Bridging Biological Ontologies and Biosimulation The Ontology of

Bridging Biological Ontologies and Biosimulation: The Ontology of Physics for Biology Daniel L. Cook 1, 2 John H. Gennari 3 Jose L. V. Mejino 2 Maxwell L. Neal 3 1 Physiology & Biophysics, 2 Biological Structure 3 Biomedical and Health Informatics University of Washington, Seattle AMIA 2008, Washington, DC
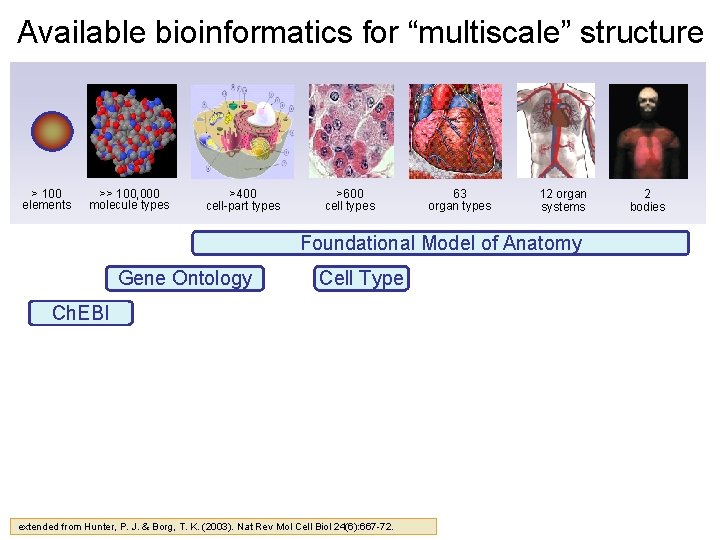
Available bioinformatics for “multiscale” structure > 100 elements >> 100, 000 molecule types >400 cell-part types >600 cell types 63 organ types 12 organ systems Foundational Model of Anatomy Gene Ontology Cell Type Ch. EBI extended from Hunter, P. J. & Borg, T. K. (2003). Nat Rev Mol Cell Biol 24(6): 667 -72. 2 bodies
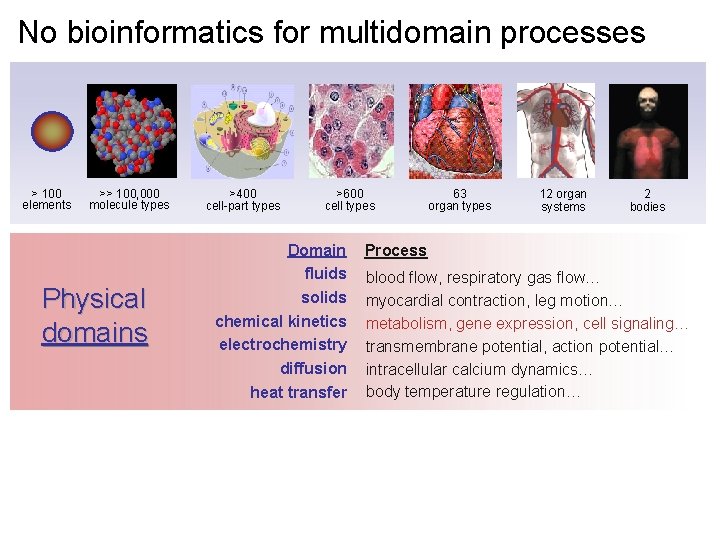
No bioinformatics for multidomain processes > 100 elements >> 100, 000 molecule types Physical domains >400 cell-part types >600 cell types Domain fluids solids chemical kinetics electrochemistry diffusion heat transfer 63 organ types 12 organ systems 2 bodies Process blood flow, respiratory gas flow… myocardial contraction, leg motion… metabolism, gene expression, cell signaling… transmembrane potential, action potential… intracellular calcium dynamics… body temperature regulation…
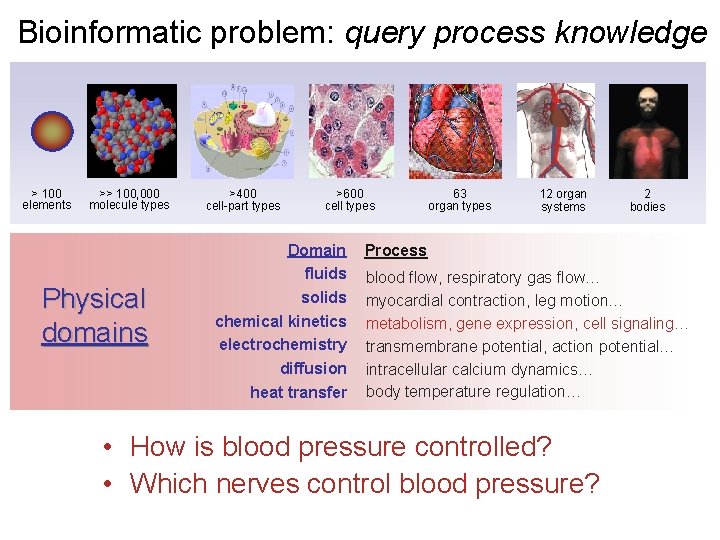
Bioinformatic problem: query process knowledge > 100 elements >> 100, 000 molecule types Physical domains >400 cell-part types >600 cell types Domain fluids solids chemical kinetics electrochemistry diffusion heat transfer 63 organ types 12 organ systems 2 bodies Process blood flow, respiratory gas flow… myocardial contraction, leg motion… metabolism, gene expression, cell signaling… transmembrane potential, action potential… intracellular calcium dynamics… body temperature regulation… • How is blood pressure controlled? • Which nerves control blood pressure?

Processes encoded as biosimulations models > 100 elements >> 100, 000 molecule types Physical domains >400 cell-part types >600 cell types Domain fluids solids chemical kinetics electrochemistry diffusion heat transfer physics-based biosimulation model 63 organ types 12 organ systems 2 bodies Process blood flow, respiratory gas flow… myocardial contraction, leg motion… metabolism, gene expression, cell signaling… transmembrane potential, action potential… intracellular calcium dynamics… body temperature regulation…

Available models constitute “physiome” > 100 elements >> 100, 000 molecule types Physical domains >400 cell-part types >600 cell types Domain fluids solids chemical kinetics electrochemistry diffusion heat transfer Hunter, P. J. & Borg, T. K. (2003). Nat Rev Mol Cell Biol 24(6): 667 -72. 63 organ types 12 organ systems 2 bodies Process blood flow, respiratory gas flow… myocardial contraction, leg motion… metabolism, gene expression, cell signaling… transmembrane potential, action potential… intracellular calcium dynamics… body temperature regulation… Physiome
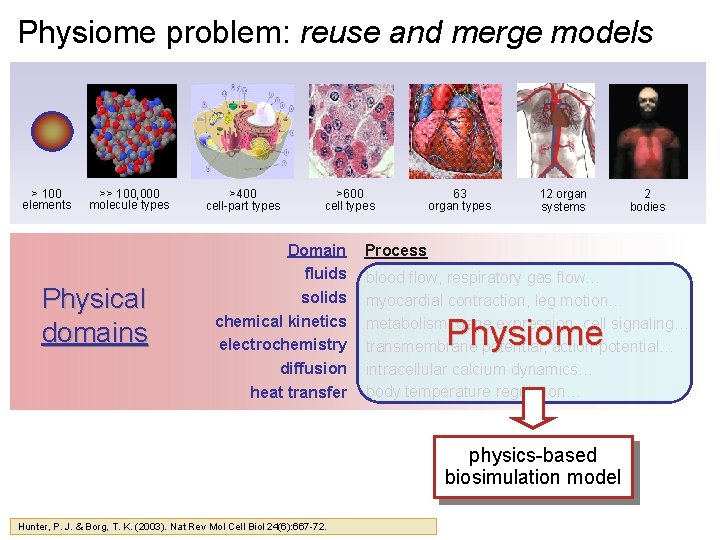
Physiome problem: reuse and merge models > 100 elements >> 100, 000 molecule types Physical domains >400 cell-part types >600 cell types Domain fluids solids chemical kinetics electrochemistry diffusion heat transfer 63 organ types 12 organ systems Process blood flow, respiratory gas flow… myocardial contraction, leg motion… metabolism, gene expression, cell signaling… transmembrane potential, action potential… intracellular calcium dynamics… body temperature regulation… Physiome physics-based biosimulation model Hunter, P. J. & Borg, T. K. (2003). Nat Rev Mol Cell Biol 24(6): 667 -72. 2 bodies
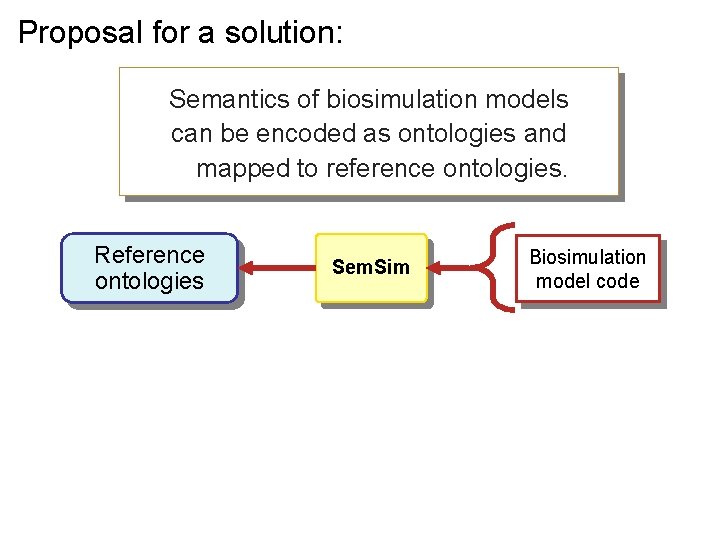
Proposal for a solution: Semantics of biosimulation models can be encoded as ontologies and mapped to reference ontologies. Reference ontologies Sem. Sim Biosimulation model code
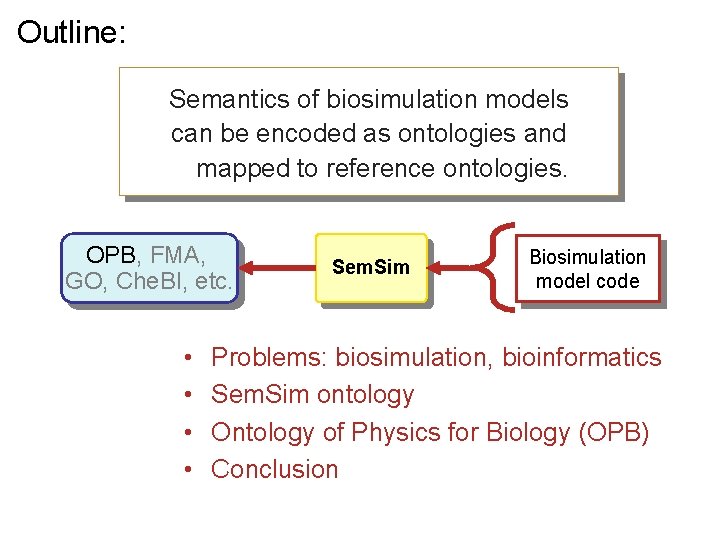
Outline: Semantics of biosimulation models can be encoded as ontologies and mapped to reference ontologies. OPB, FMA, GO, Che. BI, etc. • • Sem. Sim Biosimulation model code Problems: biosimulation, bioinformatics Sem. Sim ontology Ontology of Physics for Biology (OPB) Conclusion
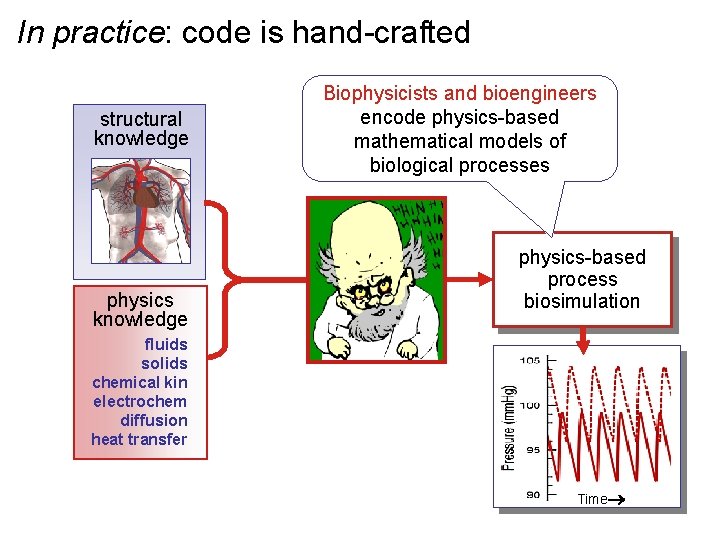
In practice: code is hand-crafted structural knowledge physics knowledge Biophysicists and bioengineers encode physics-based mathematical models of biological processes physics-based process biosimulation fluids solids chemical kin electrochem diffusion heat transfer Time®
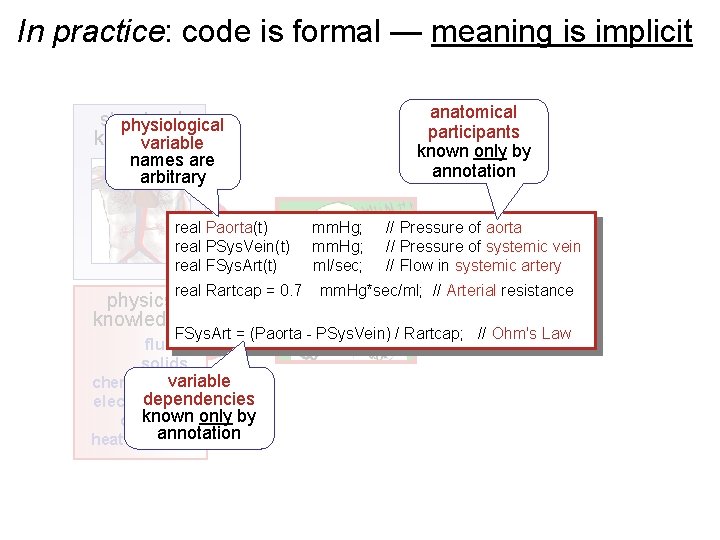
In practice: code is formal — meaning is implicit structural physiological knowledge variable names are arbitrary anatomical participants known only by annotation real Paorta(t) mm. Hg; // Pressure of aorta real PSys. Vein(t) mm. Hg; // Pressure of systemic vein real FSys. Art(t) ml/sec; // Flow in systemic artery real Rartcap = 0. 7 mm. Hg*sec/ml; // Arterial resistance physics knowledge FSys. Art = (Paorta - PSys. Vein) / Rartcap; // Ohm's Law fluids solids variable chemical kin dependencies electrochem known only by diffusion annotation heat transfer
In practice: multiple, incompatible languages structural knowledge physics knowledge fluids solids chemical kin electrochem diffusion heat transfer JSim, SBML, Cell. ML, Mat. Lab, others… physics-based process biosimulation
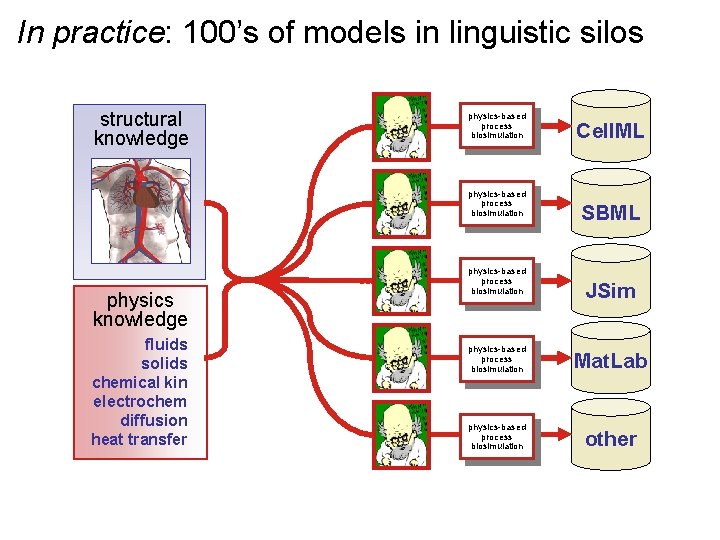
In practice: 100’s of models in linguistic silos structural knowledge physics-based process biosimulation physics knowledge fluids solids chemical kin electrochem diffusion heat transfer physics-based process biosimulation Cell. ML SBML JSim physics-based process biosimulation Mat. Lab physics-based process biosimulation other
Opportunity: a reservoir of process knowledge Cell. ML SBML JSim Mat. Lab other
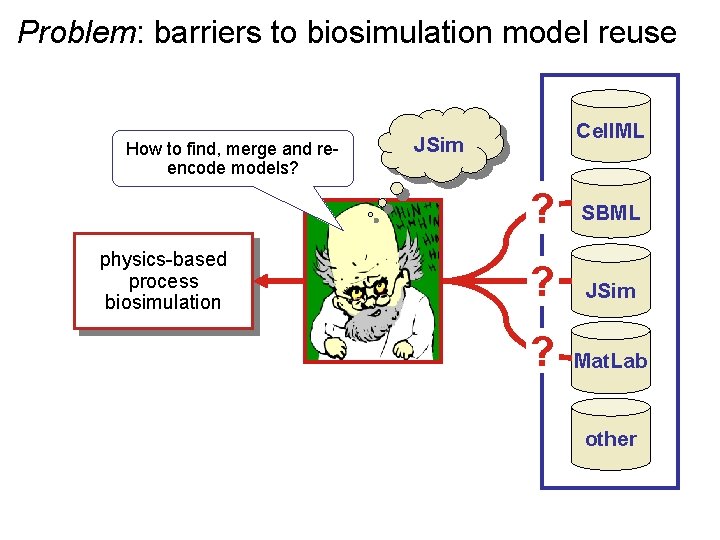
Problem: barriers to biosimulation model reuse How to find, merge and reencode models? physics-based process biosimulation Cell. ML JSim ? SBML ? JSim ? Mat. Lab other
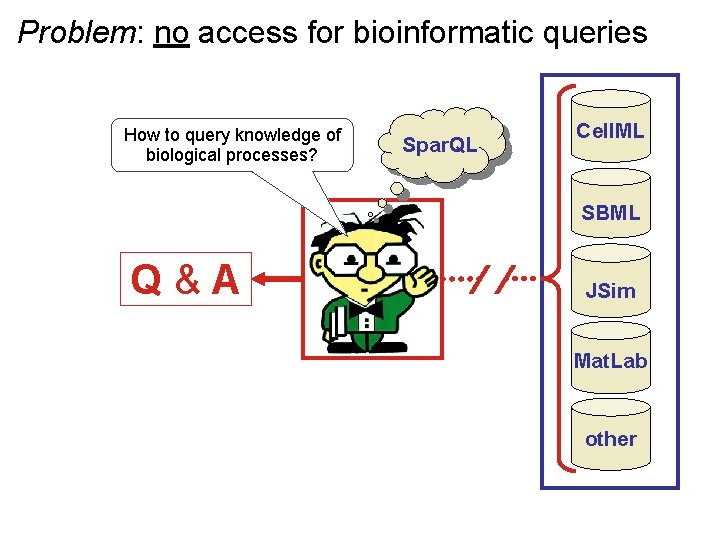
Problem: no access for bioinformatic queries How to query knowledge of biological processes? Spar. QL Cell. ML SBML Q & A JSim Mat. Lab other

Two fields, two problems: Biosimulation — re-use biosimulation models • Find models of blood pressure control. • Which models include neural-control? Bioinformatics — query process knowledge • How is blood pressure controlled? • Which nerves control blood pressure?
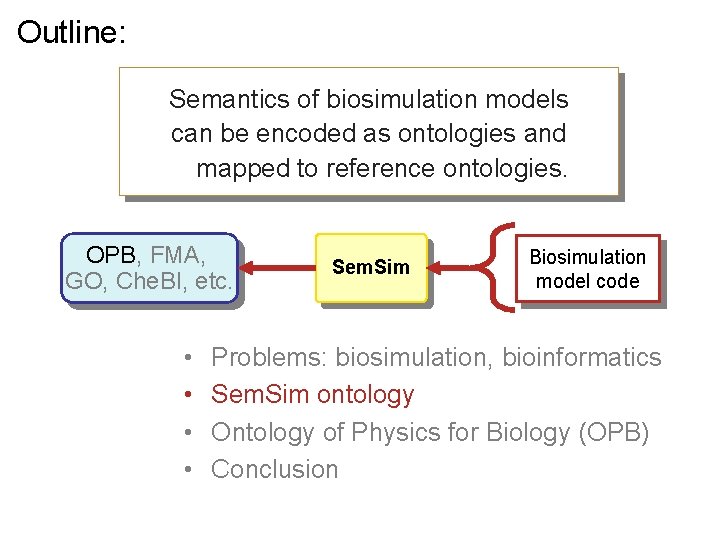
Outline: Semantics of biosimulation models can be encoded as ontologies and mapped to reference ontologies. OPB, FMA, GO, Che. BI, etc. • • Sem. Sim Biosimulation model code Problems: biosimulation, bioinformatics Sem. Sim ontology Ontology of Physics for Biology (OPB) Conclusion
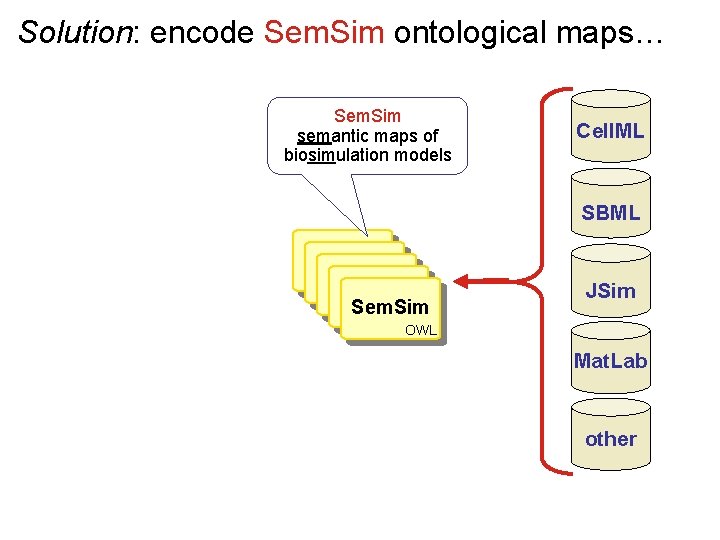
Solution: encode Sem. Sim ontological maps… Sem. Sim semantic maps of biosimulation models Cell. ML SBML Sem. Sim JSim OWL Mat. Lab other
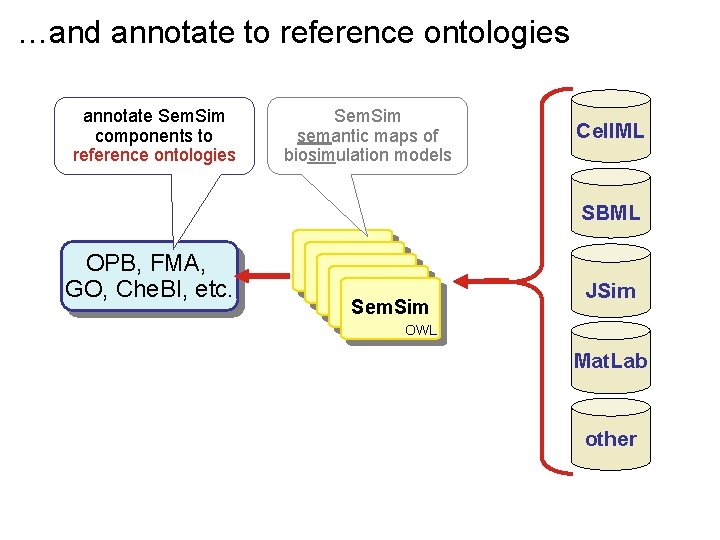
…and annotate to reference ontologies annotate Sem. Sim components to reference ontologies Sem. Sim semantic maps of biosimulation models Cell. ML SBML OPB, FMA, GO, Che. BI, etc. Sem. Sim JSim OWL Mat. Lab other
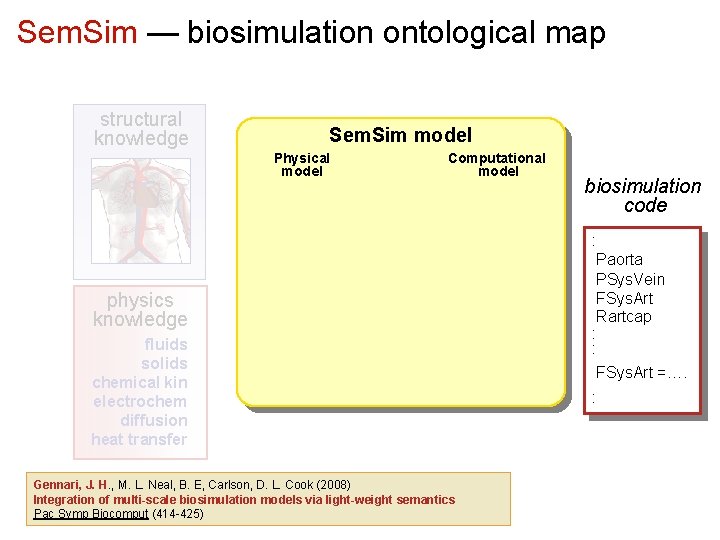
Sem. Sim — biosimulation ontological map structural knowledge Sem. Sim model Physical model Computational model physics knowledge fluids solids chemical kin electrochem diffusion heat transfer Gennari, J. H. , M. L. Neal, B. E, Carlson, D. L. Cook (2008) Integration of multi-scale biosimulation models via light-weight semantics Pac Symp Biocomput (414 -425) biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
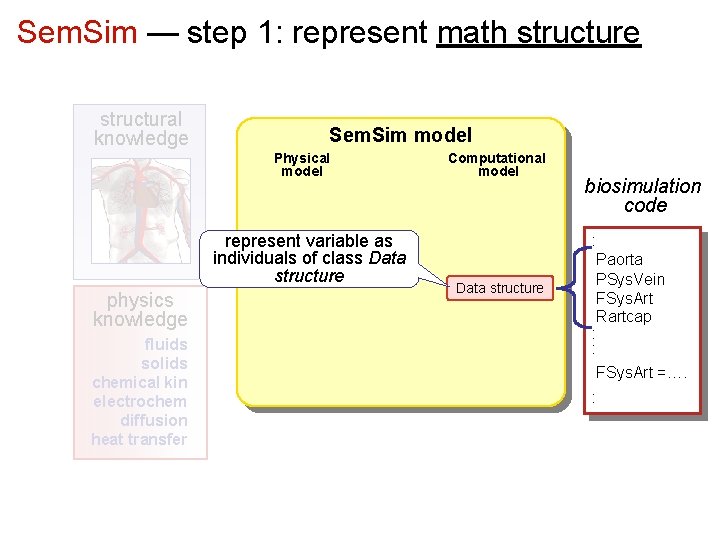
Sem. Sim — step 1: represent math structure structural knowledge Sem. Sim model Physical model represent variable as individuals of class Data structure physics knowledge fluids solids chemical kin electrochem diffusion heat transfer Computational model Data structure biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
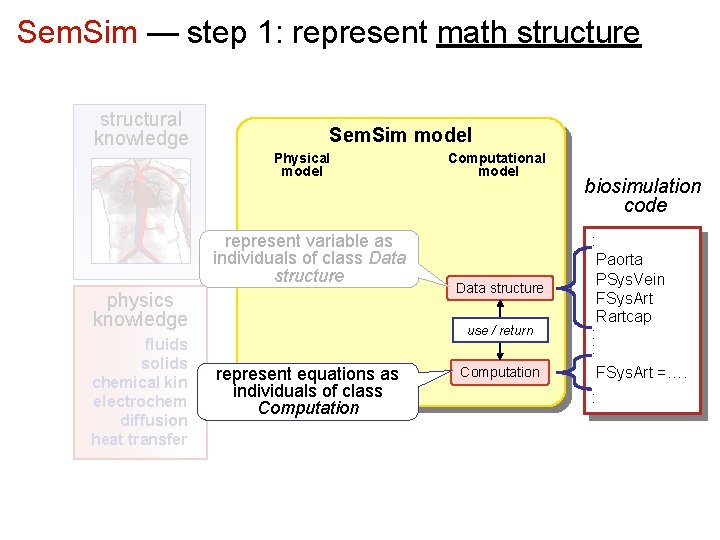
Sem. Sim — step 1: represent math structure structural knowledge Sem. Sim model Physical model represent variable as individuals of class Data structure physics knowledge fluids solids chemical kin electrochem diffusion heat transfer Computational model Data structure use / return represent equations as individuals of class Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
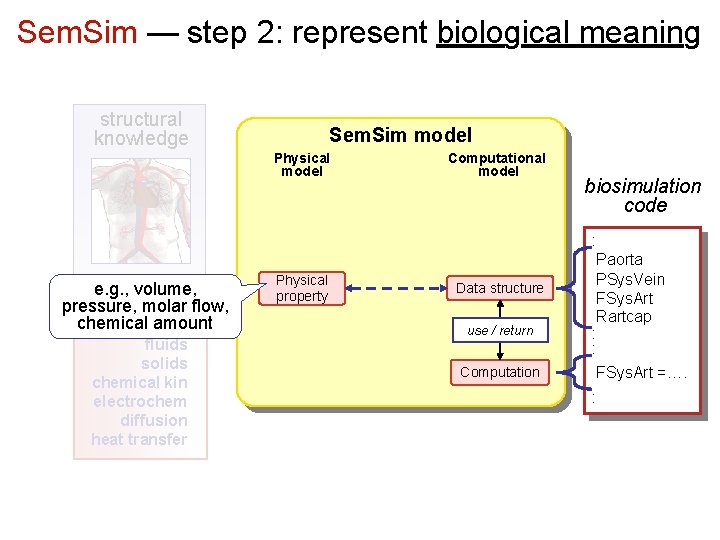
Sem. Sim — step 2: represent biological meaning structural knowledge Sem. Sim model Physical model e. g. , volume, physics pressure, molar flow, knowledge chemical amount fluids solids chemical kin electrochem diffusion heat transfer Physical property Computational model Data structure use / return Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
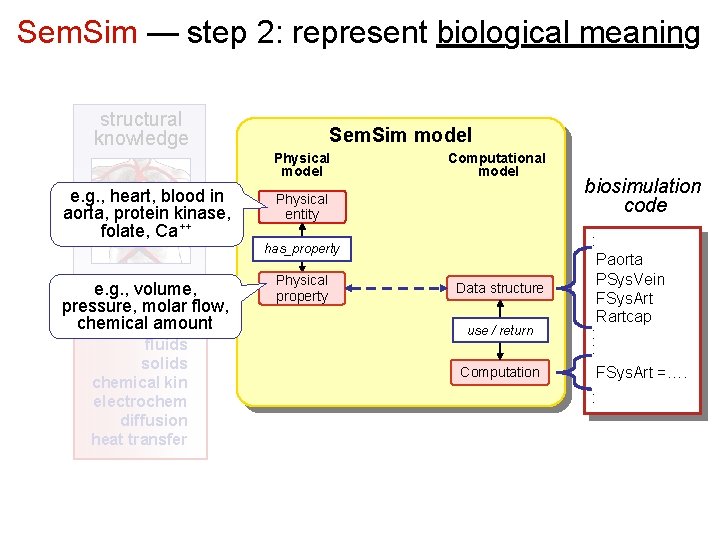
Sem. Sim — step 2: represent biological meaning structural knowledge Sem. Sim model Physical model e. g. , heart, blood in aorta, protein kinase, folate, Ca++ Computational model Physical entity has_property e. g. , volume, physics pressure, molar flow, knowledge chemical amount fluids solids chemical kin electrochem diffusion heat transfer Physical property Data structure use / return Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
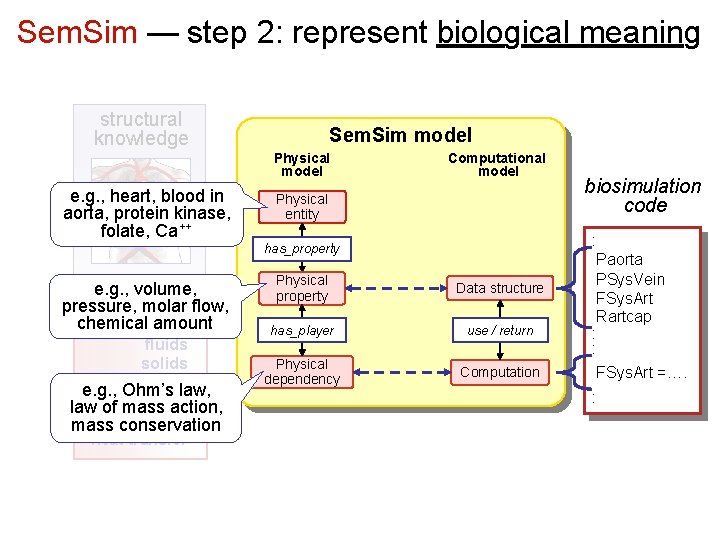
Sem. Sim — step 2: represent biological meaning structural knowledge Sem. Sim model Physical model e. g. , heart, blood in aorta, protein kinase, folate, Ca++ Computational model Physical entity has_property e. g. , volume, physics pressure, molar flow, knowledge chemical amount fluids solids chemical kin e. g. , Ohm’s law, electrochem law of mass action, diffusion mass conservation heat transfer Physical property Data structure has_player use / return Physical dependency Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
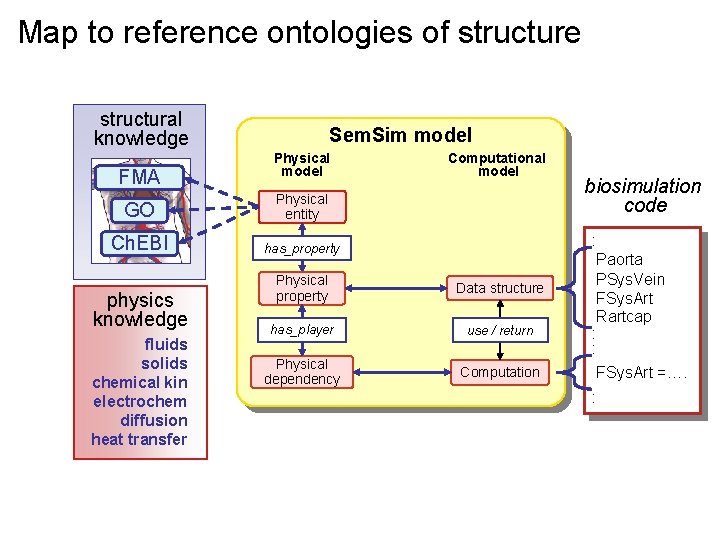
Map to reference ontologies of structure structural knowledge FMA Sem. Sim model Physical model GO Physical entity Ch. EBI has_property physics knowledge fluids solids chemical kin electrochem diffusion heat transfer Computational model Physical property Data structure has_player use / return Physical dependency Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
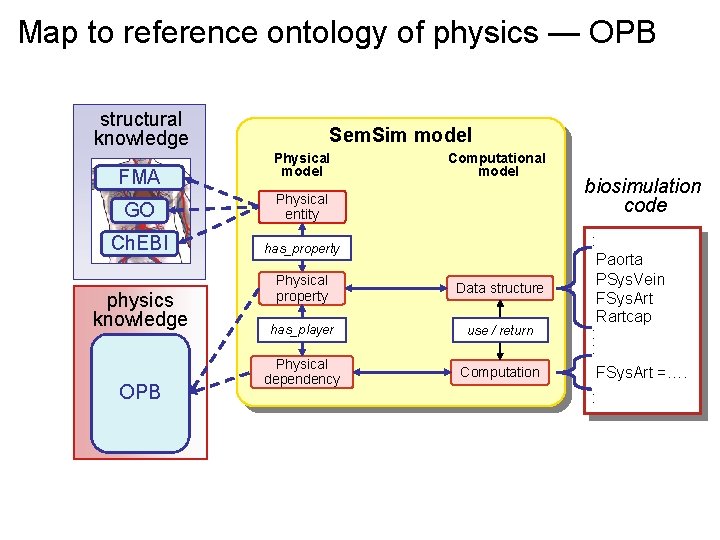
Map to reference ontology of physics — OPB structural knowledge FMA Sem. Sim model Physical model GO Physical entity Ch. EBI has_property physics knowledge fluids solids chemical kin OPB electrochem diffusion heat transfer Computational model Physical property Data structure has_player use / return Physical dependency Computation biosimulation code : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. :
Outline: Semantics of biosimulation models can be encoded as ontologies and mapped to reference ontologies. OPB, FMA, GO, Che. BI, etc. • • Sem. Sim Biosimulation model code Problems: biosimulation, bioinformatics Sem. Sim ontology Ontology of Physics for Biology (OPB) Conclusion

OPB foundational theory — system dynamics Engineering system dynamics • Bond graph theory Karnopp, Margolis, Rosenberg (1968) • Eng. Math - Ontology for Engineering Mathematics Gruber, Olsen (1994) • PHYSYS - Physical Systems Ontology Borst, Top, Akkermans (1994) Biochemical system dynamics • Network thermodynamics Oster, Perelson, �Katchalsky 1971) ( Mickulecky (1983) Beard, Qian (2008)

OPB representational goals • Represent abstractions used in physics-based biosimulations—not a theory of “reality”. • Adhere to OBO principles. • Implement in OWL; deploy to OBO and Bio. Portal.

OPB: Physics analytical entity A Physics analytical entity is an abstraction of the real world created within the science of classical physics for the description of physical entities and the analysis of physical processes. OPB
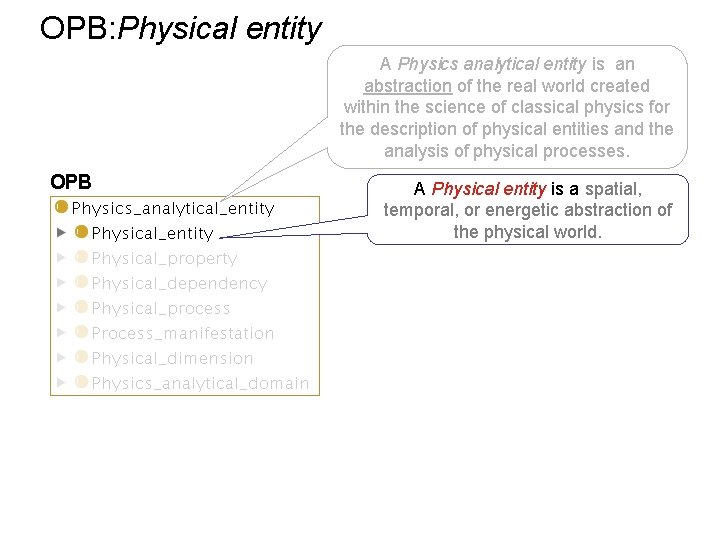
OPB: Physical entity A Physics analytical entity is an abstraction of the real world created within the science of classical physics for the description of physical entities and the analysis of physical processes. OPB A Physical entity is a spatial, temporal, or energetic abstraction of the physical world.
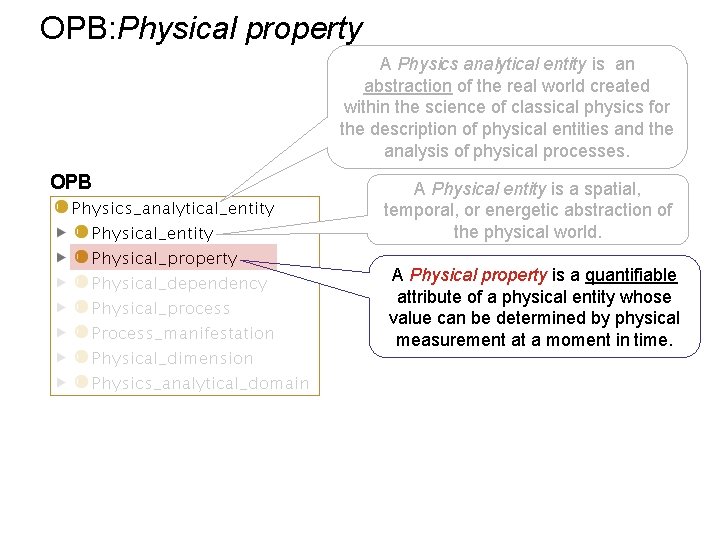
OPB: Physical property A Physics analytical entity is an abstraction of the real world created within the science of classical physics for the description of physical entities and the analysis of physical processes. OPB A Physical entity is a spatial, temporal, or energetic abstraction of the physical world. A Physical property is a quantifiable attribute of a physical entity whose value can be determined by physical measurement at a moment in time.

Physical property organizing principle Physical domain fluids volume flow pressure volume pressure momentum solids velocity force displacement solid momentum chemical kinetics molar flow chemical potential chemical amount ---- electrophysiology ionic current voltage charge ---- diffusion particle flow chemical potential particle number ---- heat flow temperature heat amount ---- heat transfer
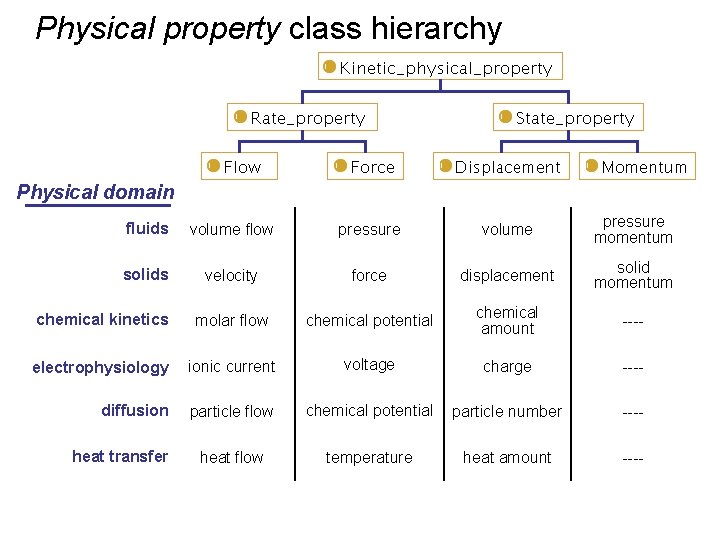
Physical property class hierarchy Physical domain fluids volume flow pressure volume pressure momentum solids velocity force displacement solid momentum chemical kinetics molar flow chemical potential chemical amount ---- electrophysiology ionic current voltage charge ---- diffusion particle flow chemical potential particle number ---- heat flow temperature heat amount ---- heat transfer
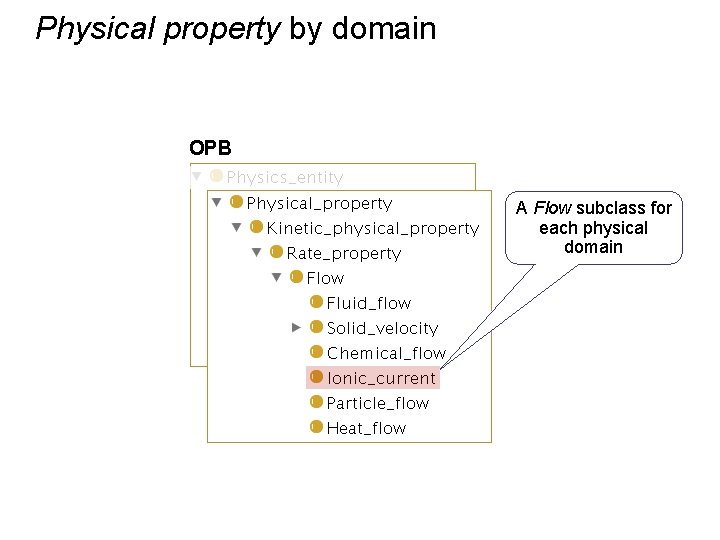
Physical property by domain OPB A Flow subclass for each physical domain

Physical dependency A Physics analytical entity is an abstraction of the real world created within the science of classical physics for the description of physical entities and the analysis of physical processes. OPB A Physical entity is a spatial, temporal, or energetic abstraction of the physical world. A Physical property is a quantifiable attribute of a physical entity whose value can be determined by physical measurement at a moment in time. A Physical dependency is a quantitative dependency between the magnitudes of two or more physical properties according to a physical law.
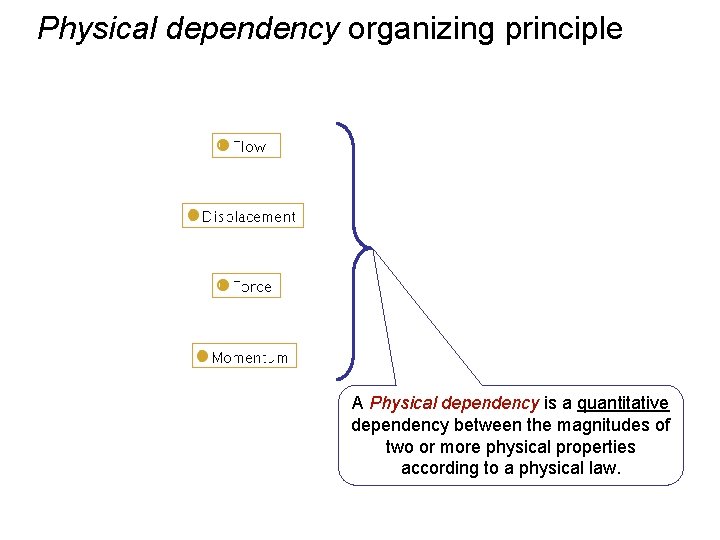
Physical dependency organizing principle A Physical dependency is a quantitative dependency between the magnitudes of two or more physical properties according to a physical law.
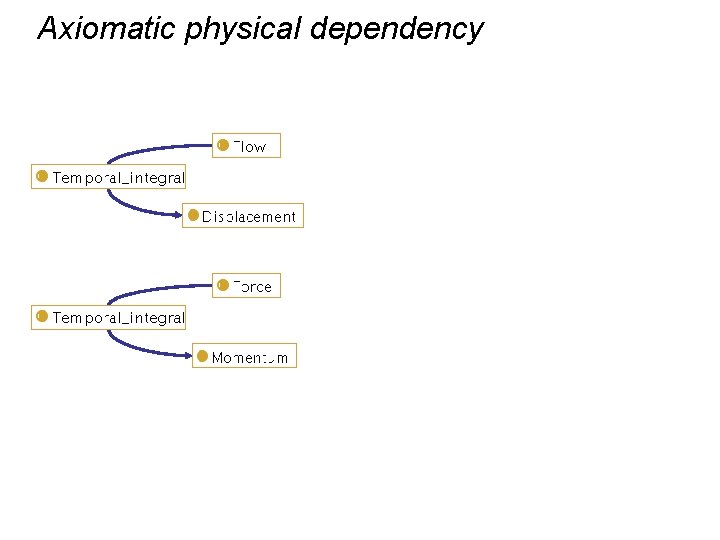
Axiomatic physical dependency
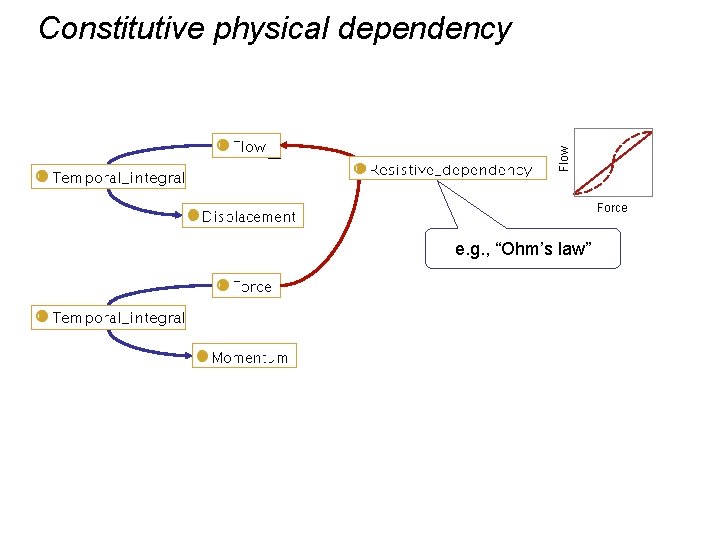
Flow Constitutive physical dependency Force e. g. , “Ohm’s law”
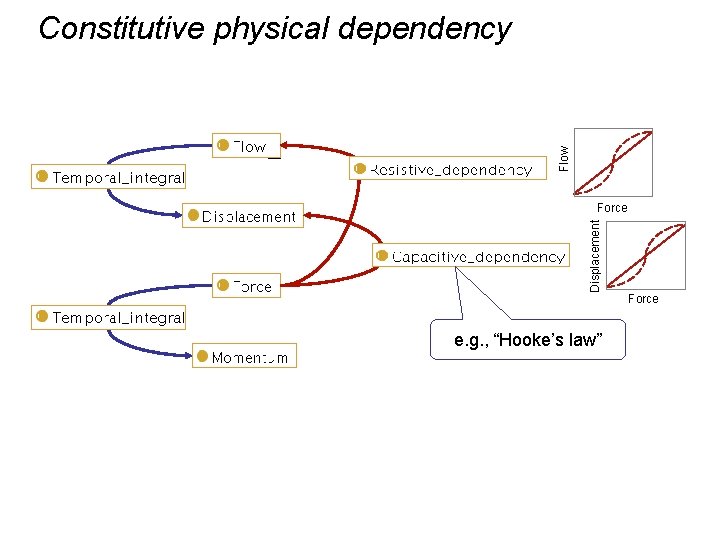
Flow Constitutive physical dependency Displacement Force e. g. , “Hooke’s law” Force
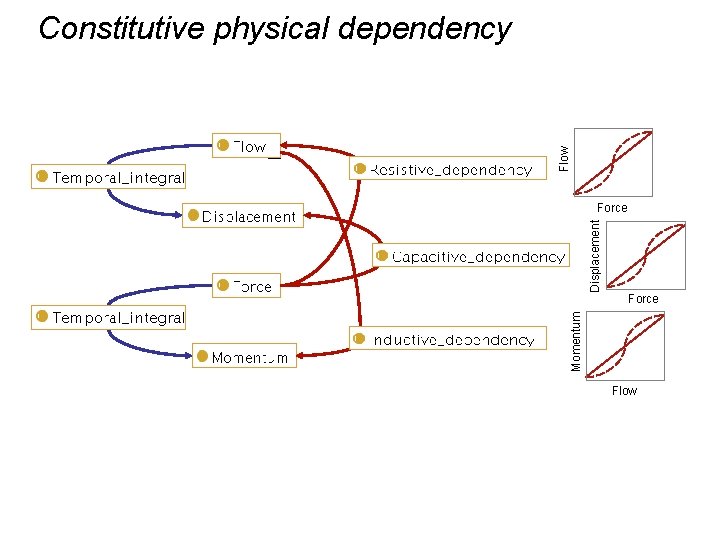
Flow Constitutive physical dependency Force Momentum Displacement Force Flow

Physical dependency class hierarchy OPB
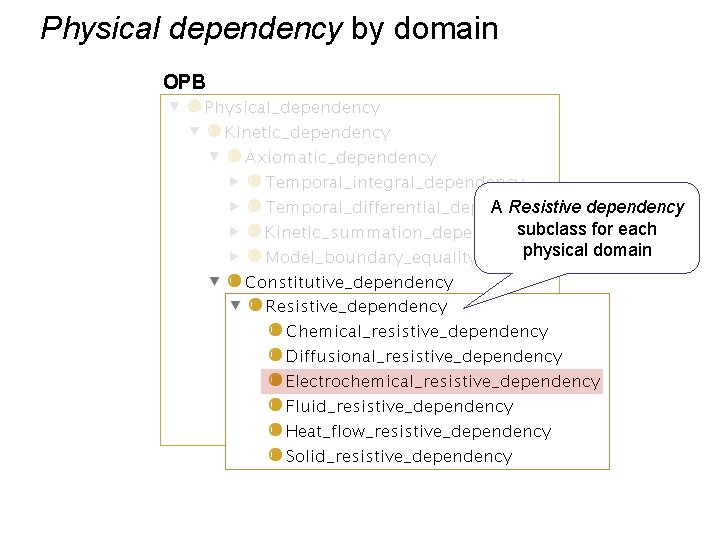
Physical dependency by domain OPB A Resistive dependency subclass for each physical domain
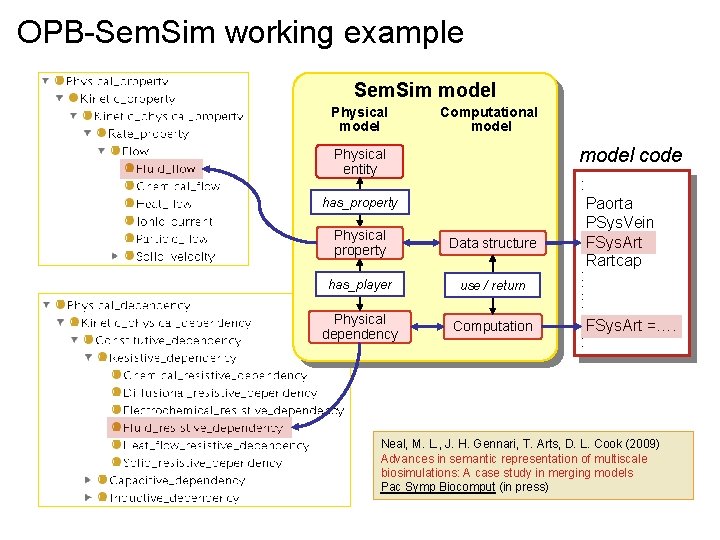
OPB-Sem. Sim working example Sem. Sim model Physical model Computational model code Physical entity has_property Physical property Data structure has_player use / return Physical dependency Computation : Paorta PSys. Vein FSys. Art Rartcap : : : FSys. Art =…. : Neal, M. L. , J. H. Gennari, T. Arts, D. L. Cook (2009) Advances in semantic representation of multiscale biosimulations: A case study in merging models Pac Symp Biocomput (in press)
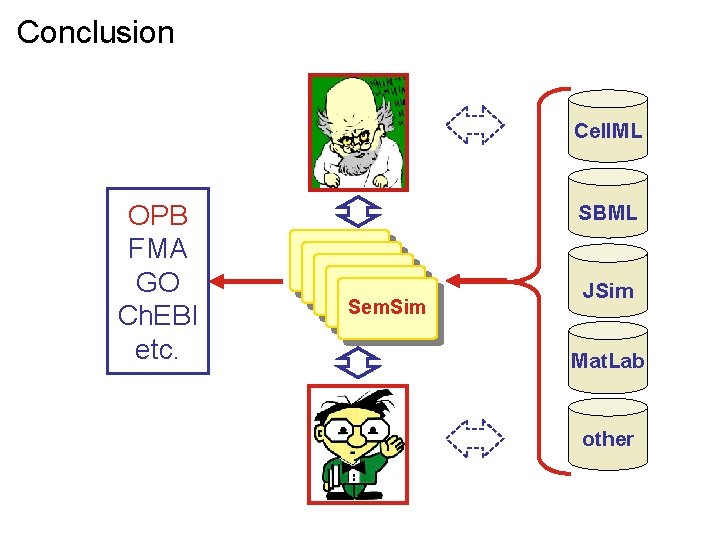
Conclusion Cell. ML OPB FMA GO Ch. EBI etc. SBML Sem. Sim JSim Mat. Lab other

Acknowledgements Sem. Sim / OPB team • Maxwell L. Neal (Grad student) • Michal Galdzicki (Grad student) • John H. Gennari, Ph. D (Assoc Prof) • Daniel L. Cook, MD, Ph. D (Res Prof) UW contributors Bioinformatics • Cornelius Rosse • Onard Mejino • James Brinkley • Todd Detwiler Partial funding from NIH MLN, MG: T 15 LM 007442 -06 DLC, JHG: R 01 HL 087706 -01 Biophysics / biosimulation • James B. Bassingthwaighte • Herbert Sauro • Erik Butterworth • Hong Qian • Adriana Emmi • Fred Bookstein
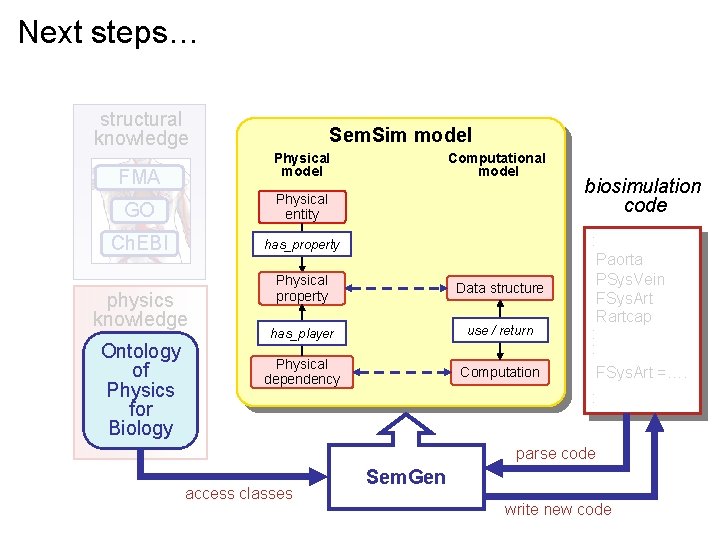
Next steps… structural knowledge Sem. Sim model Physical model FMA GO Physical entity Ch. EBI has_property physics knowledge fluids Ontology solids of chemical kin Physics electrochem for diffusion Biology Computational model Physical property Data structure has_player use / return Physical dependency Computation : Paorta PSys. Vein FSys. Art Rartcap : : FSys. Art =…. : heat transfer access classes biosimulation code parse code Sem. Gen write new code
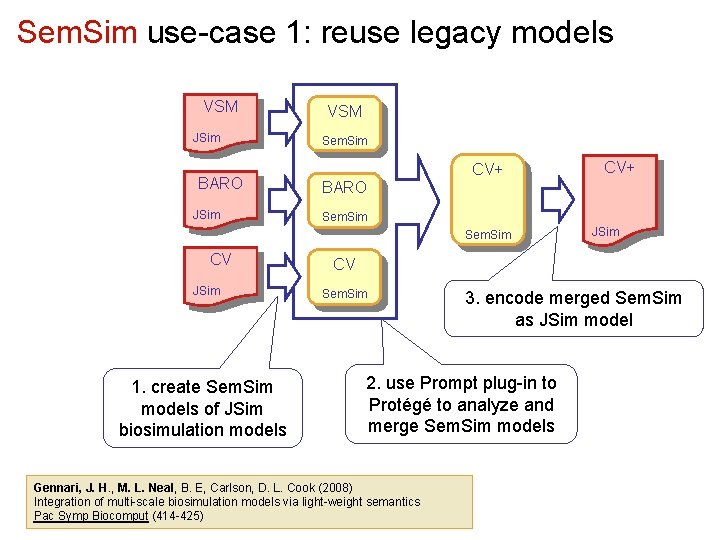
Sem. Sim use-case 1: reuse legacy models VSM JSim BARO JSim VSM Sem. Sim CV+ BARO Sem. Sim CV JSim 1. create Sem. Sim models of JSim biosimulation models CV+ JSim CV Sem. Sim 3. encode merged Sem. Sim as JSim model 2. use Prompt plug-in to Protégé to analyze and merge Sem. Sim models Gennari, J. H. , M. L. Neal, B. E, Carlson, D. L. Cook (2008) Integration of multi-scale biosimulation models via light-weight semantics Pac Symp Biocomput (414 -425)
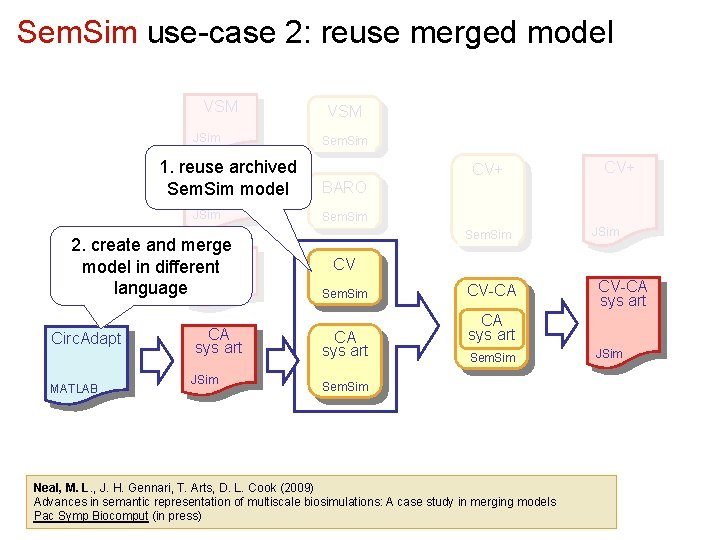
Sem. Sim use-case 2: reuse merged model VSM JSim 1. reuse archived BARO Sem. Sim model JSim 2. create and merge CV model in different language JSim Circ. Adapt MATLAB CA sys art JSim VSM Sem. Sim CV+ BARO Sem. Sim JSim CV Sem. Sim CA sys art CV-CA sys art Sem. Sim Neal, M. L. , J. H. Gennari, T. Arts, D. L. Cook (2009) Advances in semantic representation of multiscale biosimulations: A case study in merging models Pac Symp Biocomput (in press) JSim
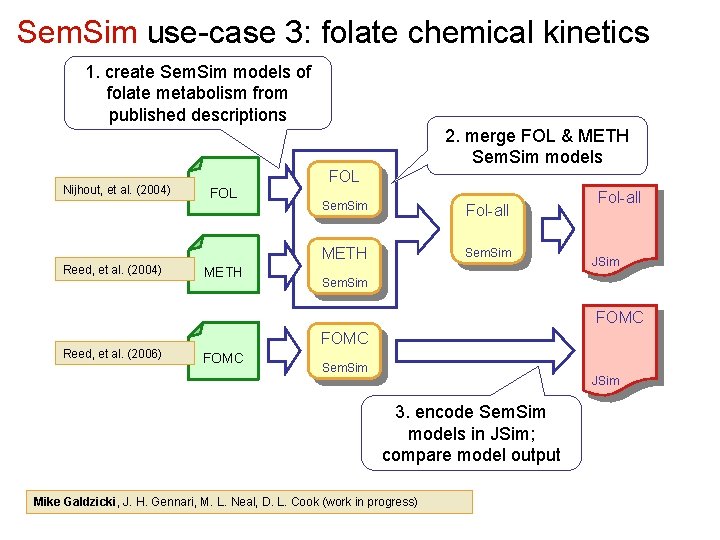
Sem. Sim use-case 3: folate chemical kinetics 1. create Sem. Sim models of folate metabolism from published descriptions 2. merge FOL & METH Sem. Sim models Nijhout, et al. (2004) Reed, et al. (2004) FOL METH FOL Sem. Sim Fol-all METH Sem. Sim Fol-all JSim Sem. Sim FOMC Reed, et al. (2006) FOMC Sem. Sim JSim 3. encode Sem. Sim models in JSim; compare model output Mike Galdzicki, J. H. Gennari, M. L. Neal, D. L. Cook (work in progress)
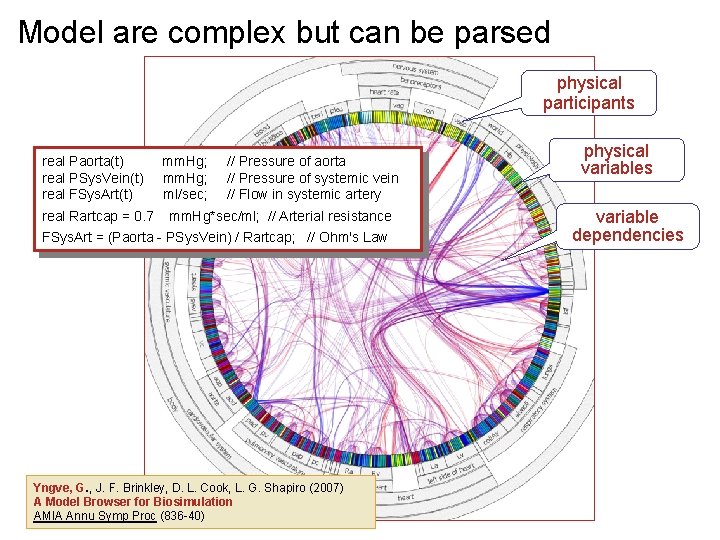
Model are complex but can be parsed physical participants real Paorta(t) mm. Hg; // Pressure of aorta real PSys. Vein(t) mm. Hg; // Pressure of systemic vein real FSys. Art(t) ml/sec; // Flow in systemic artery real Rartcap = 0. 7 mm. Hg*sec/ml; // Arterial resistance FSys. Art = (Paorta - PSys. Vein) / Rartcap; // Ohm's Law Yngve, G. , J. F. Brinkley, D. L. Cook, L. G. Shapiro (2007) A Model Browser for Biosimulation AMIA Annu Symp Proc (836 -40) physical variables variable dependencies
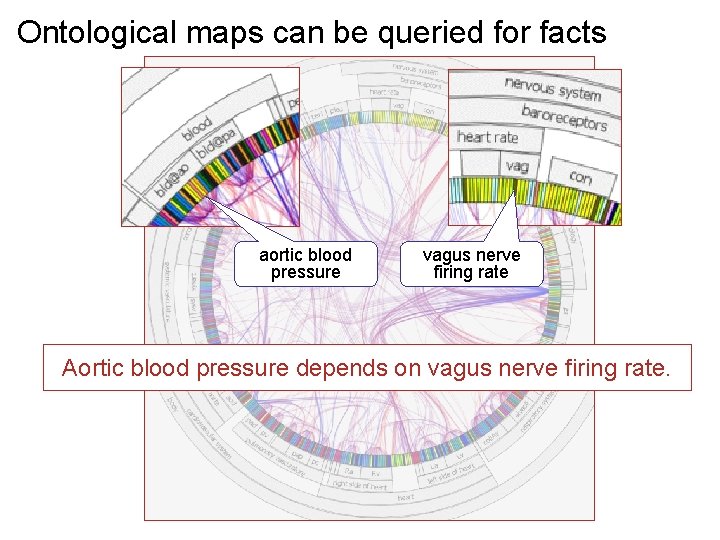
Ontological maps can be queried for facts aortic blood pressure vagus nerve firing rate Aortic blood pressure depends on vagus nerve firing rate.
- Slides: 54