Applications of GO Goals of Gene Ontology Project





















- Slides: 29

Applications of GO

Goals of Gene Ontology Project

Goals of Gene Ontology Project 1. Create controlled vocabularies – terms and definitions

Goals of Gene Ontology Project 1. Create controlled vocabularies – terms and definitions 2. Produce annotations to terms – gene product -> GO terms

Goals of Gene Ontology Project 1. Create controlled vocabularies – terms and definitions 2. Produce annotations to terms – gene product -> GO terms 3. Produce GO tools – browsing, searching and editing

Goals of Gene Ontology Project 1. Create controlled vocabularies – terms and definitions 2. Produce annotations to terms – gene product -> GO terms 3. Produce GO tools – browsing, searching and editing • Make everything publicly available

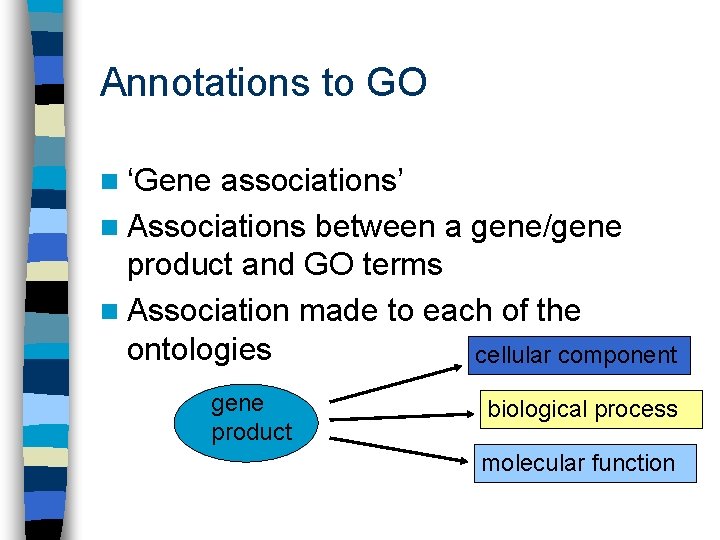
Annotations to GO n ‘Gene associations’ n Associations between a gene/gene product and GO terms n Association made to each of the ontologies cellular component gene product biological process molecular function

Annotations to GO n Three key parts: – gene name/id

Annotations to GO n Three key parts: – gene name/id – GO term

Annotations to GO n Three key parts: – gene name/id – GO term(s) – evidence for association

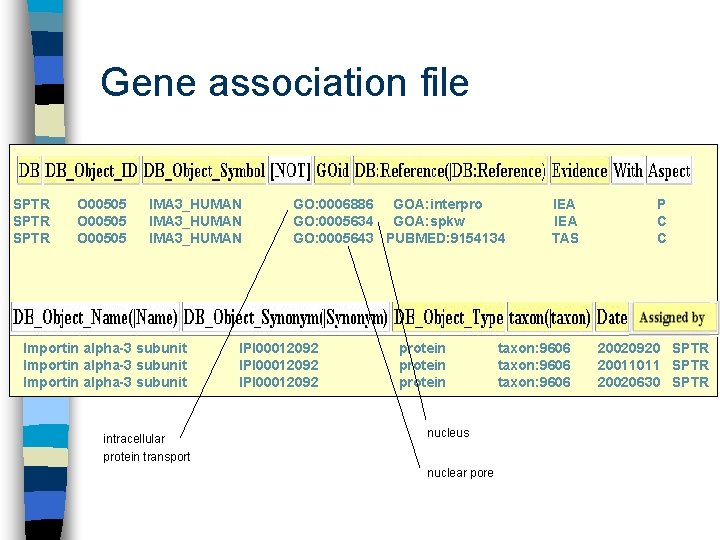
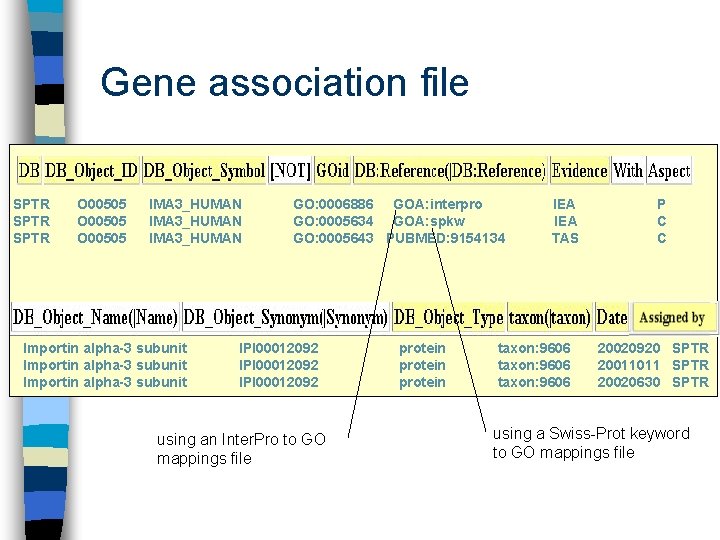
Gene association file SPTR O 00505 IMA 3_HUMAN Importin alpha-3 subunit intracellular protein transport GO: 0006886 GOA: interpro GO: 0005634 GOA: spkw GO: 0005643 PUBMED: 9154134 IPI 00012092 protein nucleus nuclear pore IEA TAS taxon: 9606 P C C 20020920 SPTR 20011011 SPTR 20020630 SPTR

Types of GO annotation: Electronic Annotation Manual Annotation

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Evidence types n ISS: Inferred from Sequence/structural Similarity IDA: Inferred from Direct Assay IPI: Inferred from Physical Interaction IMP: Inferred from Mutant Phenotype IGI: Inferred from Genetic Interaction IEP: Inferred from Expression Pattern TAS: Traceable Author Statement NAS: Non-traceable Author Statement IC: Inferred by Curator ND: No Data available n IEA: Inferred from electronic annotation n n n n

Inferred by Electronic Annotation n Annotation derived without human validation – mappings file e. g. interpro 2 go, ec 2 go. – Blast search ‘hits’ n Lower ‘quality’ than experimental codes

Mappings files Fatty acid biosynthesis ( Swiss-Prot Keyword) EC: 6. 4. 1. 2 (EC number) GO: Fatty acid biosynthesis (GO: 0006633) GO: acetyl-Co. A carboxylase activity (GO: 0003989) IPR 000438: Acetyl-Co. A carboxylase carboxyl transferase beta subunit (Inter. Pro entry) GO: acetyl-Co. A carboxylase activity (GO: 0003989)

Gene association file SPTR O 00505 IMA 3_HUMAN Importin alpha-3 subunit GO: 0006886 GOA: interpro GO: 0005634 GOA: spkw GO: 0005643 PUBMED: 9154134 IPI 00012092 using an Inter. Pro to GO mappings file protein IEA TAS taxon: 9606 P C C 20020920 SPTR 20011011 SPTR 20020630 SPTR using a Swiss-Prot keyword to GO mappings file

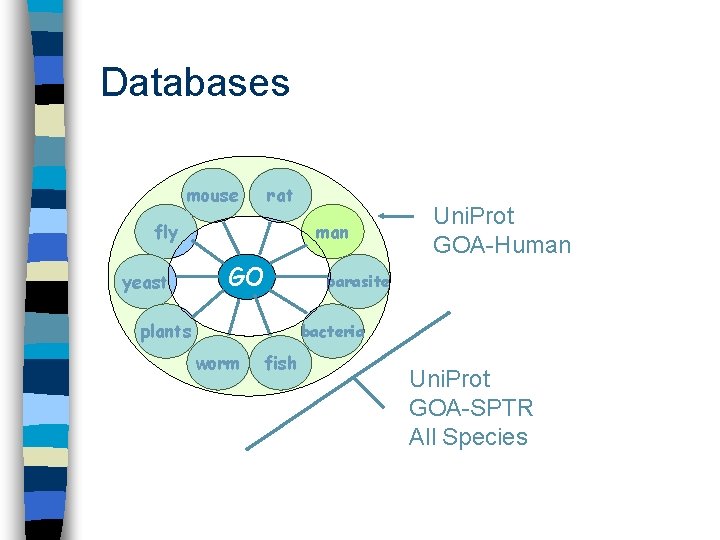
Submitting gene associations n Many model organism databases – Drosophila, mouse, Saccharomyces, rat, zebrafish, prokaryotes, Arabidopsis, slime mould, C. elegans, rice, parasites, viruses n Swiss-Prot (Uni. Prot) – Associations for >8000 species including human

Databases mouse rat fly yeast man GO plants Uni. Prot GOA-Human parasite bacteria worm fish Uni. Prot GOA-SPTR All Species

Finding GO terms In this study, we report the isolation and molecular characterization of the B. napus PERK 1 c. DNA, that is predicted to encode a novel receptor-like kinase. We have shown that like other plant RLKs, the kinase domain of PERK 1 has serine/threonine kinase activity, In addition, the location of a PERK 1 -GTP fusion protein to the plasma membrane supports the prediction that PERK 1 is an integral membrane protein…these kinases have been implicated in early stages of wound response…

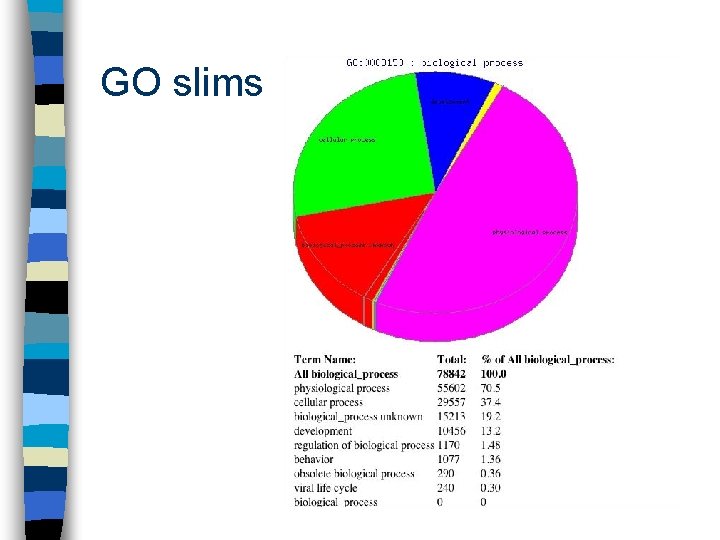
GO slims n Restricted view of the ontologies n Give broad view of gene function n Can be organism-specific or generic – plant – mammal – microbe

GO slims

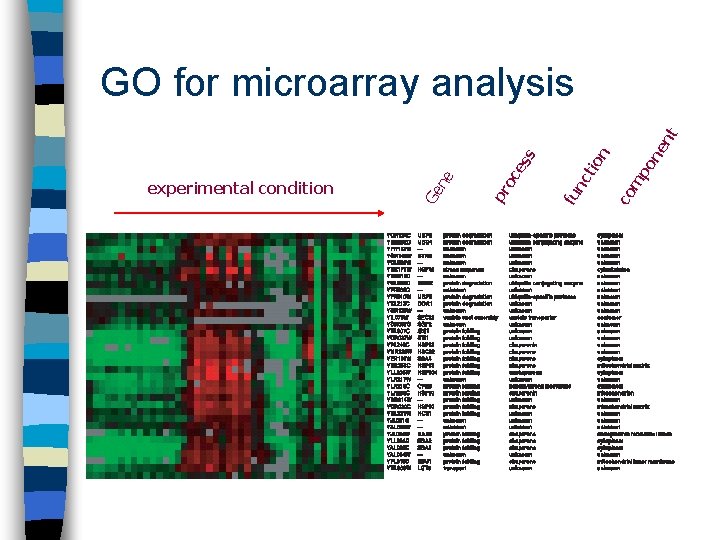
GO for microarray analysis n Annotations give ‘function’ label to genes n Ask meaningful questions of microarray data e. g. – genes involved in the same process, same/different expression patterns?

ne po m co tio n nc fu oc pr Ge experimental condition ne es s nt GO for microarray analysis

The tutorial n Part I – Navigating GO and its annotations using n Part II – Analysing microarray data using GO with