Whole Genome Alignment MUMmer and Alignment October 2

Whole Genome Alignment MUMmer and Alignment October 2 nd, 2007 Adam M Phillippy amp@cs. umd. edu

Goal of WGA w For two genomes, A and B, find a mapping from each position in A to its corresponding position in B CCGGTAGGCTATTAAACGGGGTGAGGAGCGTTGGCATAGCA 41 bp genome CCGGTAGGCTATTAAACGGGGTGAGGAGCGTTGGCATAGCA

Not so fast. . . w Genome A may have insertions, deletions, translocations, inversions, duplications or SNPs with respect to B (sometimes all of the above) CCGGTAGGATATTAAACGGGGTGAGGAGCGTTGGCATAGCA CCGCTAGGCTATTAAAACCCCGGAGGAG. . GGCTGAGCA

Visualization w How can we visualize alignments? w With an identity plot n XY plot l l n n Let x = position in genome A Let y = %similarity of Ax to corresponding position in B Plot the identity function This can reveal islands of conservation, e. g. exons

Identity plot example

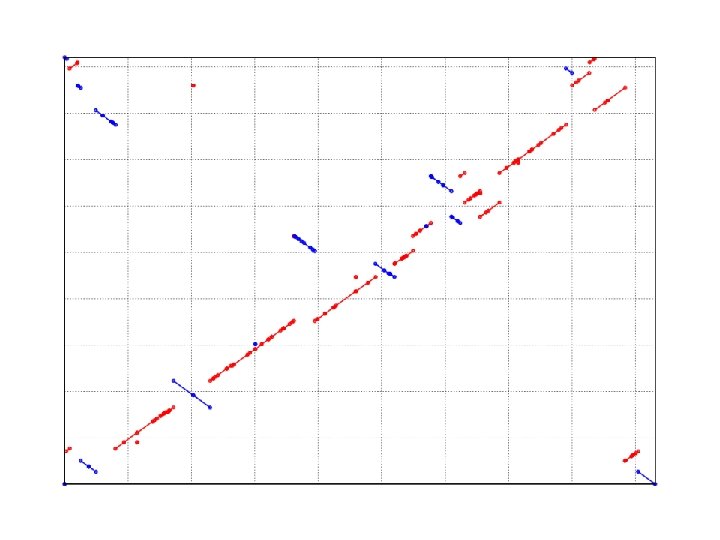
WGA visualization w How can we visualize whole genome alignments? w With an alignment dot plot n N x M matrix l l l n Let i = position in genome A Let j = position in genome B Fill cell (i, j) if Ai shows similarity to Bj A perfect alignment between A and B would completely fill the positive diagonal
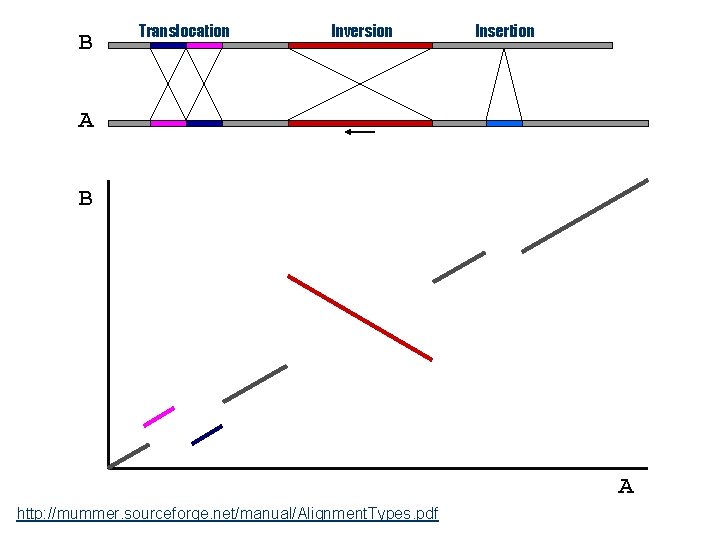
B Translocation Inversion Insertion A B A http: //mummer. sourceforge. net/manual/Alignment. Types. pdf
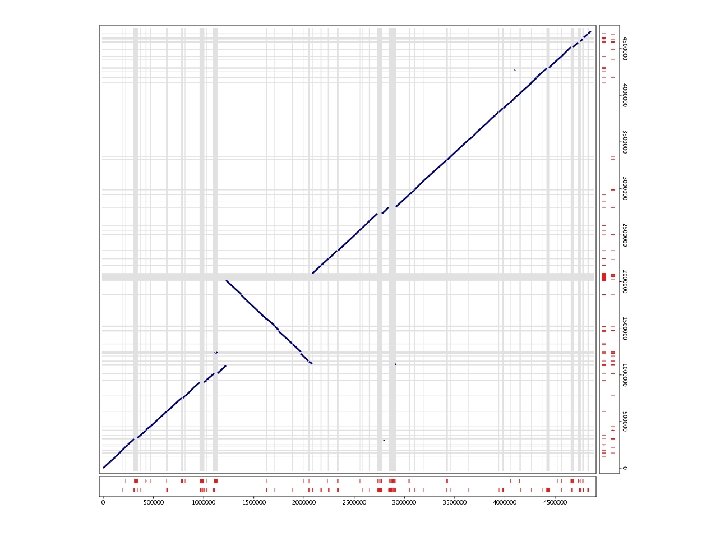

Global vs. Local w Global pairwise alignment. . . AAGCTTGGCTTAGCTGCTAGGGTAGGCTTGGG. . . AAGCTGGGCTTAGTTGCTAG. . TAGGCTTTGG. . . ^ ^ ^^ ^ w Whole genome alignment Often impossible to represent as a global alignment l We will assume a set of local alignments l w This works great for draft sequence
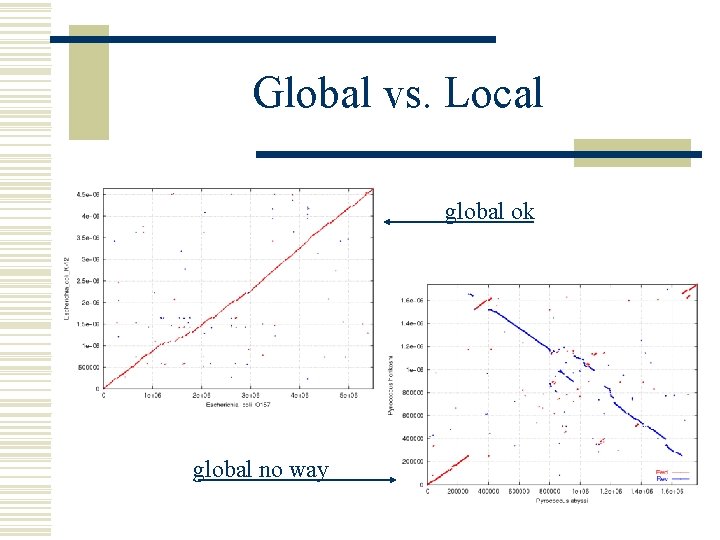
Global vs. Local global ok global no way

Alignment tools w Whole genome alignment n MUMmer* (nucmer) l n Developed, supported and available at TIGR LAGAN*, AVID l VISTA identity plots w Multiple genome alignment n MGA, MLAGAN*, DIALIGN, MAVID w Multiple alignment n Muscle*, Clustal. W* w Local sequence alignment n BLAST*, FASTA, Vmatch *open source

MUMmer w Maximal Unique Matcher (MUM) n match l n maximal l n exact match of a minimum length cannot be extended in either direction without a mismatch unique l l l occurs only once in both sequences (MUM) occurs only once in a single sequence (MAM) occurs one or more times in either sequence (MEM)
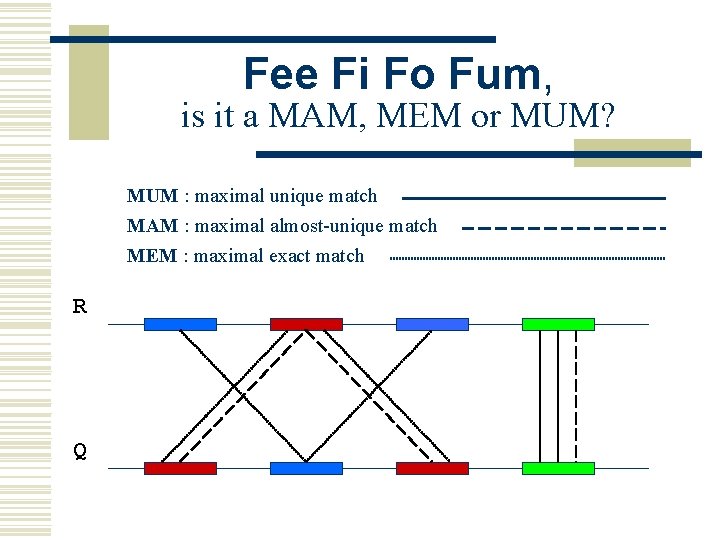
Fee Fi Fo Fum, is it a MAM, MEM or MUM? MUM : maximal unique match MAM : maximal almost-unique match MEM : maximal exact match R Q

Seed and extend w How can we make MUMs BIGGER? n Find MUMs using a suffix tree n Cluster MUMs using size, gap and distance parameters n Extend clusters using modified Smith-Waterman algorithm
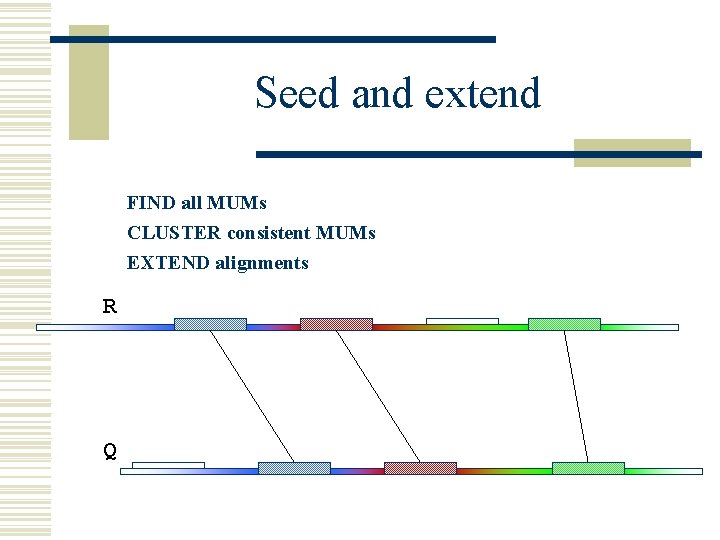
Seed and extend FIND all MUMs CLUSTER consistent MUMs EXTEND alignments R Q
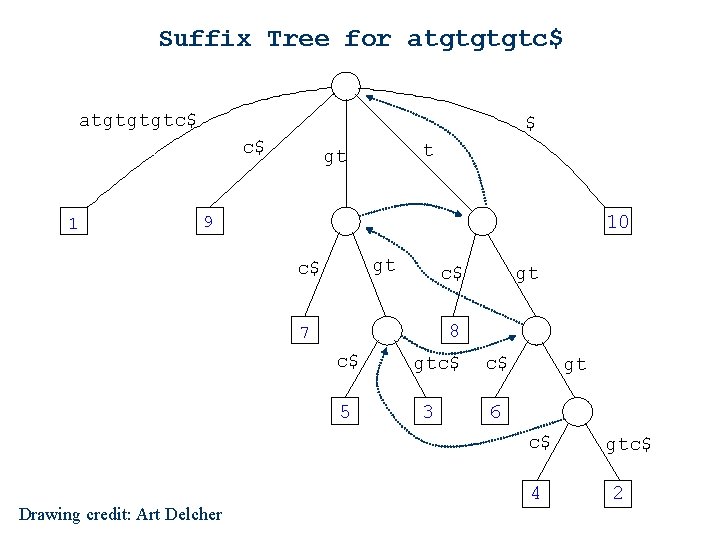
Suffix Tree for atgtgtgtc$ $ c$ 1 t gt 10 9 gt c$ c$ gt 8 7 c$ 5 gtc$ 3 c$ gt 6 c$ Drawing credit: Art Delcher 4 gtc$ 2
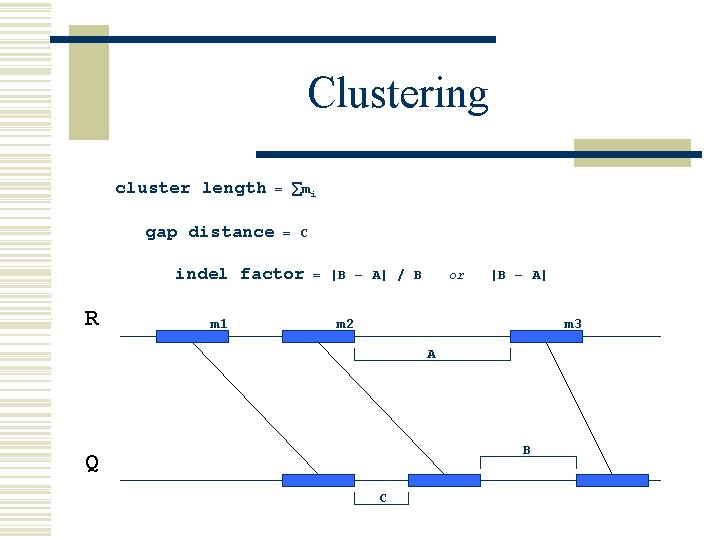
Clustering cluster length gap distance = ∑mi = C indel factor R m 1 = |B – A| / B or |B – A| m 2 m 3 A B Q C
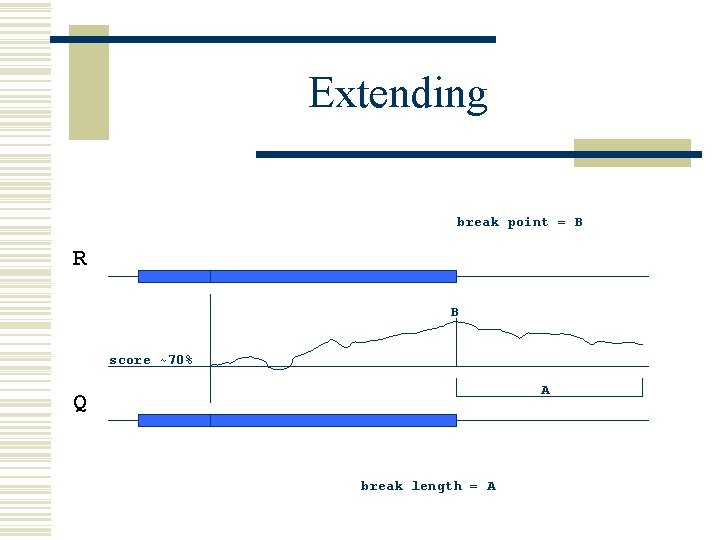
Extending break point = B R B score ~70% A Q break length = A
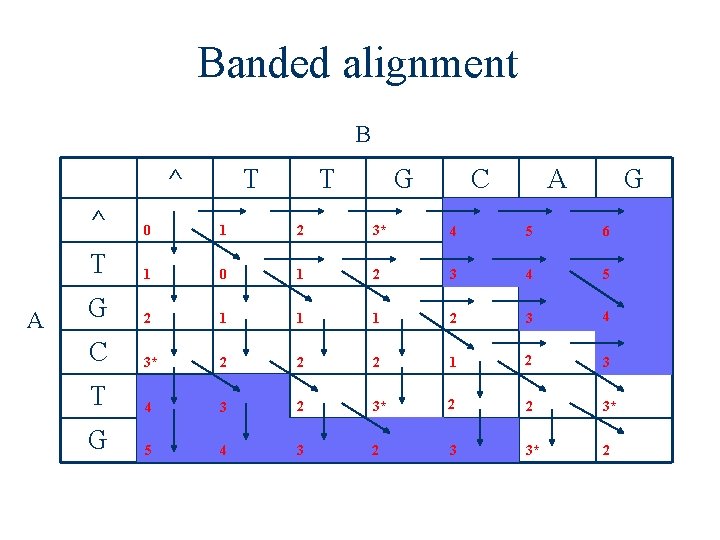
Banded alignment B ^ A T T G C A G ^ 0 1 2 3* 4 5 6 T 1 0 1 2 3 4 5 G 2 1 1 1 2 3 4 C 3* 2 2 2 1 2 3 T 4 3 2 3* 2 2 3* G 5 4 3 2 3 3* 2

MUMmer suite n mummer w exact matching n n nucmer alignment plotter § draft sequence mapping § w DNA multi-Fast. A input w whole genome alignment n w mummerplot § alignment visualization w show-coords § alignment summary w delta-filter § alignment filter w show-aligns § pairwise alignments promer w DNA multi-Fast. A input w whole genome alignment n n n run-mummer 1 w Fast. A input w global alignment run-mummer 3 w Fast. A input w/ draft w whole genome alignment exact-tandems w Fast. A input w exact tandem repeats NUCmer / PROmer utilities w mapview n System utilities w gnuplot w xfig

mummer w Primary uses l l exact matching (seeding) dot plotting w Pros l very efficient O(n) time and space w ~17 bytes per bp of reference sequence w E. coli K 12 vs. E. coli O 157: H 7 (~5 Mbp each) § 17 seconds using 77 MB RAM l multi-Fast. A input w Cons l exact matches only

nucmer w Primary uses l l whole genome alignment and analysis draft sequence alignment w Pros l l l multi-Fast. A inputs well suited for genome and contig mapping convenient helper utilities w show-coords, delta-filter, mummerplot w Cons l low sensitivity (w default parameters) with respect to BLAST

WGA example w Yersina pestis CO 92 vs. Yersina pestis KIM n High nucleotide similarity, 99. 86% l n Extensive genome shuffling l n Two strains of the same species Global alignment will not work Highly repetitive l Will confuse local alignment (e. g. BLAST)

COMMAND whole genome alignment nucmer –maxmatch CO 92. fasta KIM. fasta -maxmatch Find maximal exact matches (MEMs) delta-filter –m out. delta > out. filter. m -m -1 Many-to-many mapping One-to-one mapping show-coords -r out. delta. m > out. coords -r Sort alignments by reference position mummerplot --large --fat out. delta. m --large Large plot --fat Nice layout for multi-fasta files --x 11 Default, draw using x 11 (--postscript, --png) *requires gnuplot
![show-coords output n [S 1] start of the alignment region in the reference sequence show-coords output n [S 1] start of the alignment region in the reference sequence](http://slidetodoc.com/presentation_image_h/d1cdedb2ee1dcf818817a13d8d194fab/image-26.jpg)
show-coords output n [S 1] start of the alignment region in the reference sequence [E 1] end of the alignment region in the reference sequence [S 2] start of the alignment region in the query sequence [E 2] end of the alignment region in the query sequence [LEN 1] length of the alignment region in the reference sequence [LEN 2] length of the alignment region in the query sequence [% IDY] percent identity of the alignment [% SIM] percent similarity of the alignment [% STP] percent of stop codons in the alignment [LEN R] length of the reference sequence [LEN Q] length of the query sequence [COV R] percent alignment coverage in the reference sequence [COV Q] percent alignment coverage in the query sequence [FRM] reading frame for the reference and query sequence alignments respectively [TAGS] the reference and query Fast. A IDs respectively. n All output coordinates and lengths are relative to the forward strand n n n n

Comparative assembly w Assembly n Orient and place sequencing reads l Using overlaps and mate-pair information w Scaffolding n Order and orient draft contigs l Using mate-pair information and experimental validation w Comparative assembly and scaffolding n Orient and place reads and contigs l l Using a reference genome and alignment mapping e. g. AMOScmp (nucmer)
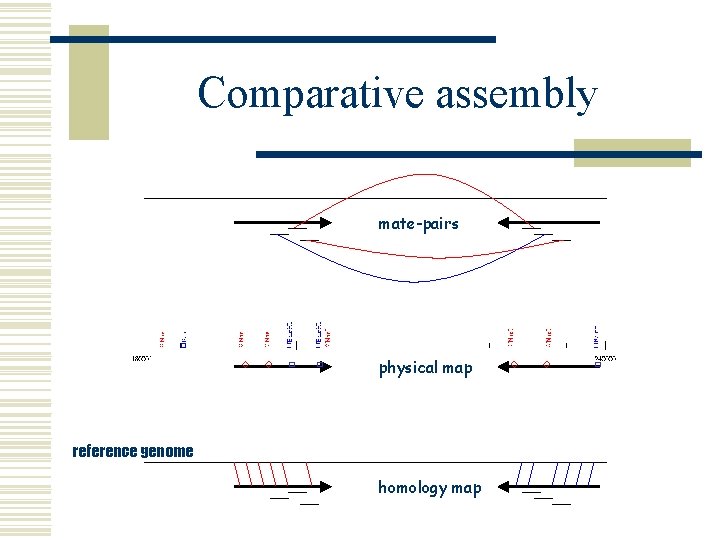
Comparative assembly mate-pairs physical map reference genome homology map
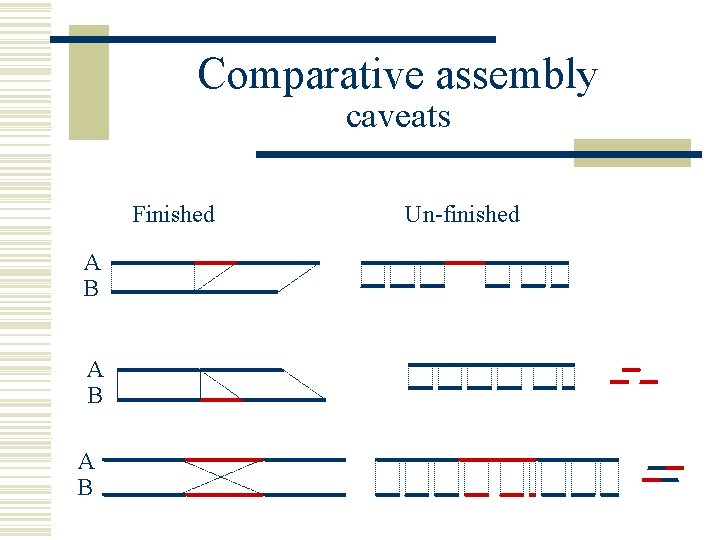
Comparative assembly caveats Finished A B A B Un-finished

COMMAND nucmer read/contig mapping nucmer –maxmatch REF. fasta QRY. fasta -maxmatch REF QRY Find maximal exact matches (MEMs) Reference sequence (genome) Query sequence to be mapped (reads, contigs) delta-filter –q out. delta > out. delta. q -q Best one-to-one mapping for each query show-coords –rcl out. delta. q > out. coords -r -c -l Sort alignments by reference Display alignment coverage percentage Display sequence length
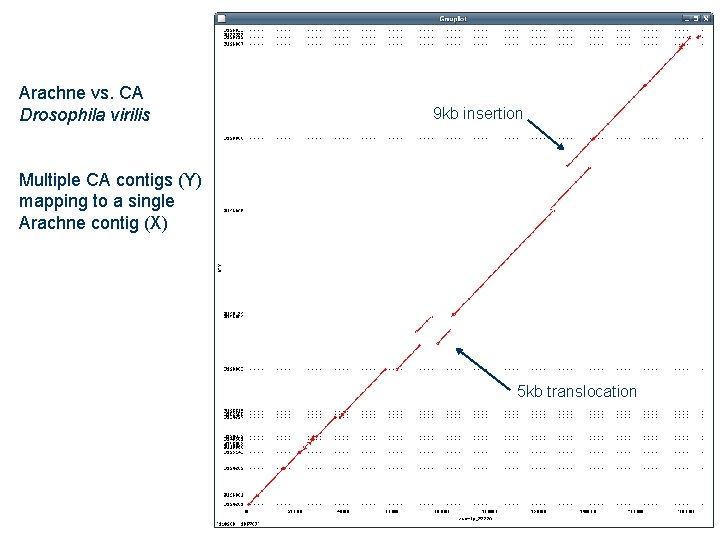
Arachne vs. CA Drosophila virilis 9 kb insertion Multiple CA contigs (Y) mapping to a single Arachne contig (X) 5 kb translocation

Gene reassembly* w nucmer -maxmatch genes. fasta reads. fasta w delta-filter -q out. delta. q w show-coords -THqcl out. delta. q > out. coords w Define matching criteria n %identity, %coverage, max gap size w Assemble matching reads n AMOS, minimus, hawkeye

References n Documentation l http: //mummer. sourceforge. net § l http: //mummer. sourceforge. net/manual § l thorough documentation http: //mummer. sourceforge. net/examples § n publication listing walkthroughs Email l l mummer-help (at) lists. sourceforge. net mummer-users (at) lists. sourceforge. net

Acknowledgements Art Delcher Steven Salzberg Stefan Kurtz Mike Schatz Mihai Pop
- Slides: 35