Ontology Databases Detecting Inconsistencies in the Gene Ontology

Ontology Databases: Detecting Inconsistencies in the Gene Ontology using Not-gadgets Paea Le. Pendu University of Oregon Talk: National Center for Biomedical Ontology ◦ Stanford University ◦ September, 2009

General Interests Logic Programming Languages Automated Reasoning Databases

Outline • Ontology-based Data Management – Background, Motivation – Theory – Benchmarking – Application Domain, Query Answering • Inconsistency Detection – Theory – The serotonin example – GO plus ZFIN, MGI annotations
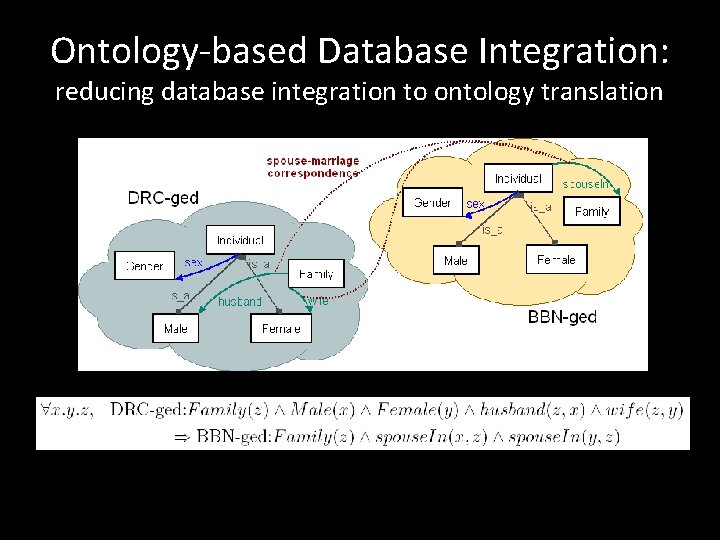
Ontology-based Database Integration: reducing database integration to ontology translation

Ontology-based Database Integration: reducing database integration to ontology translation

Ontology-based Data Management Class Property Datatype Sub. Class Restriction Relation Individual Attribute Datatype Keys Constraint View Trigger Tuple
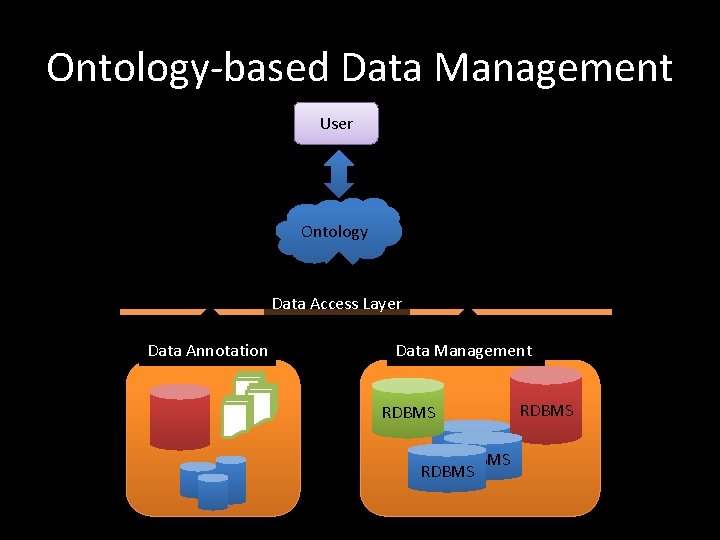
Ontology-based Data Management User Ontology Data Access Layer Data Annotation Data Management RDBMS RDBMS

Example: sisters-siblings This is what we know : All sisters are siblings. Hilary and Lynn are sisters. This is what we want to know : Who are siblings? { <x, y> | sibling. Of(x, y) } Obviously, the answer should be : Hilary and Lynn are siblings. { <Hilary, Lynn> }

Example: sisters-siblings

Example: sisters-siblings
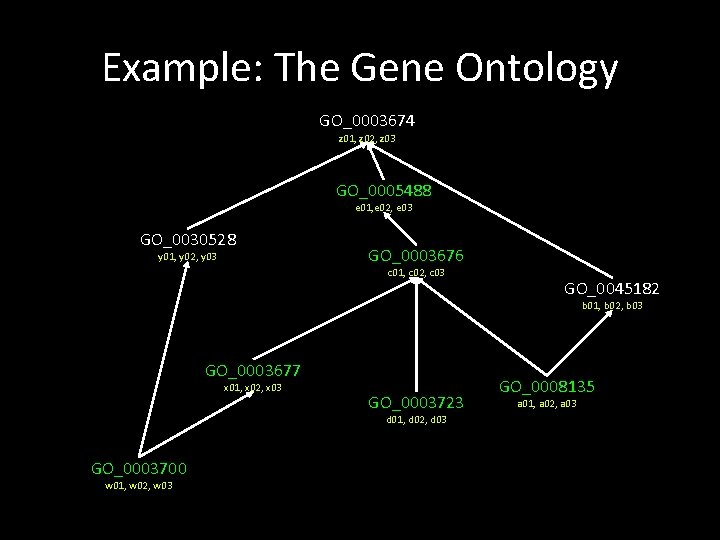
Example: The Gene Ontology GO_0003674 z 01, z 02, z 03 GO_0005488 e 01, e 02, e 03 GO_0030528 y 01, y 02, y 03 GO_0003676 c 01, c 02, c 03 GO_0045182 b 01, b 02, b 03 GO_0003677 x 01, x 02, x 03 GO_0003723 d 01, d 02, d 03 GO_0003700 w 01, w 02, w 03 GO_0008135 a 01, a 02, a 03
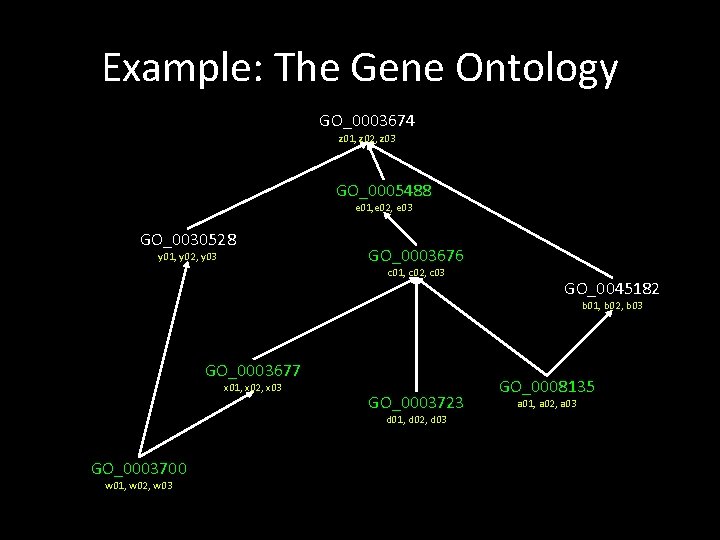
Example: The Gene Ontology GO_0003674 z 01, z 02, z 03 GO_0005488 e 01, e 02, e 03 GO_0030528 y 01, y 02, y 03 GO_0003676 c 01, c 02, c 03 GO_0045182 b 01, b 02, b 03 GO_0003677 x 01, x 02, x 03 GO_0003723 d 01, d 02, d 03 GO_0003700 w 01, w 02, w 03 GO_0008135 a 01, a 02, a 03

Ontology Databases: General Models for Database Designs • Generality is important – Avoid rewriting • Scalability of KB is important – Persistence, caching and indexing • Major generic models – Horizontal Models – Vertical Models – Decomposition Storage Models
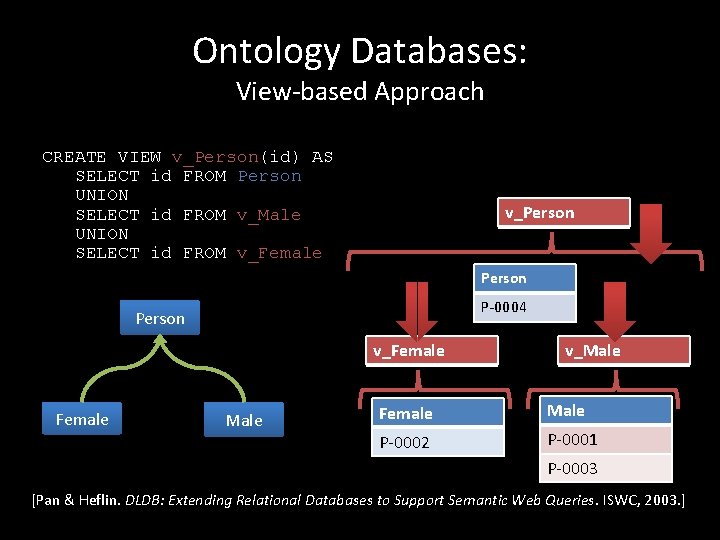
Ontology Databases: View-based Approach CREATE VIEW v_Person(id) AS SELECT id FROM Person UNION SELECT id FROM v_Male UNION SELECT id FROM v_Female v_Person P-0004 Person v_Female Male v_Male Female Male P-0002 P-0001 P-0003 [Pan & Heflin. DLDB: Extending Relational Databases to Support Semantic Web Queries. ISWC, 2003. ]

Ontology Databases: Active Database Approach ON INSERT into Male INSERT into Person On INSERT into Female INSERT into Person Female Male [Le. Pendu, et al. Ontology Database: a New Method for Semantic Modeling and an Application to Brainwave Data. SSDBM, 2008. ]
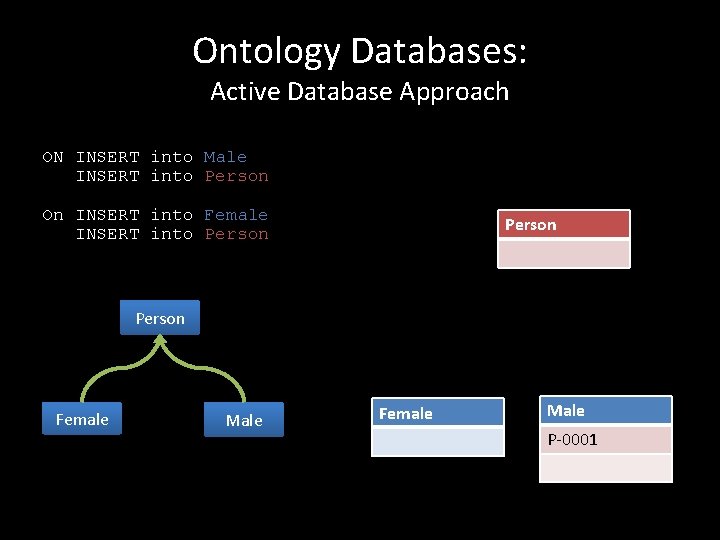
Ontology Databases: Active Database Approach ON INSERT into Male INSERT into Person On INSERT into Female INSERT into Person Female Male P-0001
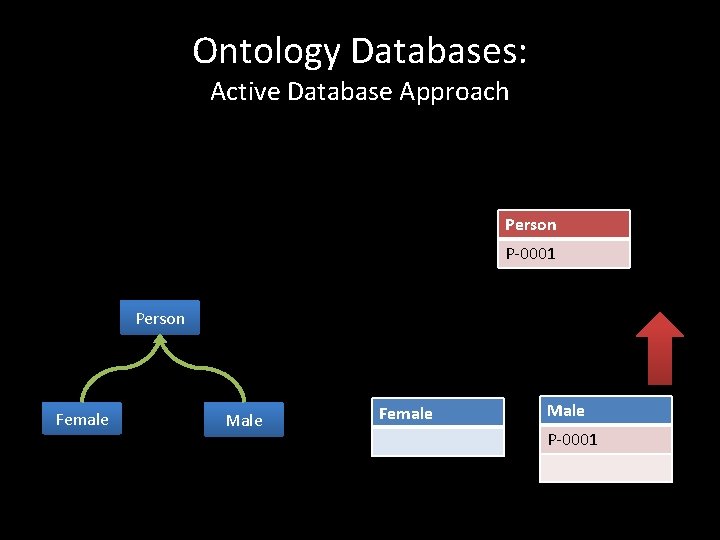
Ontology Databases: Active Database Approach Person P-0001 Person Female Male P-0001

Ontology Databases: Active Database Approach Person P-0001 Person Female Male P-0002 P-0001
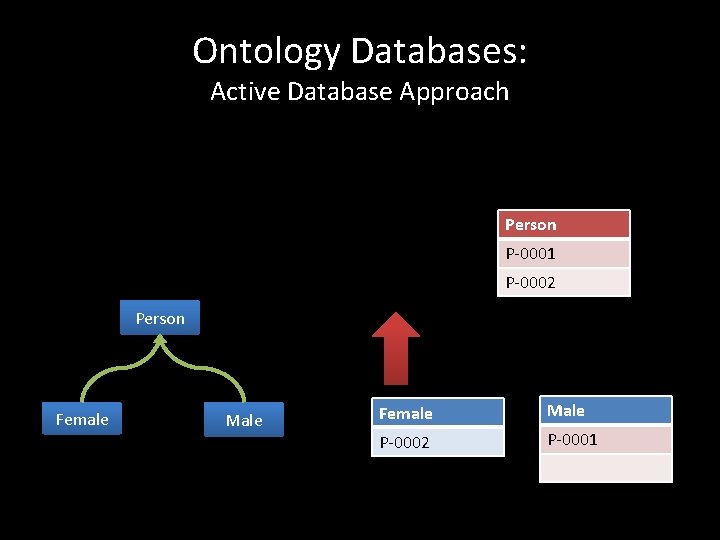
Ontology Databases: Active Database Approach Person P-0001 P-0002 Person Female Male P-0002 P-0001
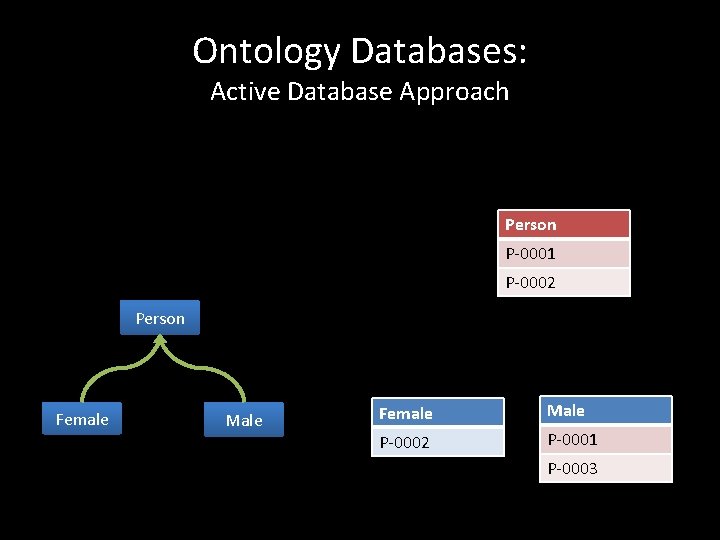
Ontology Databases: Active Database Approach Person P-0001 P-0002 Person Female Male P-0002 P-0001 P-0003

Ontology Databases: Active Database Approach Person P-0001 P-0002 P-0003 Person Female Male P-0002 P-0001 P-0003
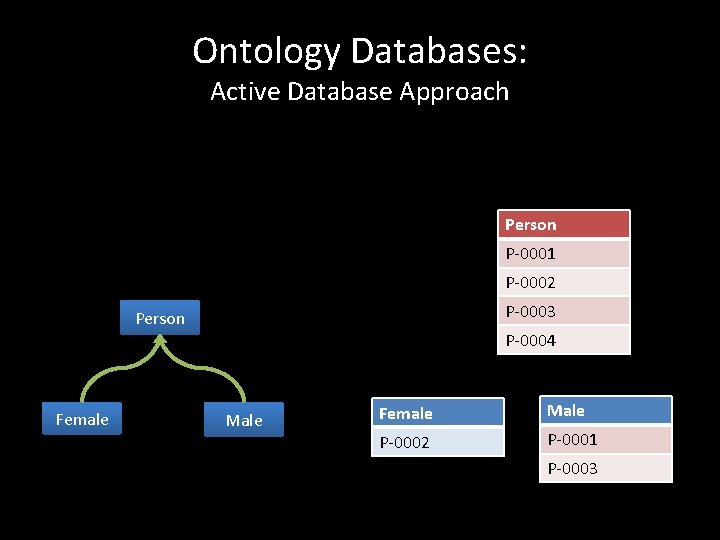
Ontology Databases: Active Database Approach Person P-0001 P-0002 P-0003 Person Female P-0004 Male Female Male P-0002 P-0001 P-0003

Example: sisters-siblings (revisited) This is what we know : All sisters are siblings. Hilary and Lynn are sisters. This is what we want to know : Who are siblings? Obviously, the answer should be : Hilary and Lynn are siblings.

Example: sisters-siblings (revisited) This is what we know : Sibling. Of All sisters are siblings. Hilary and Lynn are sisters. This is what we want to know : Who are siblings? Obviously, the answer should be : Hilary and Lynn are siblings. Sister. Of Hilary Lynn

Example: sisters-siblings (revisited) This is what we know : All sisters are siblings. Hilary and Lynn are sisters. This is what we want to know : Who are siblings? Obviously, the answer should be : Hilary and Lynn are siblings. Sibling. Of Hilary Lynn Sister. Of Hilary Lynn

Example: sisters-siblings (revisited) This is what we know : All sisters are siblings. Hilary and Lynn are sisters. Sibling. Of Hilary Lynn Sister. Of Hilary Lynn This is what we want to know : Who are siblings? Obviously, the answer should be : Hilary and Lynn are siblings. { <x, y> | sibling. Of(x, y) }

Example: sisters-siblings (revisited) This is what we know : All sisters are siblings. Hilary and Lynn are sisters. Sibling. Of Hilary Lynn Sister. Of Hilary Lynn This is what we want to know : Who are siblings? { <x, y> | sibling. Of(x, y) } Just look it up! Obviously, the answer should be : Hilary and Lynn are siblings.

Example: sisters-siblings (revisited) This is what we know : All sisters are siblings. Hilary and Lynn are sisters. Sibling. Of Hilary Lynn Sister. Of Hilary Lynn This is what we want to know : Who are siblings? { <x, y> | sibling. Of(x, y) } Just look it up! Obviously, the answer should be : Hilary and Lynn are siblings. { <Hilary, Lynn> }
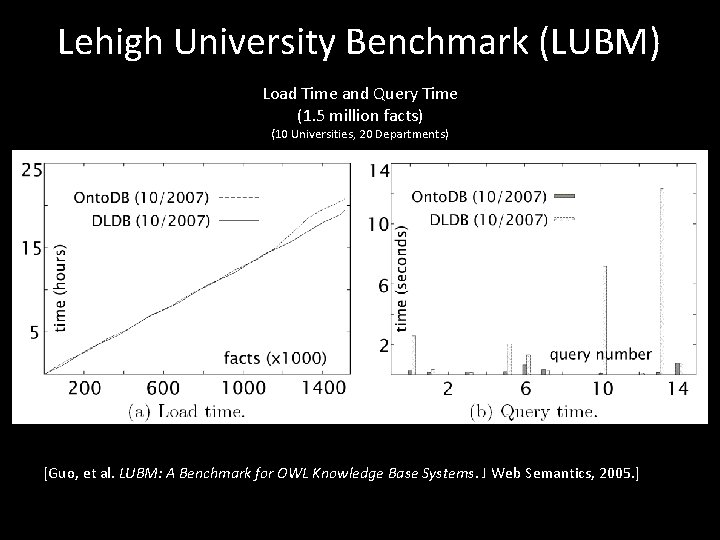
Lehigh University Benchmark (LUBM) Load Time and Query Time (1. 5 million facts) (10 Universities, 20 Departments) [Guo, et al. LUBM: A Benchmark for OWL Knowledge Base Systems. J Web Semantics, 2005. ]
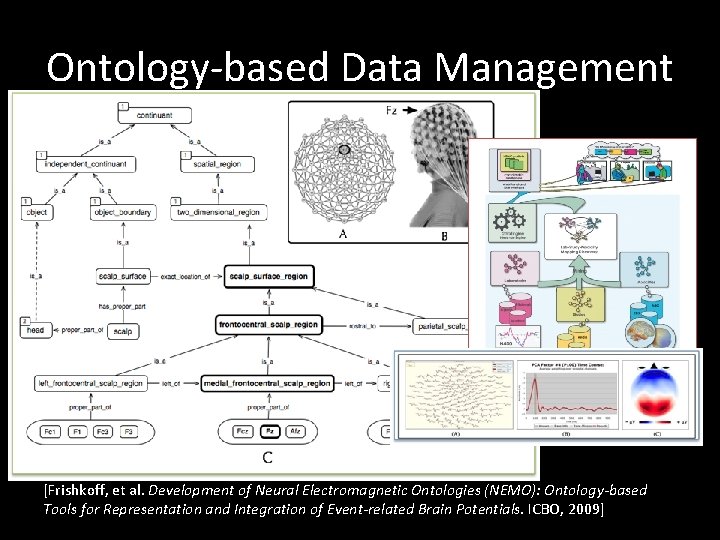
Ontology-based Data Management [Frishkoff, et al. Development of Neural Electromagnetic Ontologies (NEMO): Ontology-based Tools for Representation and Integration of Event-related Brain Potentials. ICBO, 2009]

Ontology-based Query Answering Return all data instances that belong to ERP pattern classes which have a surface positivity over frontal regions of interest and are earlier than the N 400. Which patterns have a region of interest that is left-occipital and manifests between 220 and 300 ms? What is the range of intensity mean for the region of interest for N 100? Show the region of interest for all ERP patterns that occur between 0 and 300 ms. Which PCA factor do P 100 patterns most often appear in? What is the range of intensity mean for the region of interest for N 100 patterns? Show the patterns whose region of interest is left occipital and occurs between 220 and 300 ms.

Inconsistency Detection • Background and Motivation – Expressiveness – From disjunctions to negations • Theory – Not-gadgets • Motivation – Serotonin example – ATP-gated cation channel activity • Results from ZFIN and MGI Annotations


Example: inconsistency detection "Annotations in this way sometimes point to errors in the type relationships described in the ontology. An example is the recent removal of the type serotonin secretion as an is_a child of neurotransmitter secretion from the GO Biological Process ontology. This modification was made as a result of an annotation from a paper showing that serotonin can be secreted by cells of the immune system where it does not act as a neurotransmitter. “ [Hill, et al. Gene Ontology annotations: what they mean and where they come from. BMC Bioinformatics, 2008]

Example: serotonin secretion gene-x not-gadget fail! gene-x
![Example: GO: 0004931 ATP-gated cation channel activity (as of 3/09): [Term] id: GO: 0004931 Example: GO: 0004931 ATP-gated cation channel activity (as of 3/09): [Term] id: GO: 0004931](http://slidetodoc.com/presentation_image_h/b2c3c500aaa61fdeb17f6bd4dc1e632b/image-36.jpg)
Example: GO: 0004931 ATP-gated cation channel activity (as of 3/09): [Term] id: GO: 0004931 name: ATP-gated cation channel activity namespace: molecular_function def: "Catalysis of the transmembrane transfer of an ion by a channel that opens when extracellular ATP has been bound by the channel complex or one of its constituent parts. " [GOC: mah, PMID: 9755289] comment: Note that this term refers to an activity and not a gene product. Consider also annotating to the molecular function term 'purinergic nucleotide receptor activity ; GO: 0001614'. synonym: "P 2 X activity" RELATED [] synonym: "purinoceptor" BROAD [] synonym: "purinoreceptor" BROAD [] is_a: GO: 0005231 ! excitatory extracellular ligand-gated ion channel activity is_a: GO: 0005261 ! cation channel activity

Example: GO: 0004931 GO: 0004391 sub-graph (using Jambalaya):

Example: GO: 0004931 What is so interesting about GO: 0004391? ZFIN ZDB-GENE-030319 -2 p 2 rx 2 NOT GO: 0004931 ZFIN: ZDB-PUB -031031 -8|PMID: 14580944 IDA F purinergic receptor P 2 X, ligand-gated ion channel, 2 gene taxon: 7955 20071005 ZFIN ZDB-GENE-030319 -2 p 2 rx 2 GO: 0004931 ZFIN: ZDB-PUB -031031 -8|PMID: 14580944 IGI ZFIN: ZDB-GENE-000427 -3 F purinergic receptor P 2 X, ligand-gated ion channel, 2 gene taxon: 7955 20071005 ZFIN Source: [1/13/2009] http: //www. geneontology. org/gene-associations/

Example: GO: 0004931 The not-gadget will raise a logical inconsistency. p 2 rx 2 NOTGO: 0004931 p 2 rx 2 GO: 0004931 GO_0004931 _GO_0004931 p 2 rx 2 not-gadget fail! * Tables starting with an '_' are negations.
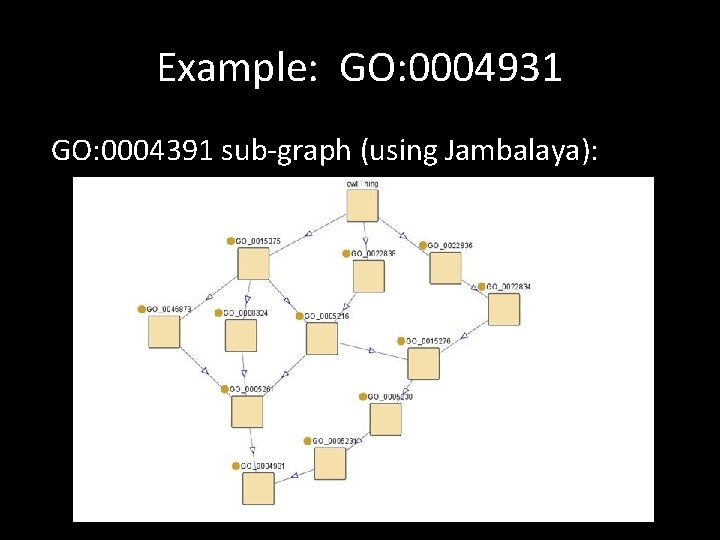
Example: GO: 0004931 GO: 0004391 sub-graph (using Jambalaya):
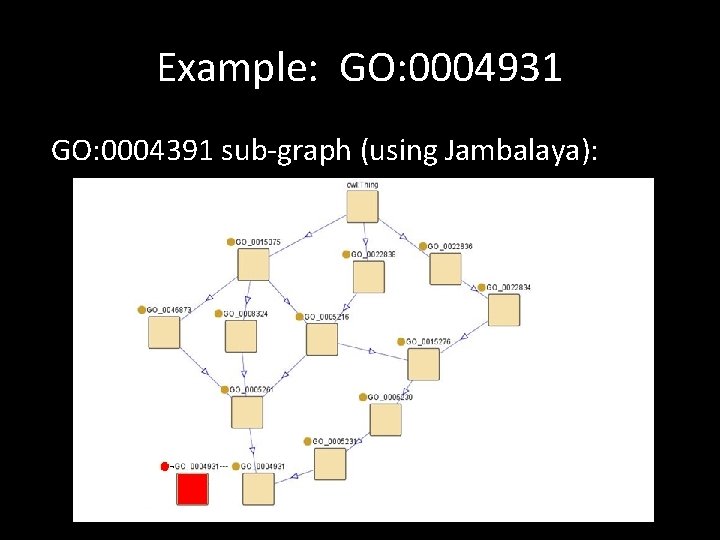
Example: GO: 0004931 GO: 0004391 sub-graph (using Jambalaya):

Example: GO: 0004931 GO: 0004391 sub-graph (using Jambalaya):
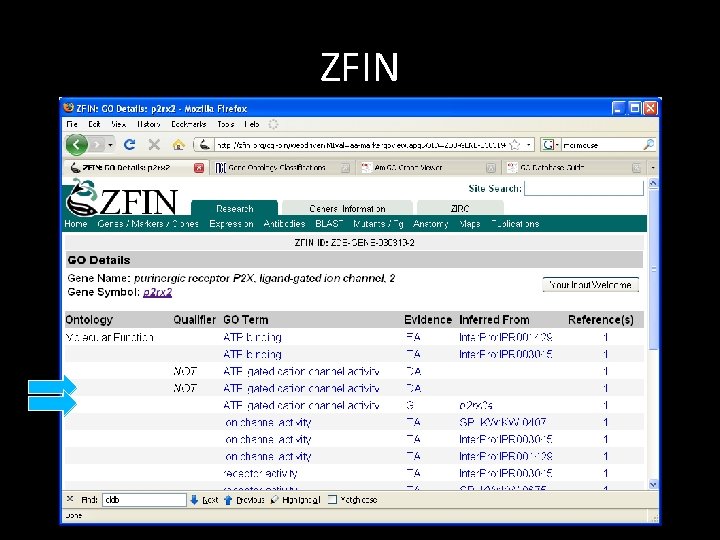
ZFIN
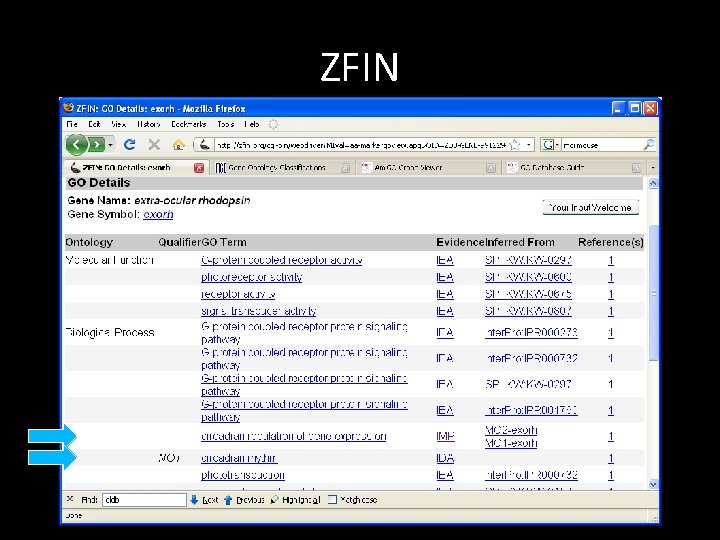
ZFIN
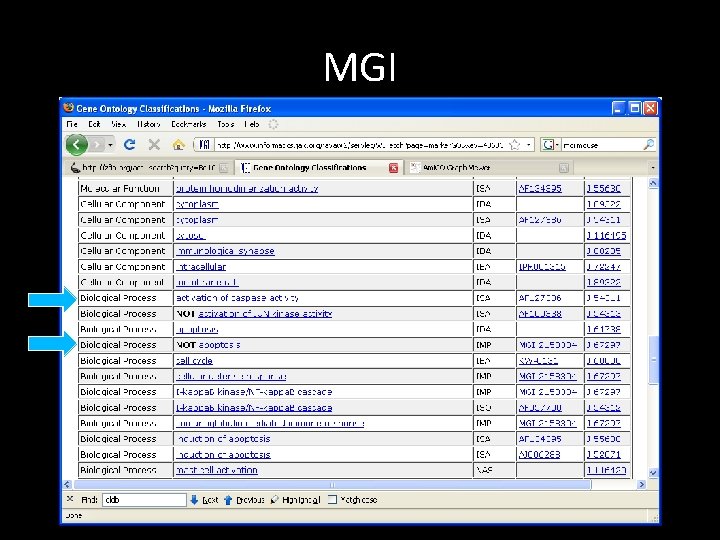
MGI
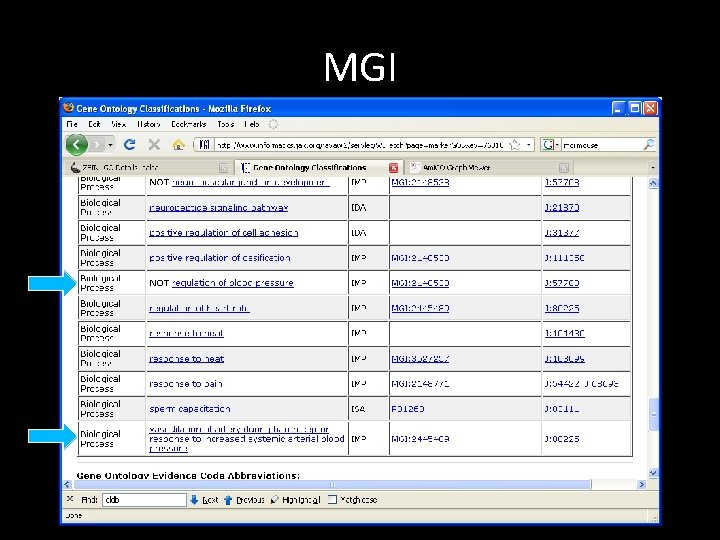
MGI
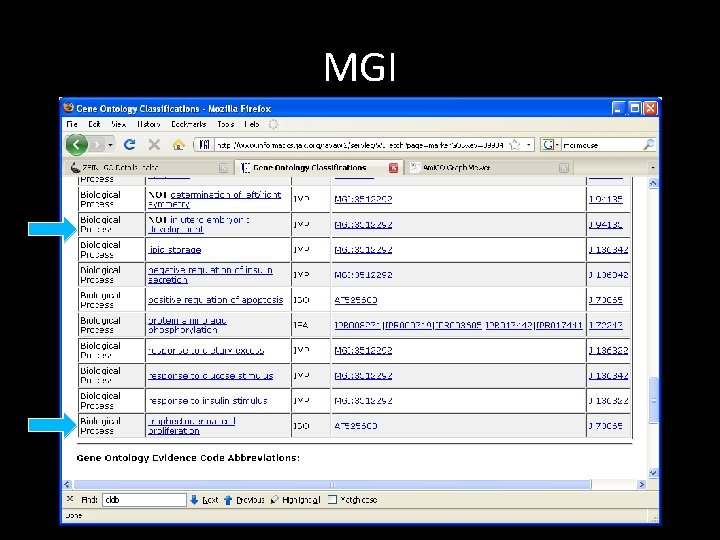
MGI
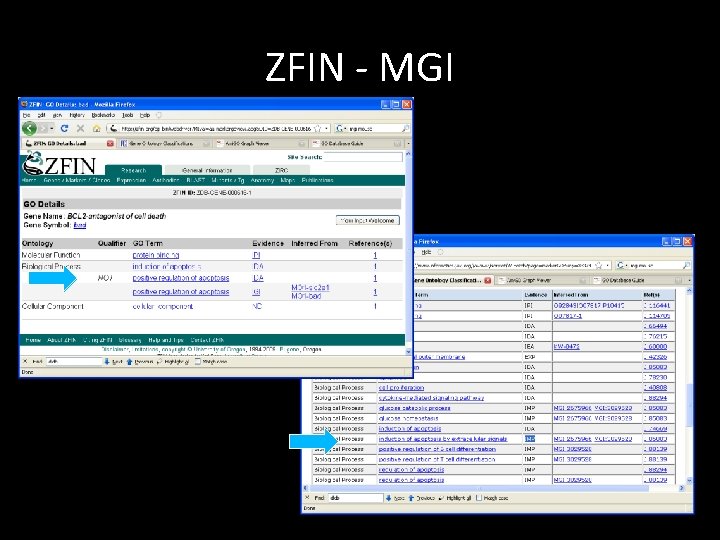
ZFIN - MGI

ZFIN
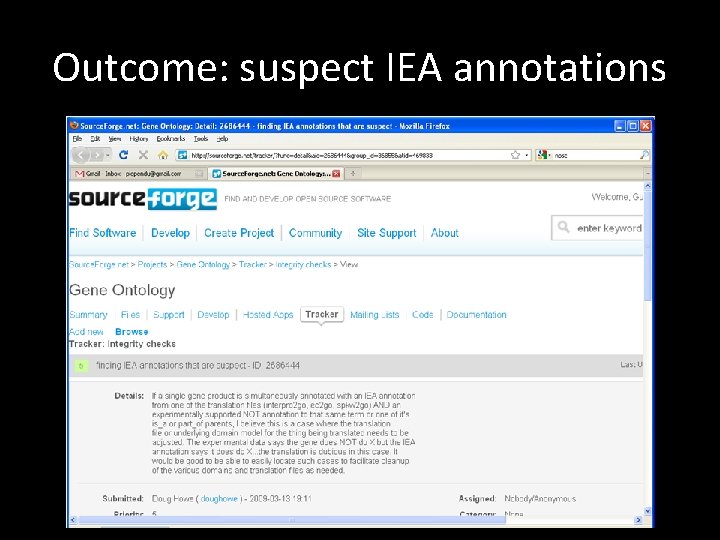
Outcome: suspect IEA annotations
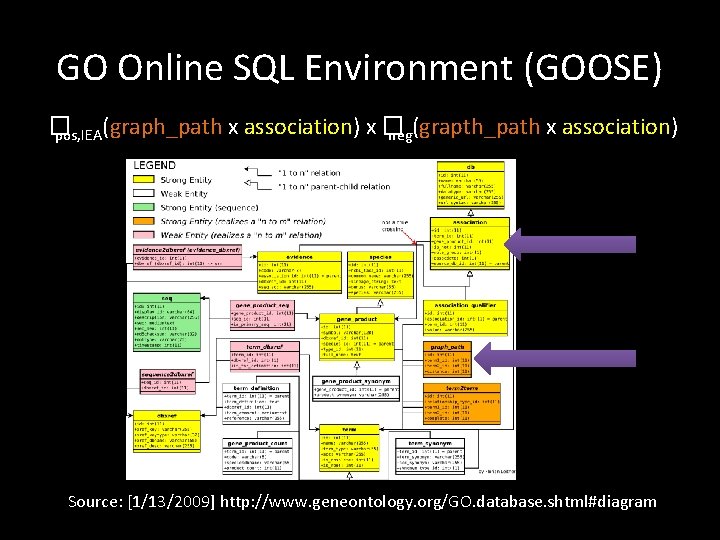
GO Online SQL Environment (GOOSE) � pos, IEA(graph_path x association) x � neg(grapth_path x association) Source: [1/13/2009] http: //www. geneontology. org/GO. database. shtml#diagram

What do logical inconsistencies mean? • Several possibilities: – Incorrect annotation (e. g. , suspect IEA annotations) – Incorrect relationship (e. g. , serotonin secretion) – Incomplete model: Recall: ZFIN ZDB-GENE-030319 -2 p 2 rx 2 GO: 0004931 ZFIN: ZDB-PUB-031031 -8|PMID: 14580944 IGI ZFIN: ZDB-GENE-000427 -3 F purinergic receptor P 2 X, ligand-gated ion channel, 2 gene taxon: 7955 20071005 ZFIN – Perfectly admissible!

Next Directions • Explanation and proof-reconstruction ✓ • Deep (data) annotation tools • Distributed network of Ontology Databases

Data Annotation: Neural Electro. Magnetic Ontologies frontocentral LFRON RFRON [Frishkoff, et al. ERP measures of partial semantic knowledge: Left temporal indices of skill differences and lexical quality. Biological Psychology, 2009. ]
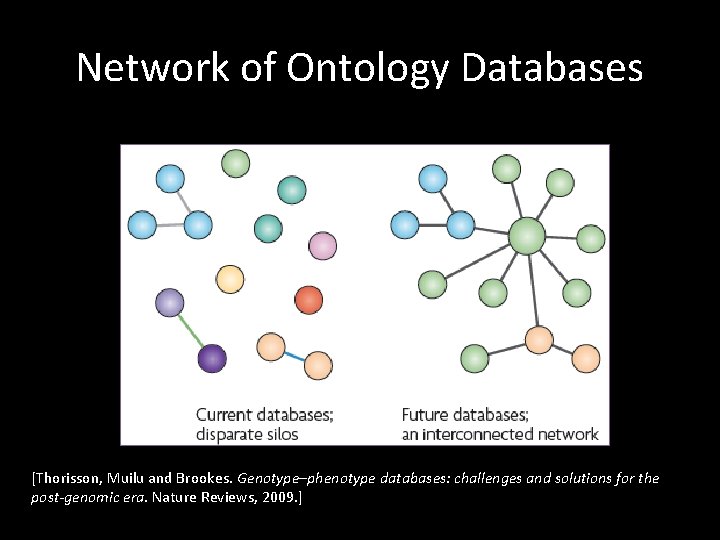
Network of Ontology Databases [Thorisson, Muilu and Brookes. Genotype–phenotype databases: challenges and solutions for the post-genomic era. Nature Reviews, 2009. ]

Thank you Questions?

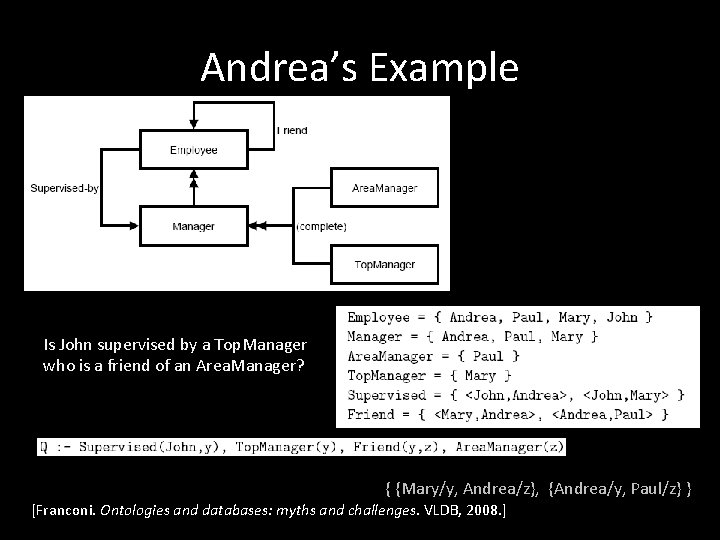
Andrea’s Example Is John supervised by a Top. Manager who is a friend of an Area. Manager? { {Mary/y, Andrea/z}, {Andrea/y, Paul/z} } [Franconi. Ontologies and databases: myths and challenges. VLDB, 2008. ]
![Raymond Reiter [Reiter. Deductive Question-Answering on Relational Data Bases. Logic and Data Bases, 1977] Raymond Reiter [Reiter. Deductive Question-Answering on Relational Data Bases. Logic and Data Bases, 1977]](http://slidetodoc.com/presentation_image_h/b2c3c500aaa61fdeb17f6bd4dc1e632b/image-59.jpg)
Raymond Reiter [Reiter. Deductive Question-Answering on Relational Data Bases. Logic and Data Bases, 1977]

Raymond Reiter
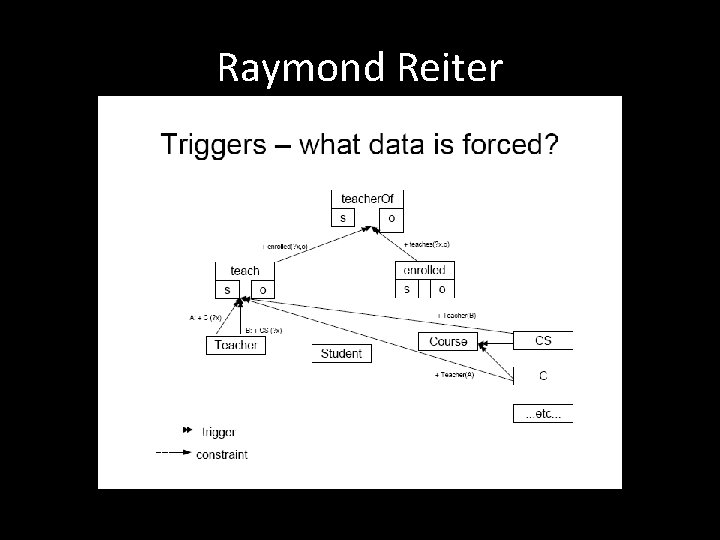
Raymond Reiter
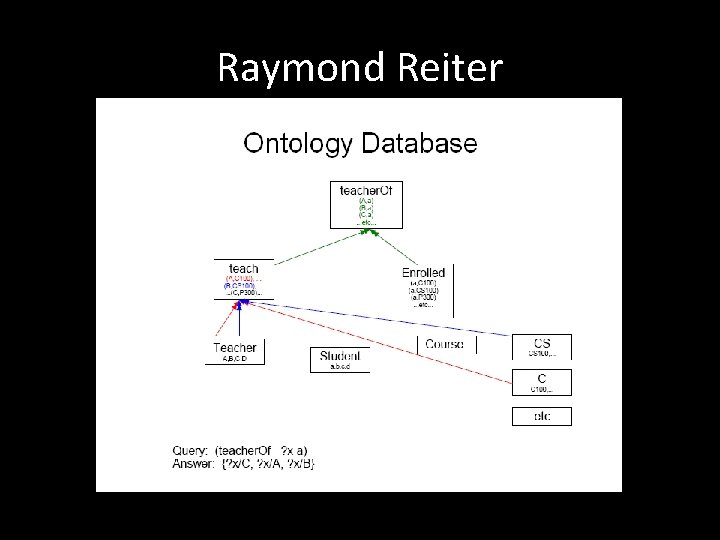
Raymond Reiter
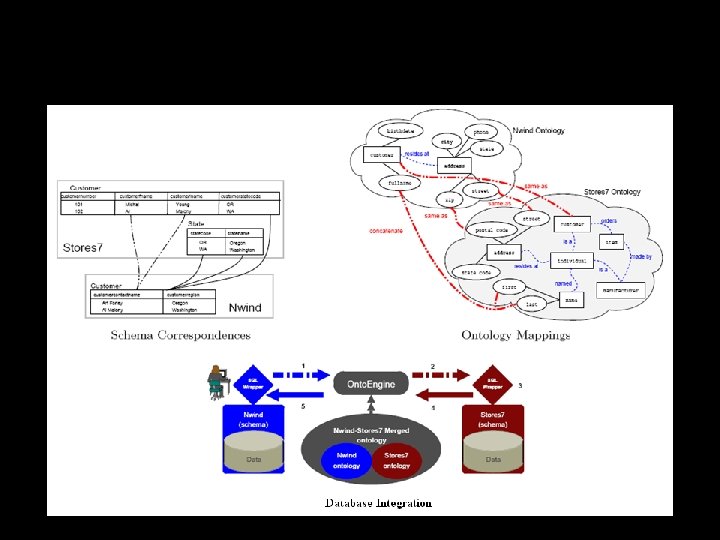

Benchmarking Suite
Origins
CIS @ UO

CIS @ UO • Research Areas in Computer Science: – – – – software engineering programming languages human-computer interaction parallel and distributed computing networking and graph theory scientific computation/visualization information integration and mining • Affiliates: – Neurosciences Institute – Computational Science Institute – Zebrafish Information Network

Ontology-based Data Access [Rodriguez-Muro , et al. Realizing Ontology Based Data Access: A plug-in for protégé. ICDEW, 2008. ]
- Slides: 68