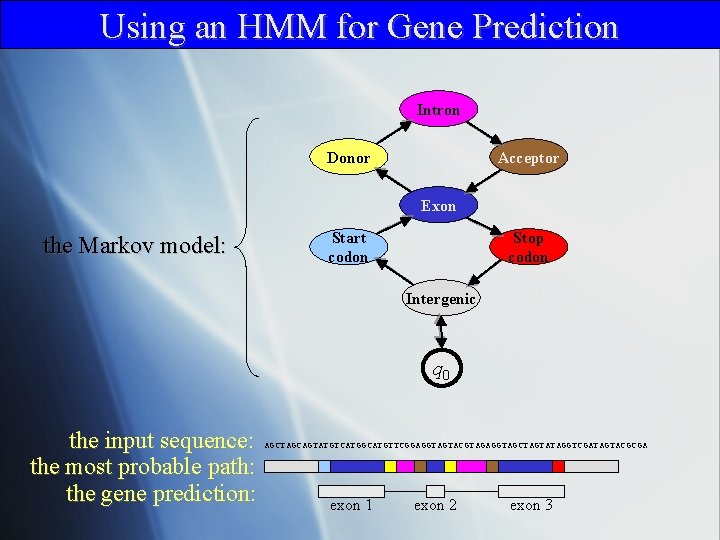
Using an HMM for Gene Prediction Intron Donor Acceptor Exon the Markov model: Start codon Stop codon Intergenic q 0 the input sequence: the most probable path: the gene prediction: AGCTAGCAGTATGTCATGGCATGTTCGGAGGTAGTACGTAGAGGTAGCTAGTATAGGTCGATAGTACGCGA exon 1 exon 2 exon 3
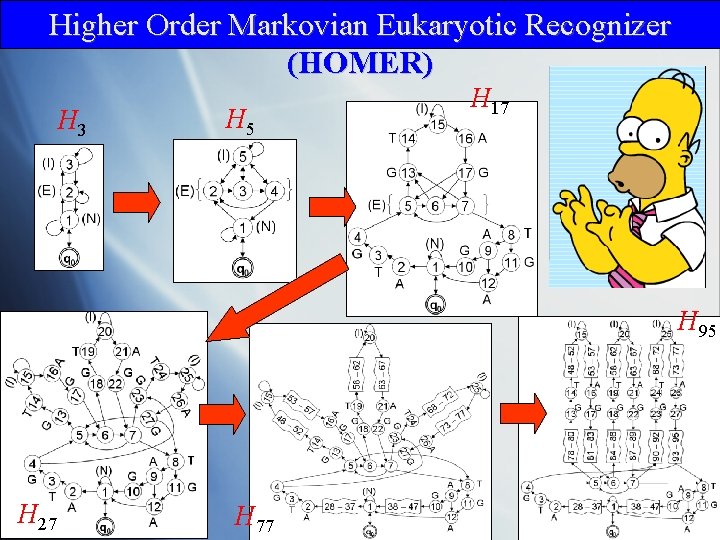
Higher Order Markovian Eukaryotic Recognizer (HOMER) H 3 H 5 H 17 H 95 H 27 H 77
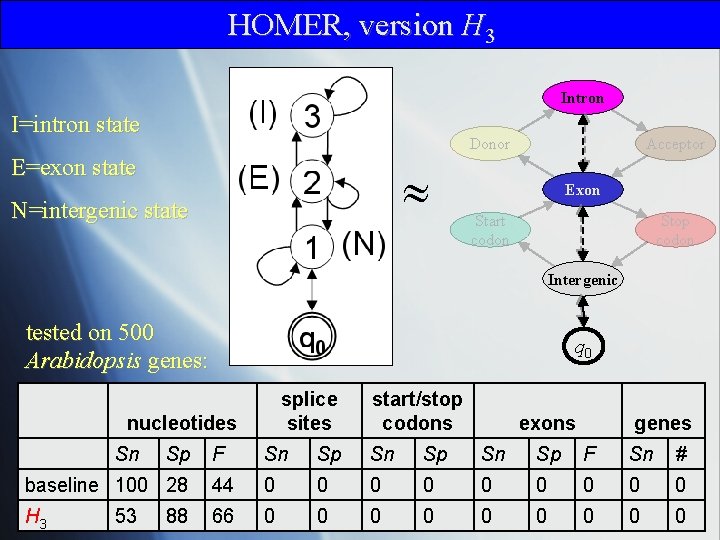
HOMER, version H 3 Intron I=intron state Donor E=exon state N=intergenic state Acceptor Exon Start codon Stop codon Intergenic tested on 500 Arabidopsis genes: q 0 splice sites nucleotides Sn exons genes F Sn Sp F Sn # baseline 100 28 44 0 0 0 0 0 H 3 66 0 0 0 0 0 53 Sp start/stop codons 88

Recall: Sensitivity and Precision NOTE: “specificity” as defined here and throughout these slides (and the text) is really precision
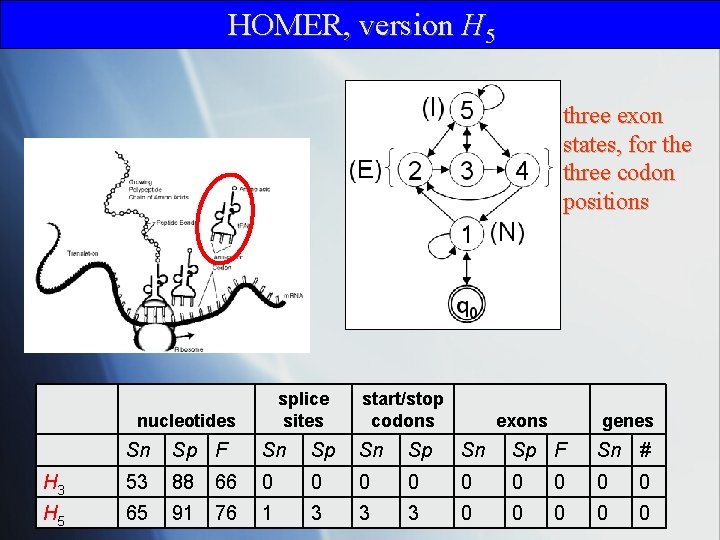
HOMER, version H 5 three exon states, for the three codon positions splice sites nucleotides start/stop codons exons genes Sn Sp F Sn Sp F Sn # H 3 53 88 66 0 0 0 0 0 H 5 65 91 76 1 3 3 3 0 0 0
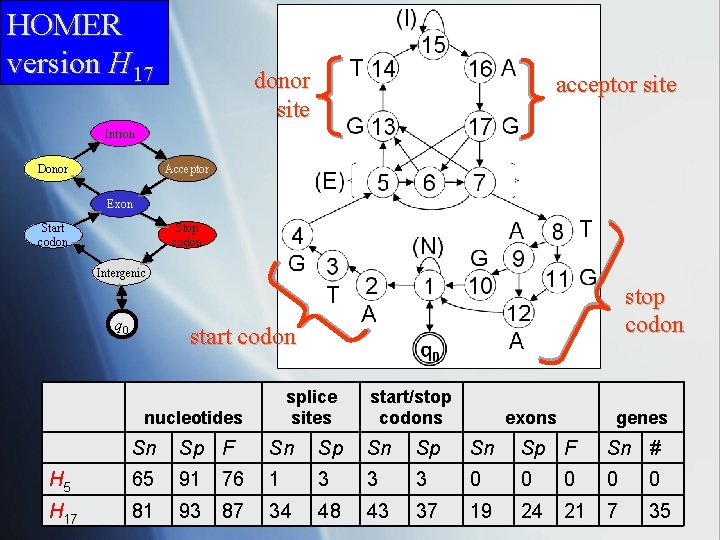
HOMER version H 17 donor site acceptor site Intron Donor Acceptor Exon Start codon Stop codon Intergenic q 0 stop codon start codon splice sites nucleotides start/stop codons exons genes Sn Sp F Sn Sp F Sn # H 5 65 91 76 1 3 3 3 0 0 0 H 17 81 93 87 34 48 43 37 19 24 21 7 35

Maintaining Phase Across an Intron phase: 012012012012 sequence: + GTATGCGATAGTCAAGAGTGATCGCTAGACC coordinates: | 0 | 5 | 10 | 15 | 20 | 25 | 30
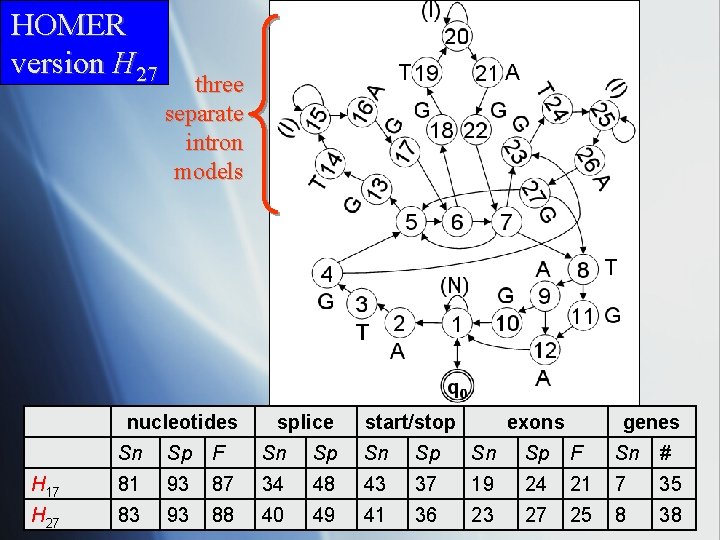
HOMER version H 27 three separate intron models nucleotides splice start/stop exons genes Sn Sp F Sn Sp F Sn # H 17 81 93 87 34 48 43 37 19 24 21 7 35 H 27 83 93 88 40 49 41 36 23 27 25 8 38
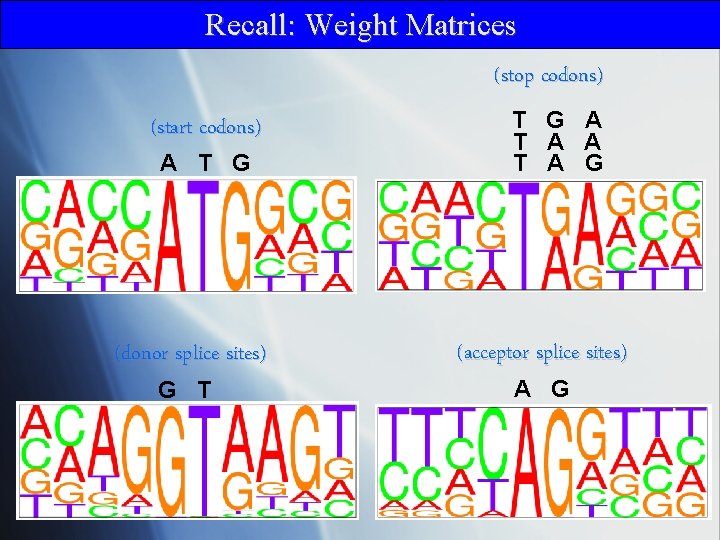
Recall: Weight Matrices (stop codons) (start codons) A T G (donor splice sites) G T T G A T A G (acceptor splice sites) A G
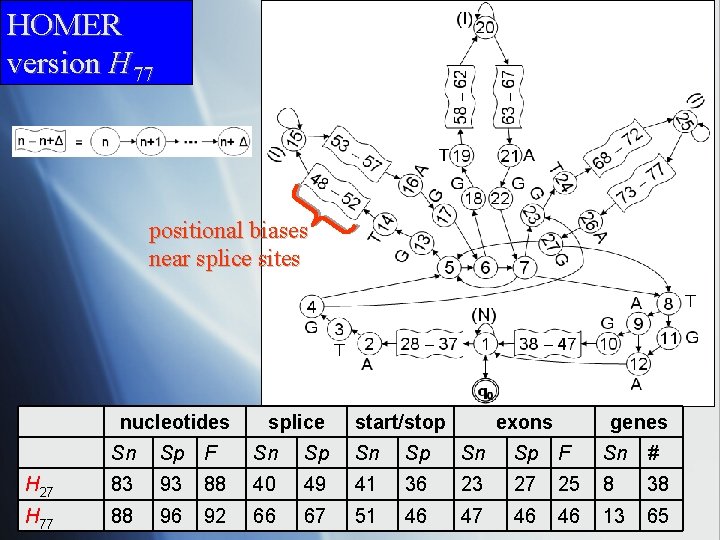
HOMER version H 77 positional biases near splice sites nucleotides splice start/stop exons genes Sn Sp F Sn Sp F Sn # H 27 83 93 88 40 49 41 36 23 27 25 8 38 H 77 88 96 92 66 67 51 46 47 46 46 13 65
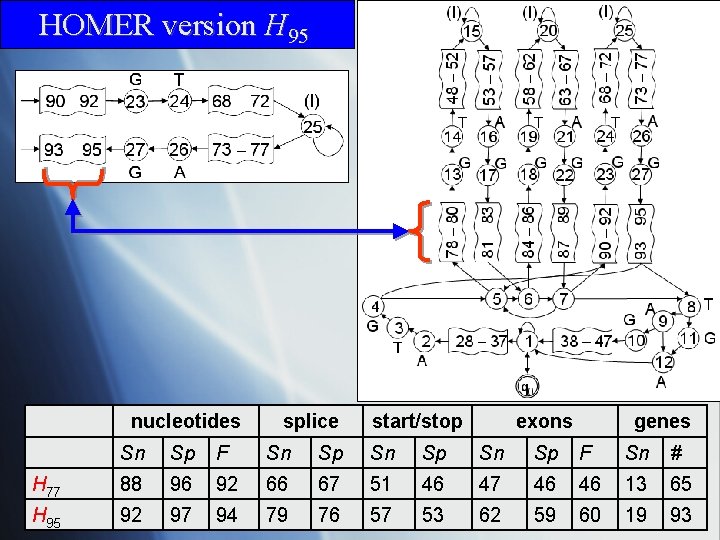
HOMER version H 95 nucleotides splice start/stop exons genes Sn Sp F Sn Sp F Sn # H 77 88 96 92 66 67 51 46 47 46 46 13 65 H 95 92 97 94 79 76 57 53 62 59 60 19 93

Summary of HOMER Results

Higher-order Markov Models P(G) 0 th order: ACGCTA P(G|C) 1 st order: ACGCTA P(G|AC) 2 nd order: ACGCTA
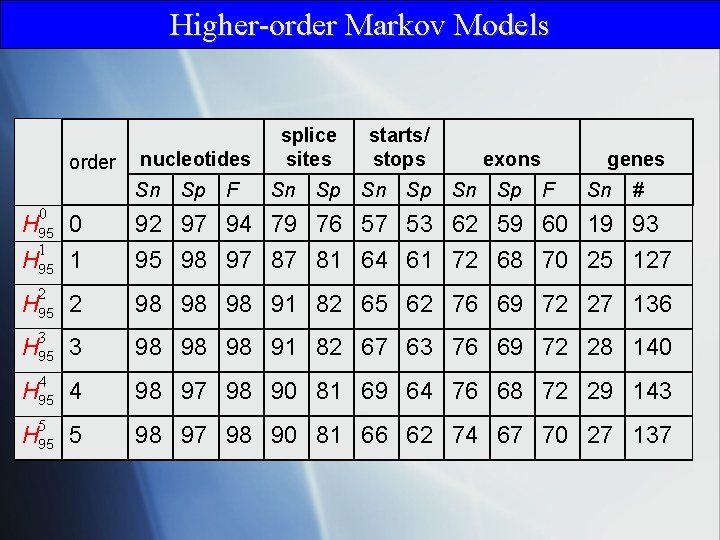
Higher-order Markov Models order nucleotides Sn Sp F splice sites Sn Sp starts/ stops Sn Sp exons Sn Sp genes F Sn # 0 92 97 94 79 76 57 53 62 59 60 19 93 95 98 97 87 81 64 61 72 68 70 25 127 2 98 98 98 91 82 65 62 76 69 72 27 136 3 98 98 98 91 82 67 63 76 69 72 28 140 4 98 97 98 90 81 69 64 76 68 72 29 143 5 98 97 98 90 81 66 62 74 67 70 27 137 H 95 0 1 H 95 2 H 95 3 H 95 4 H 95 5

Summary • An HMM is a stochastic generative model which emits sequences • Parsing with an HMM can be accomplished using a decoding algorithm (such as Viterbi) to find the most probable (MAP) path generating the input sequence • Training of unambiguous HMM’s can be accomplished using labeled sequence training • Training of ambiguous HMM’s can be accomplished using Viterbi training or the Baum-Welch algorithm • Posterior decoding can be used to estimate the probability that a given symbol or substring was generate by a particular state (next lesson. . . )
- Slides: 25