The Physiome Project Rob Mac Leod CVRTIBESCII Peter

The Physiome Project Rob Mac. Leod CVRTI/BE/SCII Peter Hunter Auckland University

What is the Physiome? “The Physiome Project is an integrated program whose mission is to archive and disseminate quantitative data and models of the functional behavior of biological molecules, cells, tissues, organs, and organisms. ” • Bassingthwaighte (1995): Advances in Experimental Medicine and Biology 1995; 382: 331 -9) – coined the term for the first time

Proposed Projects 1. Brain and CNS 2. Heart and cardiovascular system 3. Lungs and respiratory system 4. Kidney and urinary system 5. Musculo-skeletal system 6. Alimentary system 7. Reproductive system 8. Endocrine system 9. Haemolymphoid system 10. Integumental system

Physiome Bioinformatics Modeling Hierarchies • Genes • Proteins • Biophysical models • Constitutive laws • Organ model • Whole body model Databases • Genome • Protein • Physiology • Structural Bioengineering • Bioeng. Materials Clinical medicine • Clinical Molecular Biology Physiology

Mathematical Models Level 1 models: Molecular models Level 2 models: Subcellular Markov models Level 3 models: Subcellular ODE models Level 4 models: Tissue and whole organ continuum models Level 5 models: Whole body continuum models Level 6 models: Whole body system models

Visualization Tools • Interrogation of model parameters • Animated visualization of computational output • From molecular level through to the whole body • Web based • Coupled to the computational models in a user-friendly fashion.

Instrumentation • Structural measurements – geometry and tissue microstructure of organs – present methods too slow and tedious • Material property measurements – – mechanical, electrical, thermal, etc variety of species pathological conditions nonlinear, coupled parameters

Physiome Groups • Bio. No. ME (UCSD) – Biology Network of Modeling Efforts; limited activity but good pedigree – funded by Procter and Gamble for 3 years • Cardiome Project (Auckland) – the model and most active group • Microcirculatory Physiome Project (Johns Hopkins) – seems well supported and active • Endotheliome Project • Pulmonary Physiome

Physiome Links • www. ornl. gov/Tech. Resources/Human_Genome/faqs 1. html – the mother of all bioscience megaprojects • bionome. sdsc. edu/ – source of some data and info • www. physiome. org/ – the home site; poorly maintained • www. esc. auckland. ac. nz/People/Staff/Hunter/physiome. html – the best place to start the search • http: //www. esc. auckland. ac. nz/sites/physiome/index. html – home of the *ML’s • www. bme. jhu. edu/news/microphys/ – microcirculatory physiome project • http: //www. physiome. com/ – the company • www. IBB. gatech. edu/Hilton. Head/hiltonheadworkshop. html – the next relevant meeting on the physiome

Bio. No. ME Project (UCSD) • Biology Network of Modeling Efforts • Funded by Procter and Gamble for 3 years • Contains interface to models that run on remote supercomputer (at UCSD) • Only biophysical models are Beeler-Reuter and Luo-Rudy I • Anatomical models of the heart and microcirculation • Limited activity but good pedigree

Microcirculation Physiome “Thus, we propose to develop a database of the microcirculation that encompasses anatomical and functional data with mathematical and computational models, computational engines, and tools for integration. ” • Many particpants (based at JHU): – U Western Ontario, Auckland, U Tennessee, U Virginia, JHU, U Arizona • Projects – anatomy, e. g. , using GIS – functional descriptions of microcirculation, transport, etc.
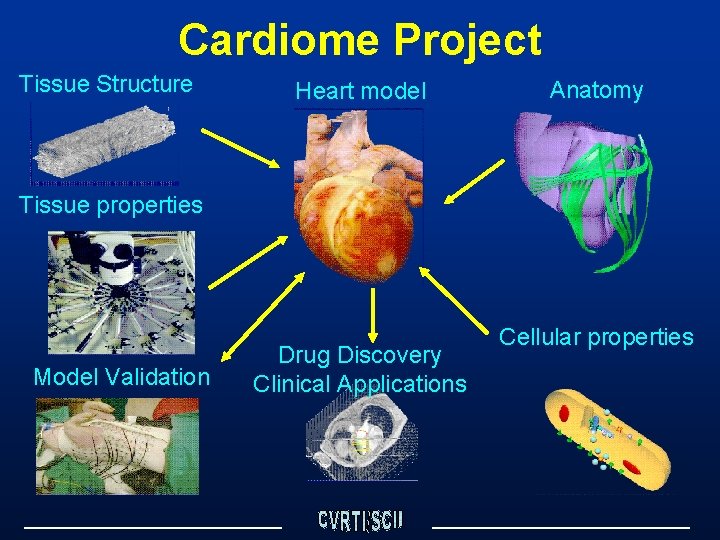
Cardiome Project Tissue Structure Heart model Anatomy Tissue properties Model Validation Drug Discovery Clinical Applications Cellular properties
Cardiome Groups AU UCSD Physiome Sciences Duke David Bullivant Peter Hunter Ian Le. Grice Denis Loiselle Poul Nielsen Andrew Pullan Bruce Smaill Alistair Young Jim Covell Andrew Mc. Culloch Jeff Omens Tom Colatsky Gang Chen Adam Muzikant Craig Henriquez et al JHU Bill Hunter Eduardo Marban Sasha Popel Rai Winslow UCSF Julius Guccione Mark Ratcliffe Tulane Natalia Trayanova et al. MIT Oxford Colin Brennan Forbes Dewey Ian Hunter Chris Bradley Martyn Nash Peter Kohl Denis Noble Nic Smith Lausanne UW (Seattle) Columbia Maastricht Theo Arts Frits Prinzen Rob Reneman Utah Chris Johnson Rob Mac. Leod Bruno Taccardi Jim Bassingthwaighte Eric Feigl Zheng Li Technion Israel Sam Sideman et al Nathalia Virag et al CWRU Yoram Rudy et al Kevin Costa Jeff Holmes

Anatomy • Completed or underway: – – – Vent. geom. & fibre-sheet structure for dog (AU) Vent. geom. & fibre-sheets for rabbit (UCSD, JHU) Coronary anatomy for pig (UCSD) Atrial geometry & structure for pig (UCSD, AU, . . . ) Cardiac valve structure (AU) Automated measurement rig (AU & MIT) • Needed soon: – Geom. & fibre-sheet structure for pig, human – Geom. & fibre-sheet structure for hypertrophy etc

Mechanics • Completed or underway: – Material properties - • biaxial tests on dog myocardium (AU) • shear testing of pig myocardium (AU) • torsion testing of rabbit pap. muscle (JHU) – – – ECM structure (UCSD, Columbia, AU, JHU) Functional studies on gene targetted mice (UCSD) Infarct modelling (UCSD, Columbia, AU) Ventricular aneurysm (UCSF) Acute ischaemia (UCSD, UWash) • Needed soon: – Microstructure & mechanical properties of cytoskeleton & ECM
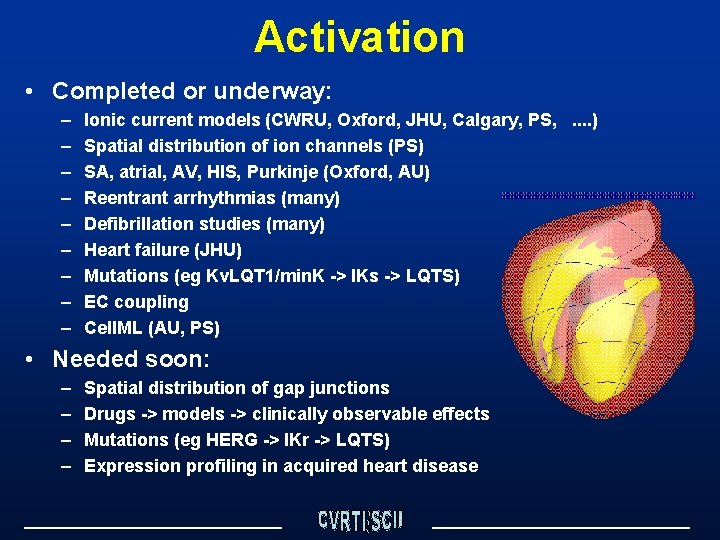
Activation • Completed or underway: – – – – – Ionic current models (CWRU, Oxford, JHU, Calgary, PS, . . ) Spatial distribution of ion channels (PS) SA, atrial, AV, HIS, Purkinje (Oxford, AU) Reentrant arrhythmias (many) Defibrillation studies (many) Heart failure (JHU) Mutations (eg Kv. LQT 1/min. K -> IKs -> LQTS) EC coupling Cell. ML (AU, PS) • Needed soon: – – Spatial distribution of gap junctions Drugs -> models -> clinically observable effects Mutations (eg HERG -> IKr -> LQTS) Expression profiling in acquired heart disease

Energy Supply & Metabolism • Completed or underway: – – – Coronary flow (AU) Coronary flow regulation (UW) Metabolism & energetics (UW, Oxford, AU) Ischaemia (Oxford, AU) Flux balance & kinetic models (UCSD, CWRU) • Needed soon: – Integration of different parts of metabolic pathway models – with energy supply & demand – Coupling to electrophysiology & generation of reentrant arrhythmias

Databases Cell • • • Tissue Organ(ism) Structure and spatial parameters Material properties Dynamic behavior Documentation Communications and interactions
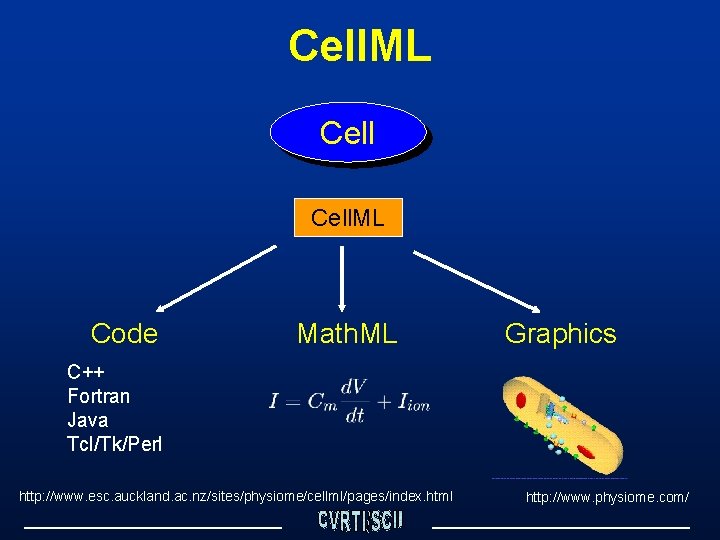
Cell. ML Code Math. ML Graphics C++ Fortran Java Tcl/Tk/Perl http: //www. esc. auckland. ac. nz/sites/physiome/cellml/pages/index. html http: //www. physiome. com/

XML www. w 3. org/XML/ • XML is a method for putting structured data in a text file • XML looks a bit like HTML but isn't HTML • XML is text, but isn't meant to be read • XML is a family of technologies • XML is verbose, but that is not a problem • XML is new, but not that new • XML is license-free, platform-independent and well-supported • (Note: supports Math. ML!)

Cell. ML Basics • Experiment (simulation) – model description – preparation (species, boundary conditions, other conditions) – variables (to overwrite those extracted from and existing database) – independent variables (time, space) – experimental results (from simulation) • Model description most advanced component
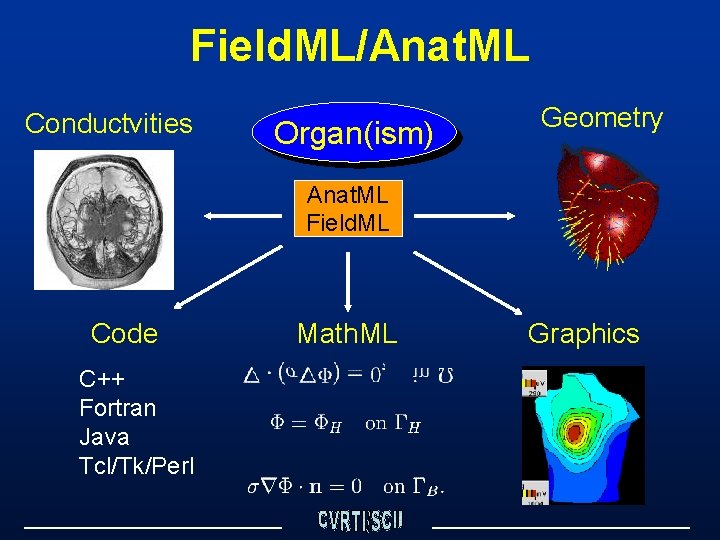
Field. ML/Anat. ML Conductvities Organ(ism) Geometry Anat. ML Field. ML Code C++ Fortran Java Tcl/Tk/Perl Math. ML Graphics

Anat. ML • <body part> – description of geometry – uses CMISS data structures • <body group> – links to other body parts • <placement> – location of the body part in space

Mesh. ML/Field. ML/Region. ML • Mesh. ML – elements of the geometry with connectivities • Field. ML – basis functions – field parameters • Region. ML – container for meshes and fields

The Future of the Physiome • • • Support from other labs Cooperation between labs Funding Technical challenges Our role? ? ?
- Slides: 25