Bioinformatics for Stem Cell Lecture 2 Debashis Sahoo

Bioinformatics for Stem Cell Lecture 2 Debashis Sahoo, Ph. D

Outline • • • Lecture 1 Recap Multivariate analysis Microarray data analysis Boolean analysis Sequencing data analysis

MULTIVARIATE ANALYSIS
Identify Markers of Human Colon Cancer and Normal Colon Piero Dalerba Tomer Kalisky 4
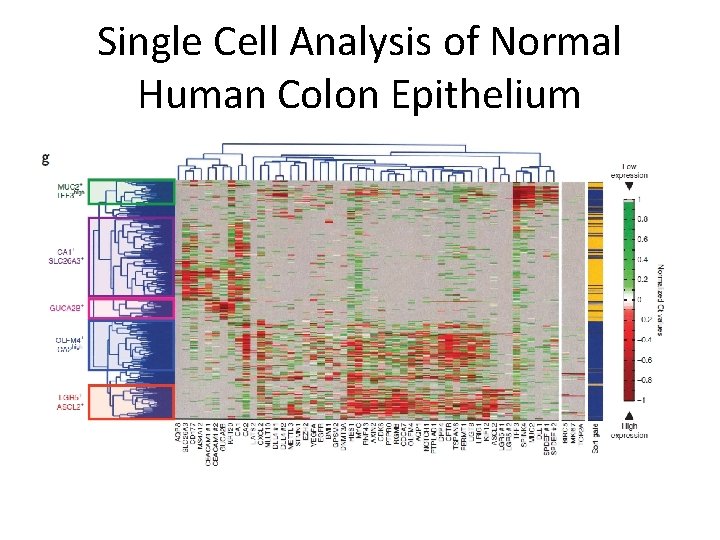
Single Cell Analysis of Normal Human Colon Epithelium
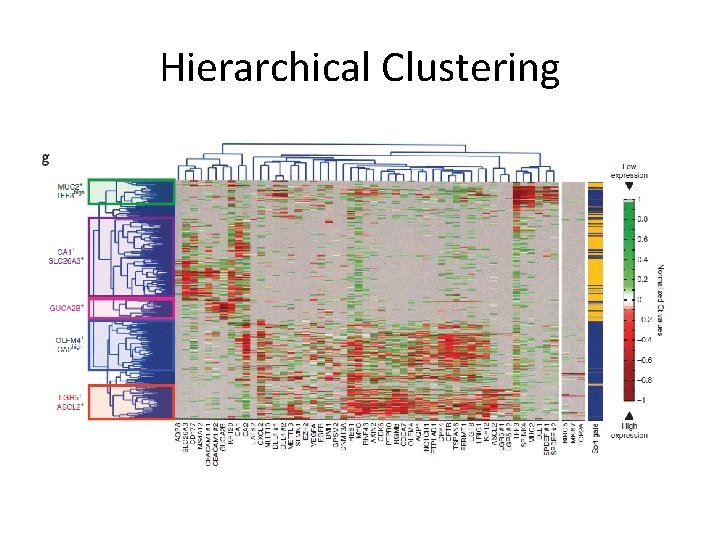
Hierarchical Clustering

Hierarchical Clustering • Cluster 3. 0 – http: //bonsai. hgc. jp/~mdehoon/software/cluster/ • Distance metric – Euclidian, Squared Euclidean, Manhattan, maximum, cosine, Pearson’s correlation • Linkage – Single, complete, average, median, centroid
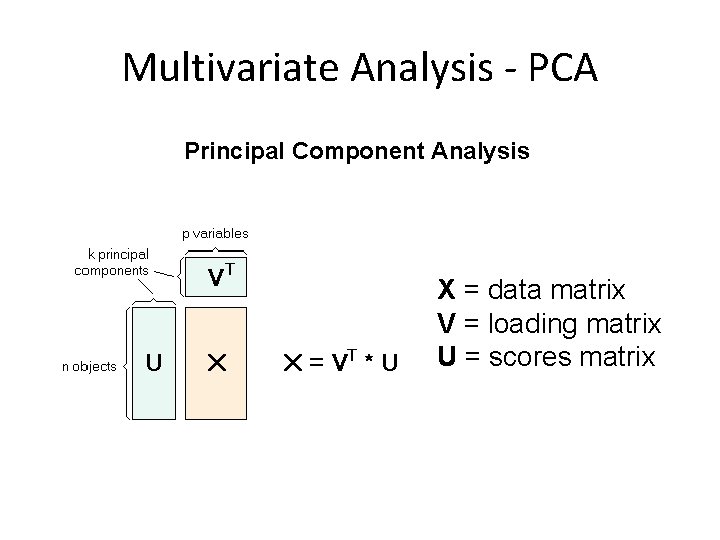
Multivariate Analysis - PCA Principal Component Analysis X = data matrix V = loading matrix U = scores matrix
Fundamentals of PCA • Reduces dimensions of the data • PCA uses orthogonal linear transformation • First principal component has the largest possible variance. • Exploratory tool to uncover unknown trends in the data
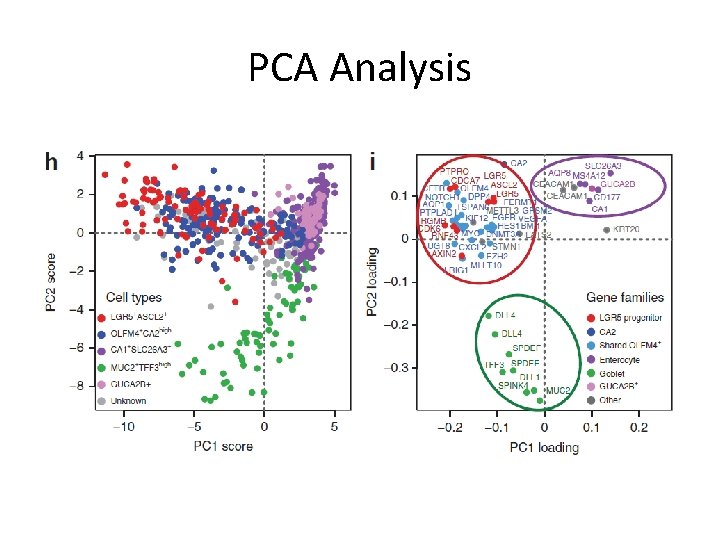
PCA Analysis

HIGH-THROUGHPUT DATA ANALYSIS

MICROARRAY ANALYSIS
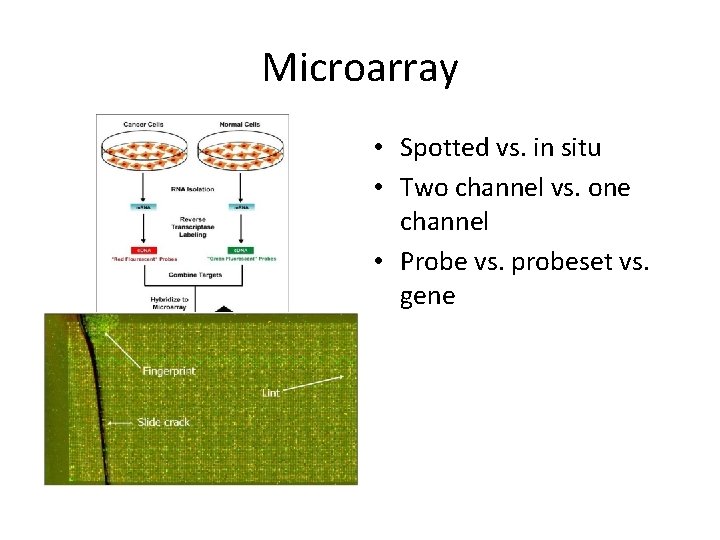
Microarray • Spotted vs. in situ • Two channel vs. one channel • Probe vs. probeset vs. gene
![Quantile Normalization Sort #1 #2 #3 Sorted. Avg Average Val(Probe_i) = Sorted. Avg[Rank(Probe_i)] Quantile Normalization Sort #1 #2 #3 Sorted. Avg Average Val(Probe_i) = Sorted. Avg[Rank(Probe_i)]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-14.jpg)
Quantile Normalization Sort #1 #2 #3 Sorted. Avg Average Val(Probe_i) = Sorted. Avg[Rank(Probe_i)]
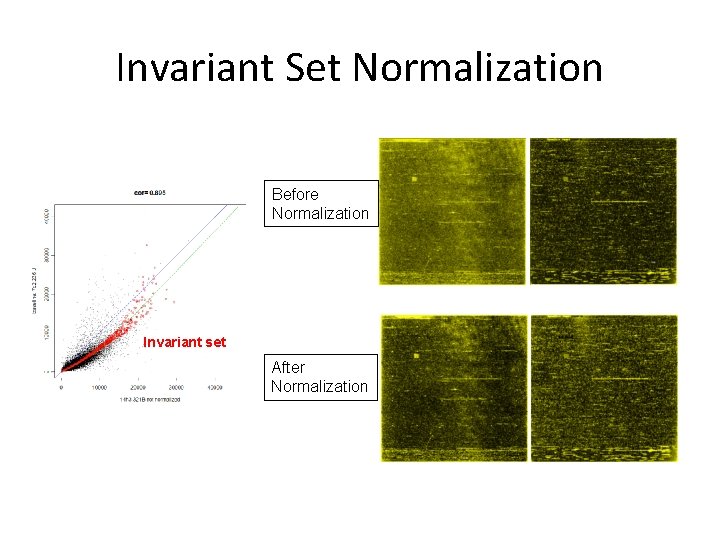
Invariant Set Normalization Before Normalization Invariant set After Normalization

Good to Check the Image
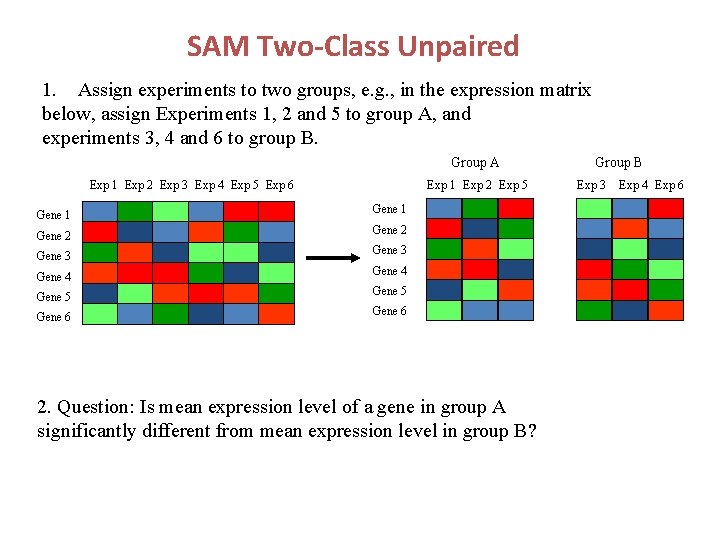
SAM Two-Class Unpaired 1. Assign experiments to two groups, e. g. , in the expression matrix below, assign Experiments 1, 2 and 5 to group A, and experiments 3, 4 and 6 to group B. Group A Exp 1 Exp 2 Exp 3 Exp 4 Exp 5 Exp 6 Gene 1 Gene 2 Gene 3 Exp 1 Exp 2 Exp 5 Gene 1 Gene 2 Gene 3 Gene 4 Gene 5 Gene 6 2. Question: Is mean expression level of a gene in group A significantly different from mean expression level in group B? Group B Exp 3 Exp 4 Exp 6

SAM Two-Class Unpaired Permutation tests i) For each gene, compute d-value (analogous to t-statistic). This is the observed d-value for that gene. ii) Rank the genes in ascending order of their d-values. iii) Randomly shuffle the values of the genes between groups A and B, such that the reshuffled groups A and B respectively have the same number of elements as the original groups A and B. Compute the d-value for each randomized gene Group A Group B Exp 1 Exp 2 Exp 5 Exp 3 Exp 4 Exp 6 Original grouping Gene 1 Group A Exp 3 Exp 2 Gene 1 Group B Exp 6 Exp 4 Exp 5 Exp 1 Randomized grouping

SAM Two-Class Unpaired iv) Rank the permuted d-values of the genes in ascending order v) Repeat steps iii) and iv) many times, so that each gene has many randomized d-values corresponding to its rank from the observed (unpermuted) d-value. Take the average of the randomized d-values for each gene. This is the expected d-value of that gene. vi) Plot the observed d-values vs. the expected d-values
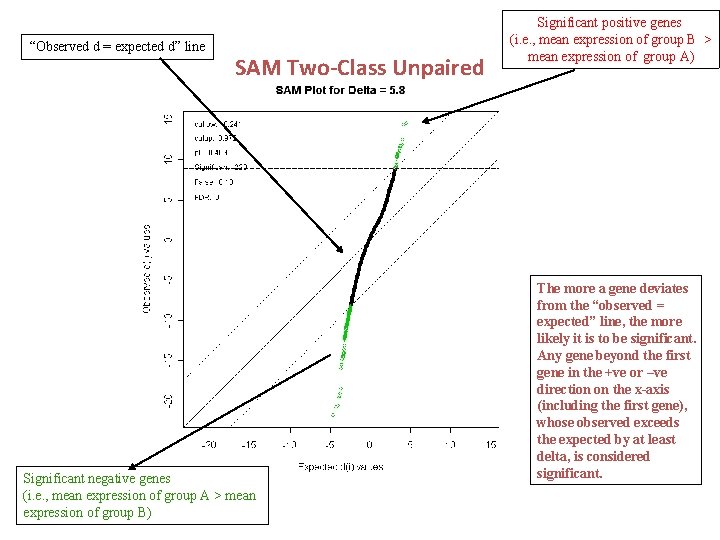
“Observed d = expected d” line SAM Two-Class Unpaired Significant negative genes (i. e. , mean expression of group A > mean expression of group B) Significant positive genes (i. e. , mean expression of group B > mean expression of group A) The more a gene deviates from the “observed = expected” line, the more likely it is to be significant. Any gene beyond the first gene in the +ve or –ve direction on the x-axis (including the first gene), whose observed exceeds the expected by at least delta, is considered significant.
Gene. Pattern http: //genepattern. broadinstitute. org/

Auto. SOME http: //jimcooperlab. mcdb. ucsb. edu/autosome/ Aaron Newman and James Cooper, BMC Bioinformatics, 2010, 11: 117
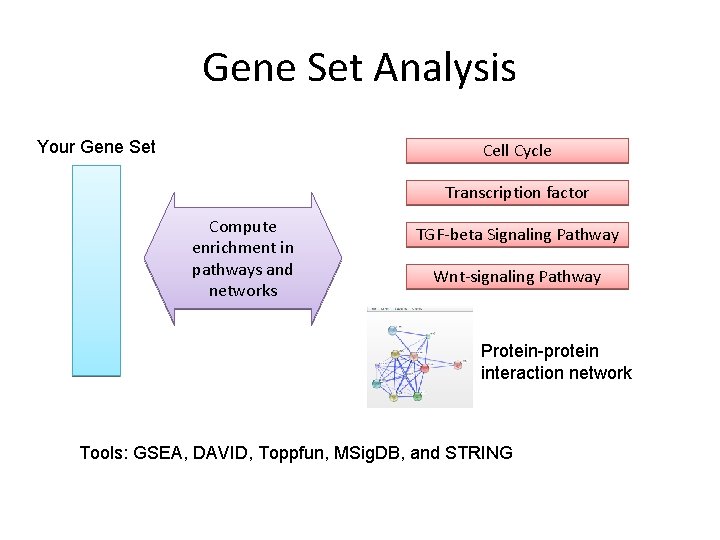
Gene Set Analysis Your Gene Set Cell Cycle Transcription factor Compute enrichment in pathways and networks TGF-beta Signaling Pathway Wnt-signaling Pathway Protein-protein interaction network Tools: GSEA, DAVID, Toppfun, MSig. DB, and STRING

BOOLEAN ANALYSIS
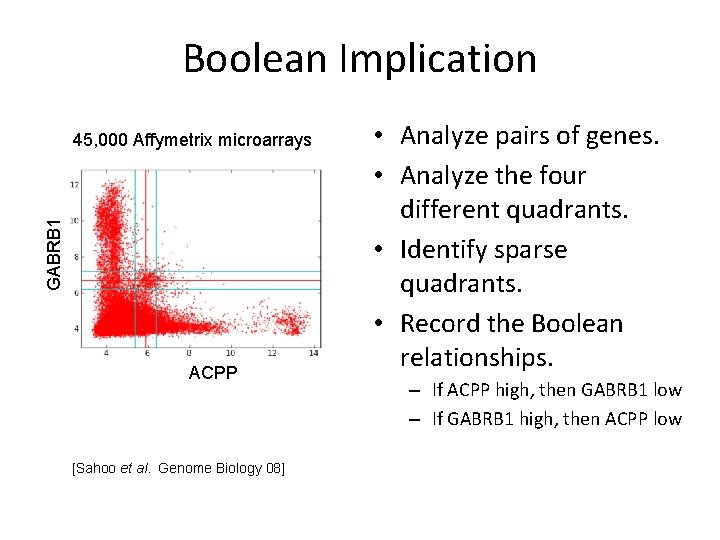
Boolean Implication GABRB 1 45, 000 Affymetrix microarrays ACPP [Sahoo et al. Genome Biology 08] • Analyze pairs of genes. • Analyze the four different quadrants. • Identify sparse quadrants. • Record the Boolean relationships. – If ACPP high, then GABRB 1 low – If GABRB 1 high, then ACPP low
![CDH expression Threshold Calculation High Intermediate Low Threshold Sorted arrays [Sahoo et al. 07] CDH expression Threshold Calculation High Intermediate Low Threshold Sorted arrays [Sahoo et al. 07]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-26.jpg)
CDH expression Threshold Calculation High Intermediate Low Threshold Sorted arrays [Sahoo et al. 07] • A threshold is determined for each gene. • The arrays are sorted by gene expression • Step. Miner is used to determine threshold

Boolean. Net Statistics B a 01 a 11 a 00 a 10 n. Alow = (a 00+ a 01), n. Blow = (a 00+ a 10) total = a 00+ a 01+ a 10+ a 11, observed = a 00 expected = (n. Alow/ total * n. Blow/ total) * total statistic = A error rate = 1 2 ((a a 00 00+ a 01) (expected – observed) √ expected + a 00 (a 00+ a 10) ) Boolean Implication = (statistic > 3, error rate < 0. 1) [Sahoo et al. Genome Biology 08]
![Six Boolean Implications [Sahoo et al. Genome Biology 08] Six Boolean Implications [Sahoo et al. Genome Biology 08]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-28.jpg)
Six Boolean Implications [Sahoo et al. Genome Biology 08]
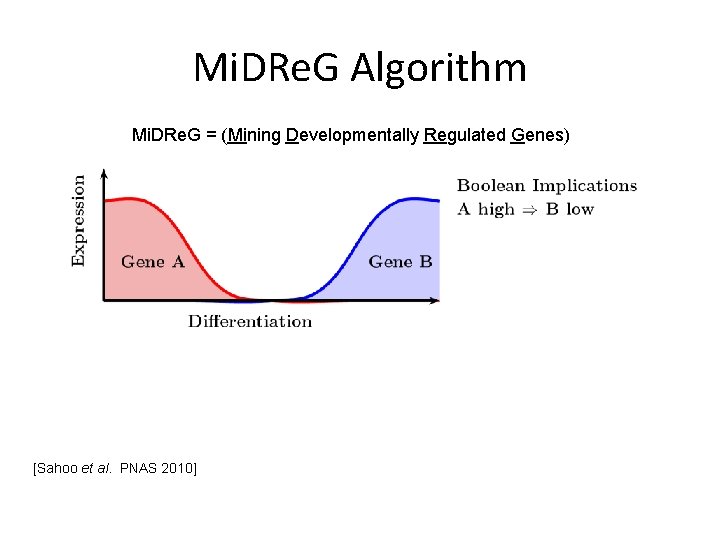
Mi. DRe. G Algorithm Mi. DRe. G = (Mining Developmentally Regulated Genes) [Sahoo et al. PNAS 2010]
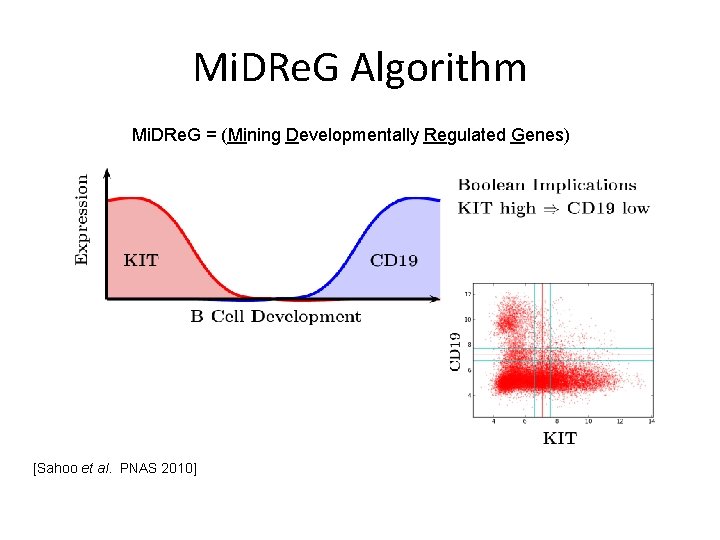
Mi. DRe. G Algorithm Mi. DRe. G = (Mining Developmentally Regulated Genes) [Sahoo et al. PNAS 2010]
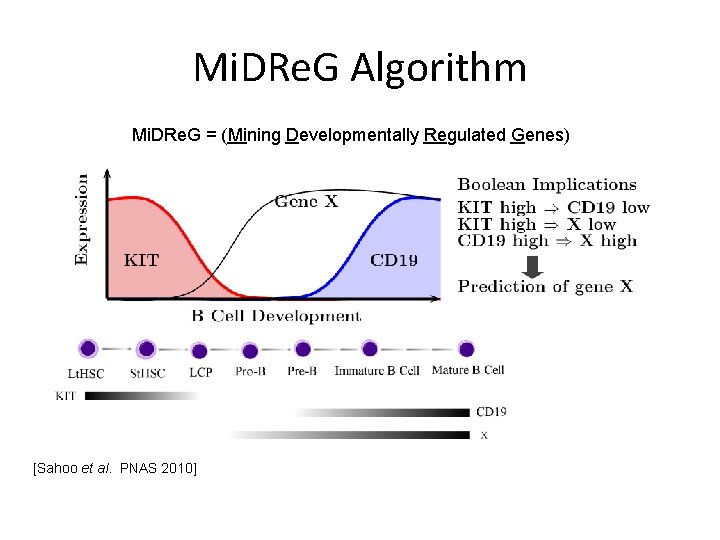
Mi. DRe. G Algorithm Mi. DRe. G = (Mining Developmentally Regulated Genes) [Sahoo et al. PNAS 2010]
![B Cell Genes KIT Boolean Implications CD 19 [Sahoo et al. PNAS 2010] B Cell Genes KIT Boolean Implications CD 19 [Sahoo et al. PNAS 2010]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-32.jpg)
B Cell Genes KIT Boolean Implications CD 19 [Sahoo et al. PNAS 2010]
![Jun Seita http: //gexc. stanford. edu [Seita, Sahoo et al. PLo. S ONE, 2012] Jun Seita http: //gexc. stanford. edu [Seita, Sahoo et al. PLo. S ONE, 2012]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-33.jpg)
Jun Seita http: //gexc. stanford. edu [Seita, Sahoo et al. PLo. S ONE, 2012]

SEQUENCING DATA ANALYSIS

Sequencing Data Format >SEQUENCE_1 MTEITAAMVKELRESTGAGMMDCKNALSETNGDFDKAVQLLREKGLGKAAKKADRLAAEG LVSVKVSDDFTIAAMRPSYLSYEDLDMTFVENEYKALVAELEKENEERRRLKDPNKPEHK IPQFASRKQLSDAILKEAEEKIKEELKAQGKPEKIWDNIIPGKMNSFIADNSQLDSKLTL MGQFYVMDDKKTVEQVIAEKEKEFGGKIKIVEFICFEVGEGLEKKTEDFAAEVAAQL >SEQUENCE_2 SATVSEINSETDFVAKNDQFIALTKDTTAHIQSNSLQSVEELHSSTINGVKFEEYLKSQI ATIGENLVVRRFATLKAGANGVVNGYIHTNGRVGVVIAAACDSAEVASKSRDLLRQICMH FASTQ S X I J L - FASTA @HWI-EAS 209: 5: 5894: 21141#ATCACG/1 TTAATTGGTAAATCTCCTAATAGCTTAGATNT +HWI-EAS 209: 5: 5894: 21141#ATCACG/1 efcfffffcfeefffcffffffddf`feed]`]_Ba Sanger Phred+33, (0, 40) Solexa+64, (-5, 40) Illumina 1. 3+ Phred+64, (0, 40) Illumina 1. 5+ Phred+64, (3, 40) Illumina 1. 8+ Phred+33, (0, 41)
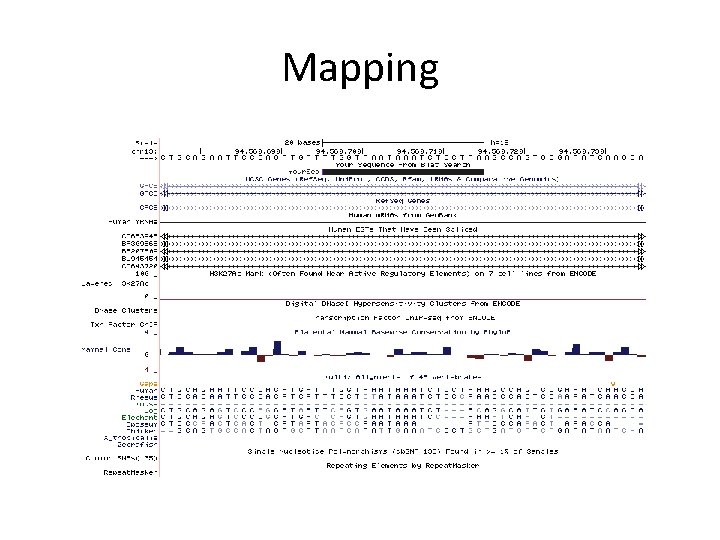
Mapping

Mapping Software • Long reads – BLAST, HMMER, SSEARCH • Short reads – BLAT – Bowtie, BWA, Partek, SOAP, Tophat, Olego, Barra. CUDA
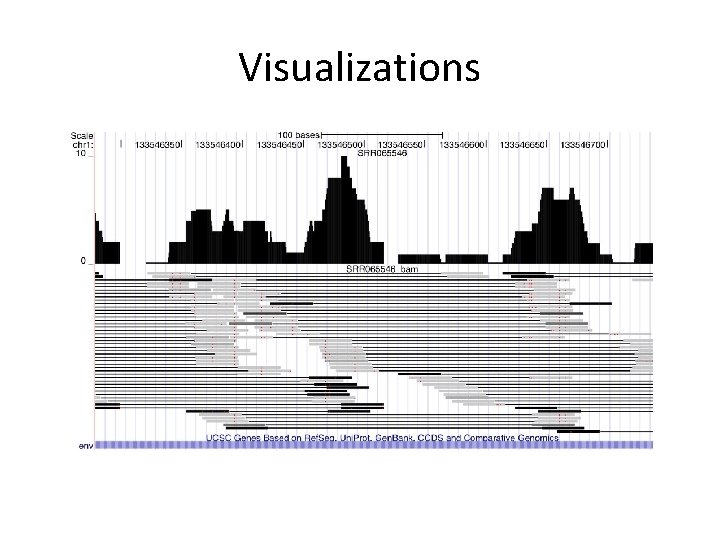
Visualizations
Visualizations • UCSC Genome Browser • Geno. Viewer, Samtools tview, Maq. View, rtracklayer, Bam. View, gbrowse 2 • Integrative Genomics Viewer (IGV)

Quantification • Peak calling – Qu. EST, MACS, Peak. Seq, T-PIC, SIPe. S, GLITR, SICER, Si. SSRs, OMT • Expression quantification – Cufflinks, NEUMA, RSEM, ABy. SS, ERANGE, RSAT, Velvet, MISO, RSEQ • SNP calling – samtools, Var. Scan, GATK, SOAP 2, real. SFS, Beagle, QCall, Ma. CH
![Peak Discovery [Pepke et al. Nature Methods 2009] Peak Discovery [Pepke et al. Nature Methods 2009]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-41.jpg)
Peak Discovery [Pepke et al. Nature Methods 2009]
![Transcript Quantification RPKM, FPKM [Pepke et al. Nature Methods 2009] Transcript Quantification RPKM, FPKM [Pepke et al. Nature Methods 2009]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-42.jpg)
Transcript Quantification RPKM, FPKM [Pepke et al. Nature Methods 2009]
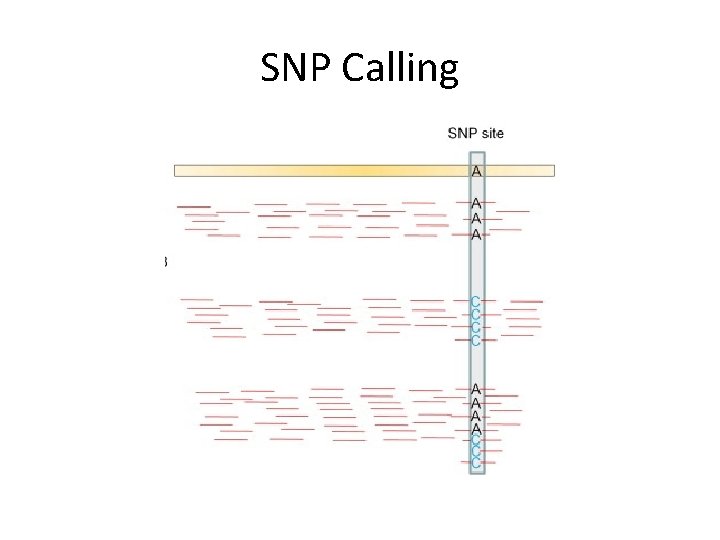
SNP Calling
![Typical RNA-seq Workflow [Trapnell et al. Nature Biotech 2010] Typical RNA-seq Workflow [Trapnell et al. Nature Biotech 2010]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-44.jpg)
Typical RNA-seq Workflow [Trapnell et al. Nature Biotech 2010]
![[Trapnell et al. Nature Biotech 2010] [Trapnell et al. Nature Biotech 2010]](http://slidetodoc.com/presentation_image_h2/97436f910b174a283db77c61fc329f59/image-45.jpg)
[Trapnell et al. Nature Biotech 2010]
- Slides: 45