Universitt Bielefeld Graduiertenkolleg Bioinformatik Multiple Genome Alignment Chaining

Universität Bielefeld Graduiertenkolleg Bioinformatik Multiple Genome Alignment: Chaining Algorithms Revisited Mohamed I. Abouelhoda University of Bielefeld Joint work with Enno Ohlebusch University of Ulm Germany 2003 1

Universität Bielefeld Graduiertenkolleg Bioinformatik Comparative Genomics The practice of analysing the genomic material of a species by comparing it with the genomic material of other species. Why is this important? Ø The next logical step after the high throughput sequencing projects. Ø Deducing the mechanism and history of genome evolution. Ø Discovering genes and regulatory elements. Ø Identifying exons in eukaryotic genes. ØRevealing the role of non-coding conserved sequences. 2
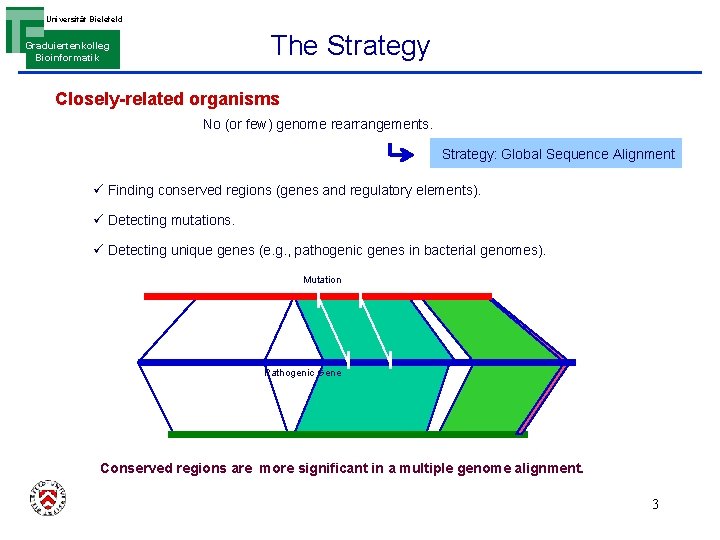
Universität Bielefeld The Strategy Graduiertenkolleg Bioinformatik Closely-related organisms No (or few) genome rearrangements. Strategy: Global Sequence Alignment ü Finding conserved regions (genes and regulatory elements). ü Detecting mutations. ü Detecting unique genes (e. g. , pathogenic genes in bacterial genomes). Mutation Pathogenic Gene Conserved regions are more significant in a multiple genome alignment. 3

Universität Bielefeld Graduiertenkolleg Bioinformatik Multiple Genome Alignment is Difficult Ø Given k genomes, each of average length N Ø Standard dynamic programming takes O(N k) N is very large inpractical Mega bases even for k=2 Ø Heuristic algorithms are therefore employed Program Year Authors Two Genomes Pip. Maker 2000 Schwartz et al. DIALIGN 2000 Morgenstern MUMmer 2002 Delcher et al. CHAOS 2002 Brudno and Morgenstern OWEN 2002 Roytberg et al. AVID 2003 Bray et al. Multiple Genomes MGA 2002 Höhl et al. These tools use anchor-based alignment method 4

Universität Bielefeld Graduiertenkolleg Bioinformatik MGA uses a strategy composed of three steps: 1. Computation of fragments (maximal multiple exact matches). 2. Computation of an optimal chain of colinear non-overlapping fragments. 3. Detailed alignment of the regions between the fragments of the optimal chain. First Genome G 1 Second Genome G 2 Third Genome G 3 5
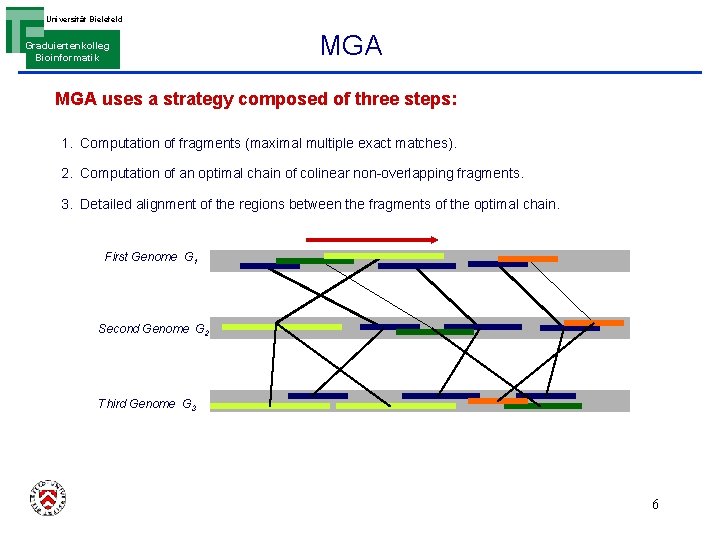
Universität Bielefeld Graduiertenkolleg Bioinformatik MGA uses a strategy composed of three steps: 1. Computation of fragments (maximal multiple exact matches). 2. Computation of an optimal chain of colinear non-overlapping fragments. 3. Detailed alignment of the regions between the fragments of the optimal chain. First Genome G 1 Second Genome G 2 Third Genome G 3 6
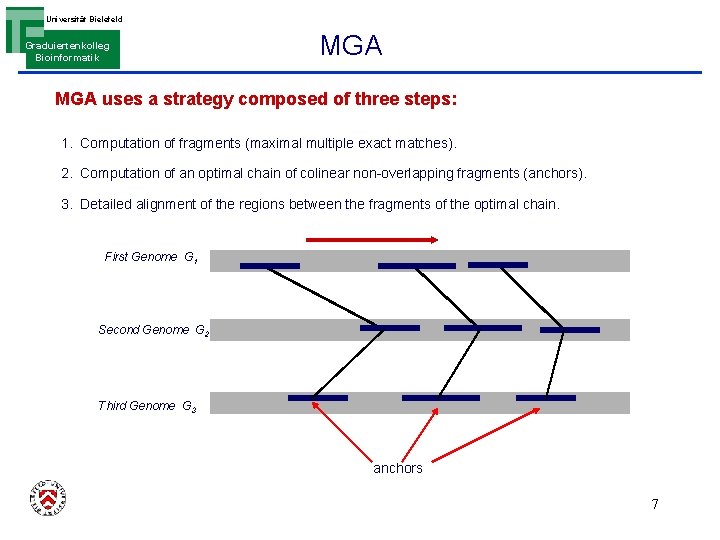
Universität Bielefeld Graduiertenkolleg Bioinformatik MGA uses a strategy composed of three steps: 1. Computation of fragments (maximal multiple exact matches). 2. Computation of an optimal chain of colinear non-overlapping fragments (anchors). 3. Detailed alignment of the regions between the fragments of the optimal chain. First Genome G 1 Second Genome G 2 Third Genome G 3 anchors 7
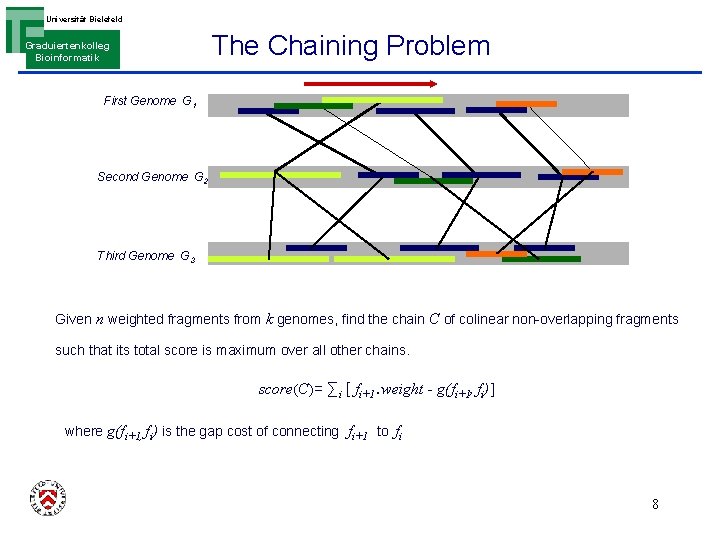
Universität Bielefeld Graduiertenkolleg Bioinformatik The Chaining Problem First Genome G 1 Second Genome G 2 Third Genome G 3 Given n weighted fragments from k genomes, find the chain C of colinear non-overlapping fragments such that its total score is maximum over all other chains. score(C)= ∑i [ fi+1. weight - g(fi+1, fi)] where g(fi+1, fi) is the gap cost of connecting fi+1 to fi 8
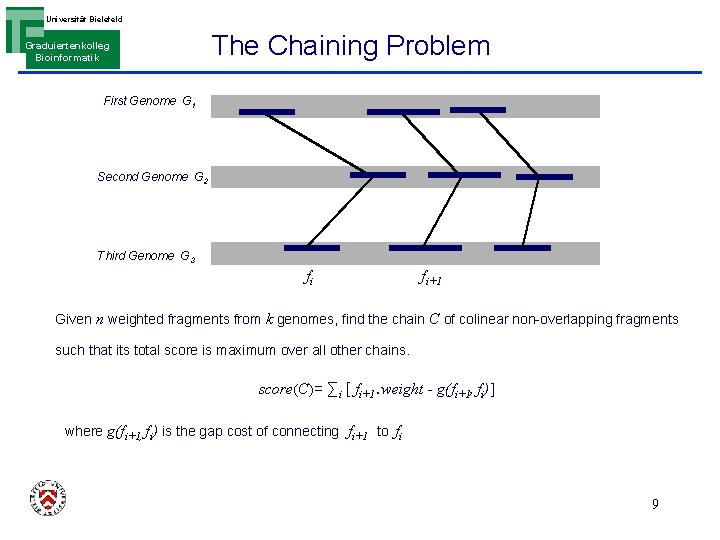
Universität Bielefeld Graduiertenkolleg Bioinformatik The Chaining Problem First Genome G 1 Second Genome G 2 Third Genome G 3 fi fi+1 Given n weighted fragments from k genomes, find the chain C of colinear non-overlapping fragments such that its total score is maximum over all other chains. score(C)= ∑i [ fi+1. weight - g(fi+1, fi)] where g(fi+1, fi) is the gap cost of connecting fi+1 to fi 9
Universität Bielefeld Graduiertenkolleg Bioinformatik Previous Work ü Graph based solution takes O(n 2) time. Soble-Martinez, 1986 Wilbur-Lipman, 1983 ü Geometric based algorithm is subquadratic (sparse dynamic programming): Eppstein-Giancarlo, 1992 1. Zhang et al. (1994) used space division with a kd-tree (no complexity analysis was given). 2. Myers and Miller (1995) used orthogonal range search with a range tree yielding a complexity of O(n log k n) time and O(n log k-1 n) space. But the result is a time bound higher by a logarithmic factor than what one would expect. David Eppstein 10

Universität Bielefeld Previous Work Graduiertenkolleg Bioinformatik ü For two genomes the complexities are also higher than those of known 2 -dim. chaining algorithms O(n log 2 n) time and O(n log n) space. We thought hard to reduce this discrepancy but have been unable to do so and the reasons appear to be fundamental ! To improve upon our result appears to be a difficult open problem ! Myers & Miller ü Here, it is improved by almost two log factors in time and one log factor in space For k>2 For k=2 O(n log k-2 n log n) time and O(n log k-2 n) space. O(n log n) time and O(n) space. 11
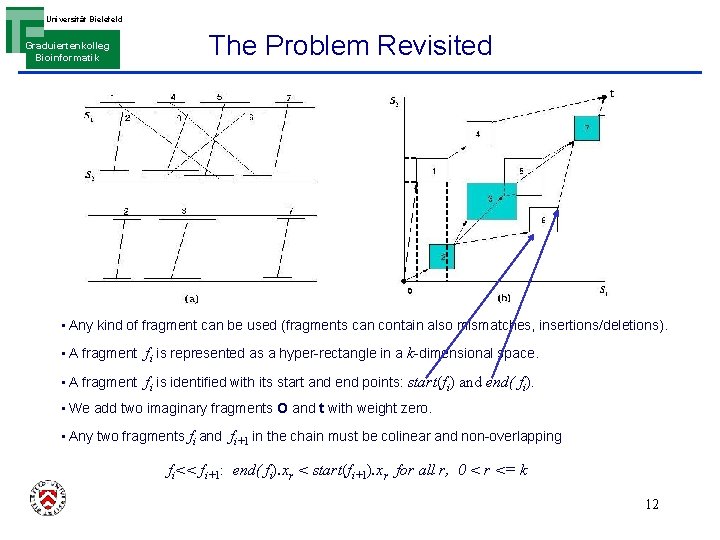
Universität Bielefeld Graduiertenkolleg Bioinformatik The Problem Revisited • Any kind of fragment can be used (fragments can contain also mismatches, insertions/deletions). • A fragment fi is represented as a hyper-rectangle in a k-dimensional space. • A fragment fi is identified with its start and end points: start(fi) and end( fi). • We add two imaginary fragments O and t with weight zero. • Any two fragments fi and fi+1 in the chain must be colinear and non-overlapping fi<< fi+1: end( fi). xr < start(fi+1). xr for all r, 0 < r <= k 12
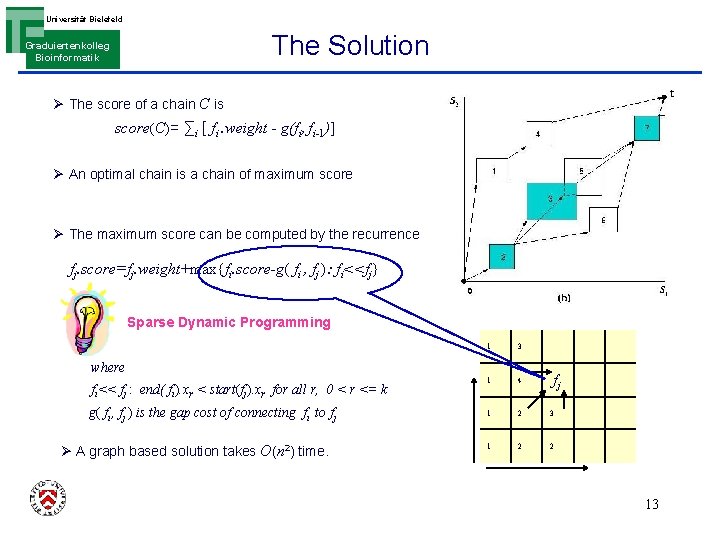
Universität Bielefeld The Solution Graduiertenkolleg Bioinformatik Ø The score of a chain C is score(C)= ∑i [ fi. weight - g(fi, fi-1)] Ø An optimal chain is a chain of maximum score Ø The maximum score can be computed by the recurrence fj. score=fj. weight+max{fi. score-g( fi , fj ): fi<<fj} Sparse Dynamic Programming 1 3 1 4 1 2 3 1 2 2 where fi<< fj : end( fi). xr < start(fj). xr for all r, 0 < r <= k g( fi , fj ) is the gap cost of connecting fi to fj Ø A graph based solution takes O(n 2) time. fj 13
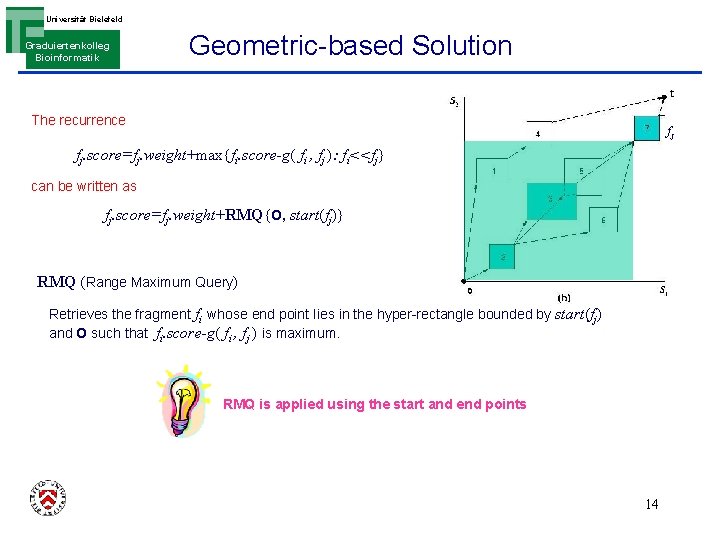
Universität Bielefeld Graduiertenkolleg Bioinformatik Geometric-based Solution The recurrence fj fj. score=fj. weight+max{fi. score-g( fi , fj ): fi<<fj} can be written as fj. score=fj. weight+RMQ{O, start(fj)} RMQ (Range Maximum Query) Retrieves the fragment fi whose end point Iies in the hyper-rectangle bounded by start(fj) and O such that fi. score-g( fi , fj ) is maximum. RMQ is applied using the start and end points 14
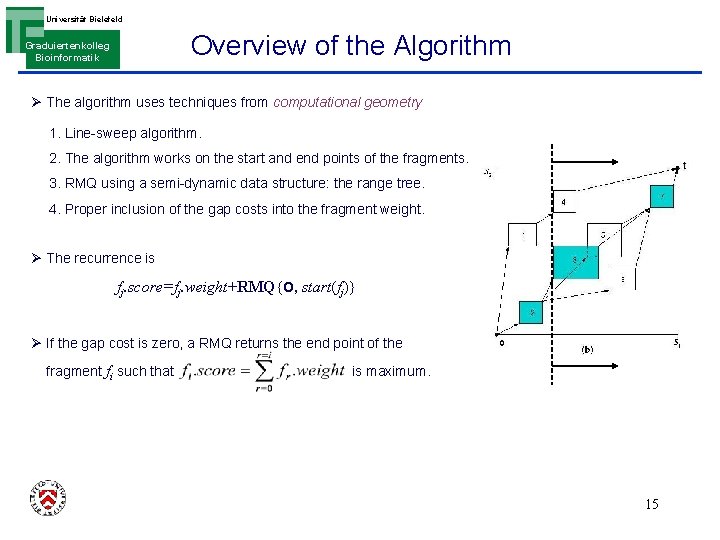
Universität Bielefeld Overview of the Algorithm Graduiertenkolleg Bioinformatik Ø The algorithm uses techniques from computational geometry 1. Line-sweep algorithm. 2. The algorithm works on the start and end points of the fragments. 3. RMQ using a semi-dynamic data structure: the range tree. 4. Proper inclusion of the gap costs into the fragment weight. Ø The recurrence is fj. score=fj. weight+RMQ{O, start(fj)} Ø If the gap cost is zero, a RMQ returns the end point of the fragment fi such that is maximum. 15
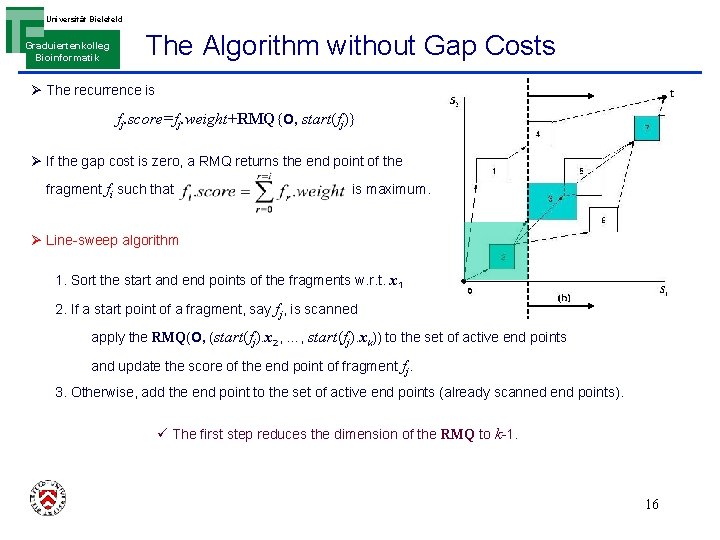
Universität Bielefeld Graduiertenkolleg Bioinformatik The Algorithm without Gap Costs Ø The recurrence is fj. score=fj. weight+RMQ{O, start(fj)} Ø If the gap cost is zero, a RMQ returns the end point of the fragment fi such that is maximum. Ø Line-sweep algorithm 1. Sort the start and end points of the fragments w. r. t. x 1 2. If a start point of a fragment, say fj, is scanned apply the RMQ(O, (start(fj). x 2, …, start(fj). xk)) to the set of active end points and update the score of the end point of fragment fj. 3. Otherwise, add the end point to the set of active end points (already scanned end points). ü The first step reduces the dimension of the RMQ to k-1. 16

Universität Bielefeld Graduiertenkolleg Bioinformatik The Complexity of the Algorithm Ø The complexity of the algorithm depends on the complexity of the RMQ in d= k-1 dimensions Ø The required data structure D to manipulate the set of points • is a semi-dynamic data structure over all end points in d= k-1 dimensions • supports the operations: 1. Activate an end point. 2. Perform a RMQ. 1. supported by fractional cascading. Ø D is implemented as a range tree 2. enhanced with priority queues. Willard, 1985 Johnson, 1982 van Emde Boas, 1977 Ø For n fragments and dimension d, the RMQ and activation takes: O(n log d-1 n log n) time and O(n log d-1 n) space Ø Since d= k-1>1, the complexity of the algorithm is O(n log k-2 n log n) time and O(n log k-2 n) space Ø For k=2, the total complexity is O(n log n) time and O(n) space 17

Universität Bielefeld Graduiertenkolleg Bioinformatik Including Gap Costs How to define the gap costs ? How to include the gap costs without affecting the complexity? Recall the recurrence fj. score=fj. weight+max{fi. score-g( fi , fj ): fi<<fj} The gap cost should be included in the RMQ, otherwise the algorithm would be quadratic. A C C XXX A C C fj. score=fj. weight+RMQ{O, start(fj)} f ACCYYACC 18
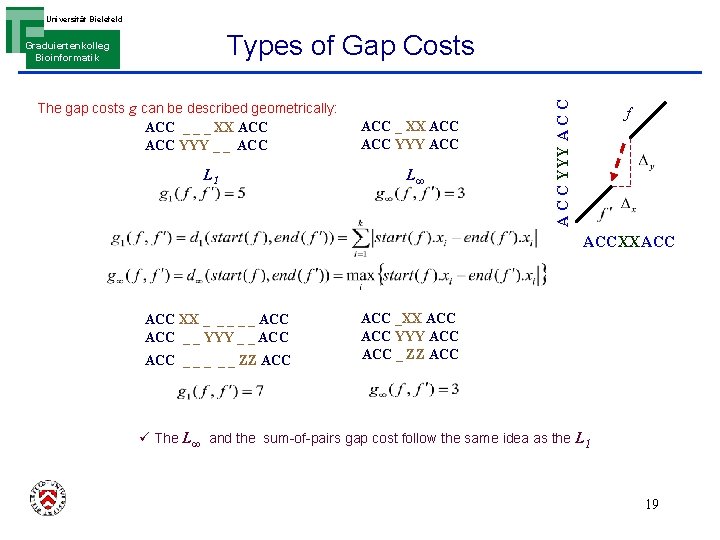
Universität Bielefeld The gap costs g can be described geometrically: ACC _ _ _ XX ACC YYY _ _ ACC L 1 ACC _ XX ACC YYY ACC L∞ A C C YYY A C C Types of Gap Costs Graduiertenkolleg Bioinformatik f ACCXXACC XX _ _ _ ACC _ _ YYY _ _ ACC _ _ _ ZZ ACC _XX ACC YYY ACC _ ZZ ACC ü The L∞ and the sum-of-pairs gap cost follow the same idea as the L 1 19
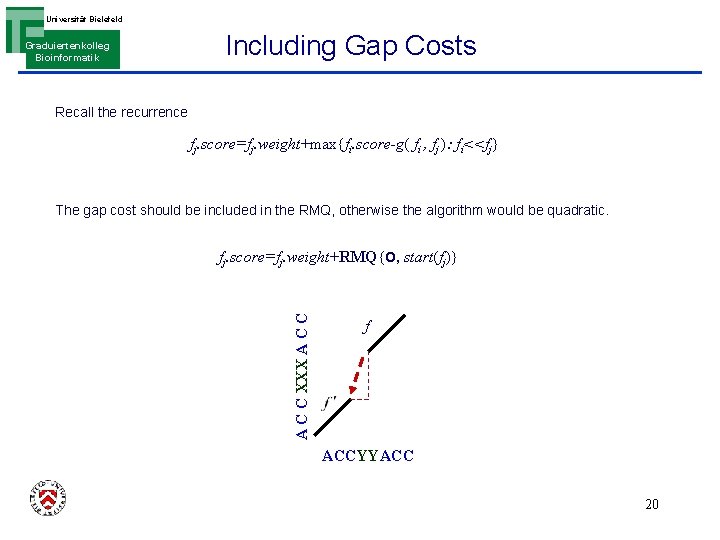
Universität Bielefeld Graduiertenkolleg Bioinformatik Including Gap Costs Recall the recurrence fj. score=fj. weight+max{fi. score-g( fi , fj ): fi<<fj} The gap cost should be included in the RMQ, otherwise the algorithm would be quadratic. A C C XXX A C C fj. score=fj. weight+RMQ{O, start(fj)} f ACCYYACC 20
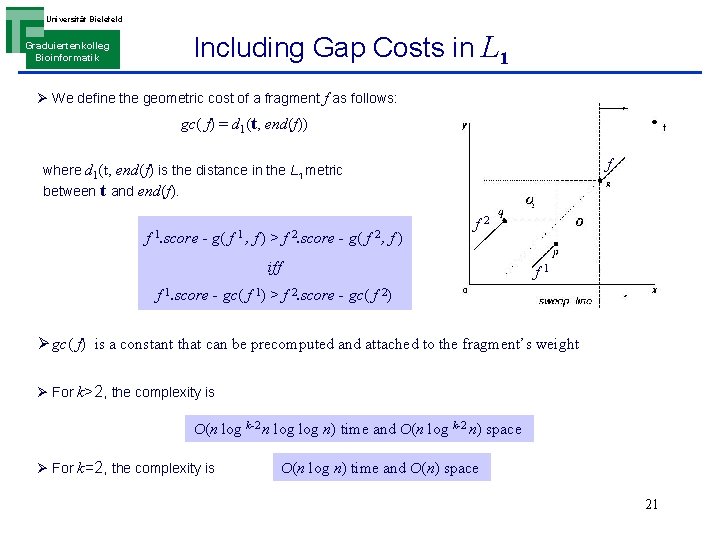
Universität Bielefeld Including Gap Costs in L 1 Graduiertenkolleg Bioinformatik Ø We define the geometric cost of a fragment f as follows: gc( f) = d 1(t, end(f)) f where d 1(t, end(f) is the distance in the L 1 metric between t and end(f). f 1. score - g( f 1, f) > f 2. score - g( f 2, f) f 2 iff f 1. score - gc( f 1) > f 2. score - gc( f 2) Ø gc( f) is a constant that can be precomputed and attached to the fragment’s weight Ø For k>2, the complexity is O(n log k-2 n log n) time and O(n log k-2 n) space Ø For k=2, the complexity is O(n log n) time and O(n) space 21
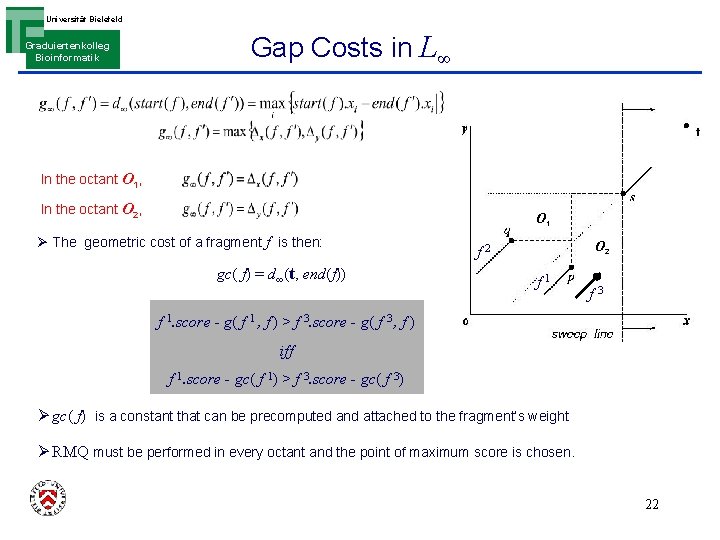
Universität Bielefeld Graduiertenkolleg Bioinformatik Gap Costs in L∞ In the octant O 1, In the octant O 2, O 1 Ø The geometric cost of a fragment f is then: gc( f) = d∞(t, end(f)) O 2 f 1 f 3 f 1. score - g( f 1 , f ) > f 3. score - g( f 3 , f ) iff f 1. score - gc( f 1) > f 3. score - gc( f 3) Ø gc( f) is a constant that can be precomputed and attached to the fragment’s weight Ø RMQ must be performed in every octant and the point of maximum score is chosen. 22

Universität Bielefeld RMQ on Octants Graduiertenkolleg Bioinformatik Ø Because the RMQ requires an orthogonal range, we use the octant-to-quadrant transformation: For the octant O 1 For the octant O 2 s O 1 q O 2 p s q p Ø The total complexity of the algorithm depends on the space divison. Ø For k>2, the complexity is O(k! n log k-2 n log n) time and O(k! n log k-2 n) space Ø For k=2, the complexity is O(2 n log n) time and O(2 n) space 23
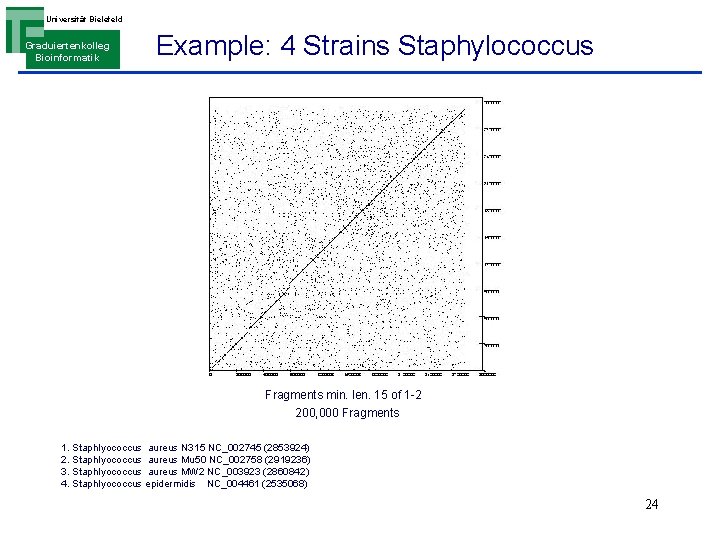
Universität Bielefeld Graduiertenkolleg Bioinformatik Example: 4 Strains Staphylococcus Fragments min. len. 15 of 1 -2 200, 000 Fragments 1. Staphlyococcus aureus N 315 NC_002745 (2853924) 2. Staphlyococcus aureus Mu 50 NC_002758 (2919236) 3. Staphlyococcus aureus MW 2 NC_003923 (2860842) 4. Staphlyococcus epidermidis NC_004461 (2535068) 24
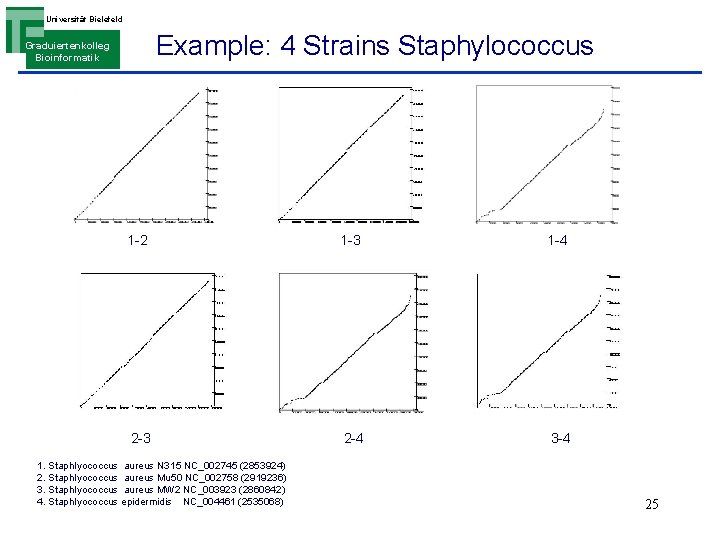
Universität Bielefeld Example: 4 Strains Staphylococcus Graduiertenkolleg Bioinformatik 1 -2 1 -3 1 -4 2 -3 2 -4 3 -4 1. Staphlyococcus aureus N 315 NC_002745 (2853924) 2. Staphlyococcus aureus Mu 50 NC_002758 (2919236) 3. Staphlyococcus aureus MW 2 NC_003923 (2860842) 4. Staphlyococcus epidermidis NC_004461 (2535068) 25
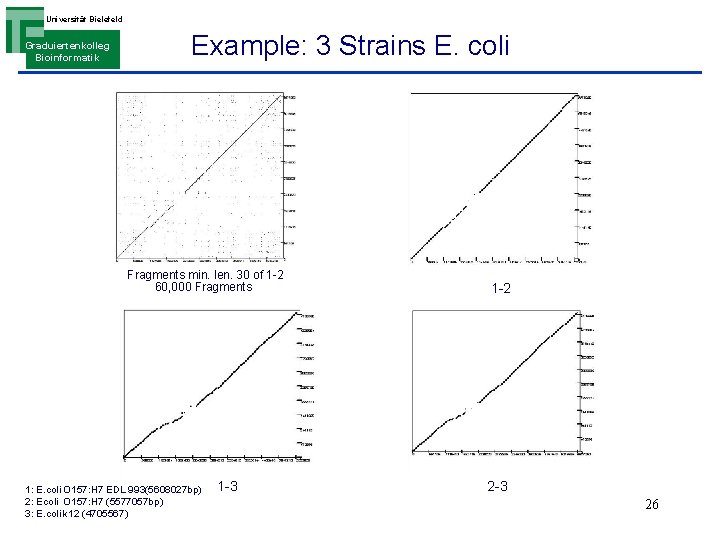
Universität Bielefeld Graduiertenkolleg Bioinformatik Example: 3 Strains E. coli Fragments min. len. 30 of 1 -2 60, 000 Fragments 1: E. coli O 157: H 7 EDL 993(5608027 bp) 2: Ecoli O 157: H 7 (5577057 bp) 3: E. coli k 12 (4705567) 1 -3 1 -2 2 -3 26

Universität Bielefeld Graduiertenkolleg Bioinformatik Conclusions Ø Our algorithm solves an open problem w. r. t. the chaining of n fragments of k genomes. Ø We reduced the time complexity by almost two log factors and the space complexity by one log factor. Ø The sum-of-pairs gap cost is addressed in the paper. Ø Other data structures than the range tree can be used, e. g. , the kd-tree. It takes for k>2 time and O(n) space Ø Other research topic: Comparison of distantly-related organisms. Many genome rearrangements. ü Detecting rearranged segments. Local sequence alignment ü Revealing the mechanisms of rearrangements. ü Better identification of the exons of eukaryotic genes. Abouelhoda-Ohlebusch, WABI 2003 27
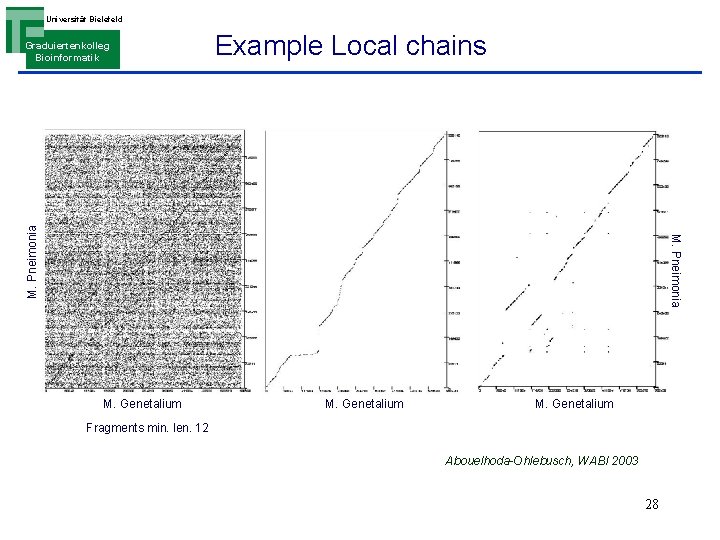
Universität Bielefeld Example Local chains M. Pneimonia Graduiertenkolleg Bioinformatik M. Genetalium Fragments min. len. 12 Abouelhoda-Ohlebusch, WABI 2003 28

Universität Bielefeld Graduiertenkolleg Bioinformatik Example Local chains V. cholera Fragments min. len. 15 E. coli 29

Universität Bielefeld Graduiertenkolleg Bioinformatik Acknowledgement Enno Ohlebusch, Ulm University Stefan Kurtz, , Hamburg University Robert Giegerich, Bielefeld University Janina Scholz, Bielefeld University This work was funded by the Graduiertenkolleg Bioinformatik, Universität Bielefeld, Germany. Thanks for your attention 30
- Slides: 30