An open platform for physiome and systems biology

An open platform for physiome and systems biology: insilico. ML and IDE as a Cell. ML compatible model sharing environment T Nomura 1, 2, Y. Asai 2, K Hagihara 2, 3 and Y Kurachi 2, 4 1. Division of Bioengineering, Graduate School of Engineering Science, Osaka University 2. The Center for Advanced Medical Engineering and Informatics, Osaka University 3. Department of Computer Science, Graduate School of Information Science and Technology 4. Department of Pathophysiology and Therapeutics, Graduate School of Medicine, Osaka University SIAM Model Sharing Panel, 4 th Aug, 2008, Montreal
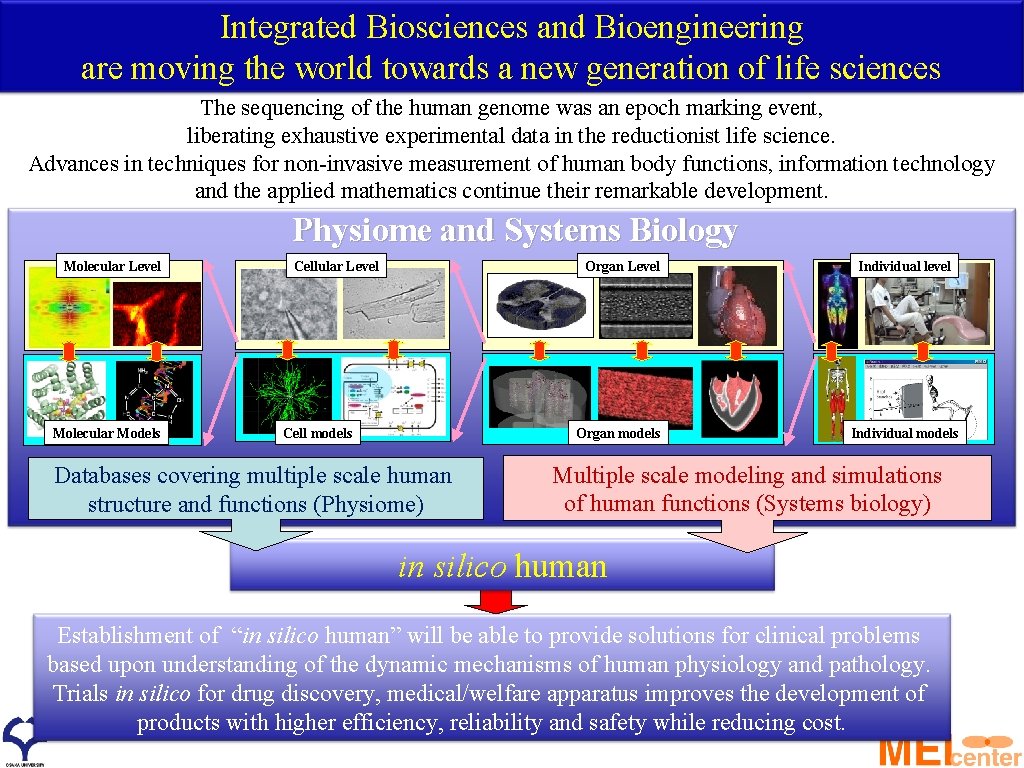
Integrated Biosciences and Bioengineering are moving the world towards a new generation of life sciences The sequencing of the human genome was an epoch marking event, liberating exhaustive experimental data in the reductionist life science. Advances in techniques for non-invasive measurement of human body functions, information technology and the applied mathematics continue their remarkable development. Physiome and Systems Biology Molecular Level Molecular Models Cellular Level Organ Level Cell models Organ models Databases covering multiple scale human structure and functions (Physiome) Individual level Individual models Multiple scale modeling and simulations of human functions (Systems biology) in silico human Establishment of “in silico human” will be able to provide solutions for clinical problems based upon understanding of the dynamic mechanisms of human physiology and pathology. Trials in silico for drug discovery, medical/welfare apparatus improves the development of products with higher efficiency, reliability and safety while reducing cost.

Physiome. jp as a Worldwide Open Platform for Physiome and Systems Biology
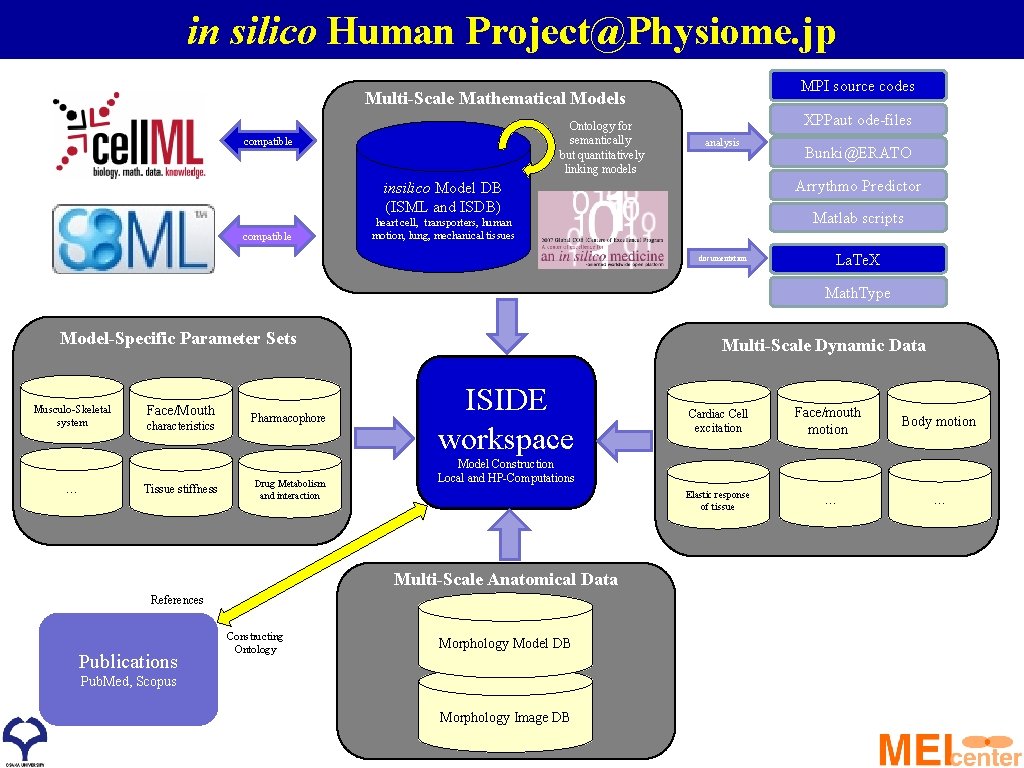
in silico Human Project@Physiome. jp MPI source codes Multi-Scale Mathematical Models Ontology for semantically but quantitatively linking models compatible XPPaut ode-files analysis Arrythmo Predictor insilico Model DB (ISML and ISDB) compatible Bunki@ERATO Matlab scripts heart cell, transporters, human motion, lung, mechanical tissues documentation La. Te. X Math. Type Model-Specific Parameter Sets Musculo-Skeletal system … Face/Mouth characteristics Tissue stiffness Pharmacophore Drug Metabolism and interaction Multi-Scale Dynamic Data ISIDE workspace References Publications Face/mouth motion Body motion Elastic response of tissue … … Model Construction Local and HP-Computations Multi-Scale Anatomical Data Constructing Ontology Cardiac Cell excitation Morphology Model DB Pub. Med, Scopus Morphology Image DB

Physiome. jp as a Model Sharing Platform - Enhancing Knowledge Integration -
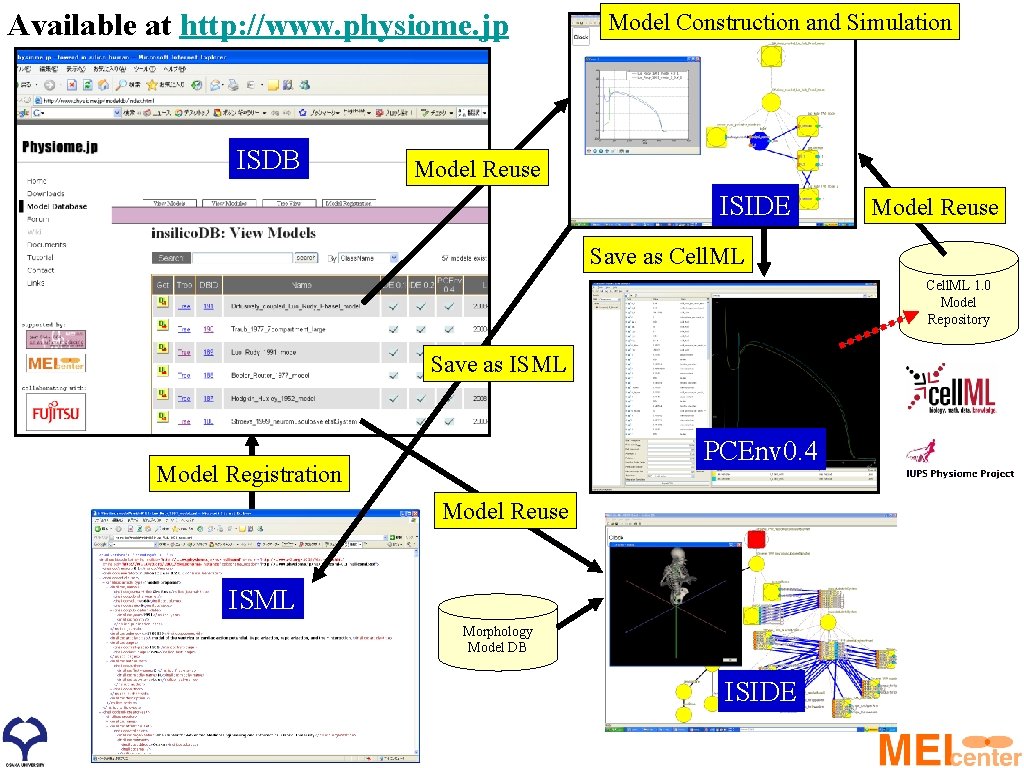
Available at http: //www. physiome. jp ISDB Model Construction and Simulation Model Reuse ISIDE Model Reuse Save as Cell. ML 1. 0 Model Repository Save as ISML PCEnv 0. 4 Model Registration Model Reuse ISML Morphology Model DB ISIDE
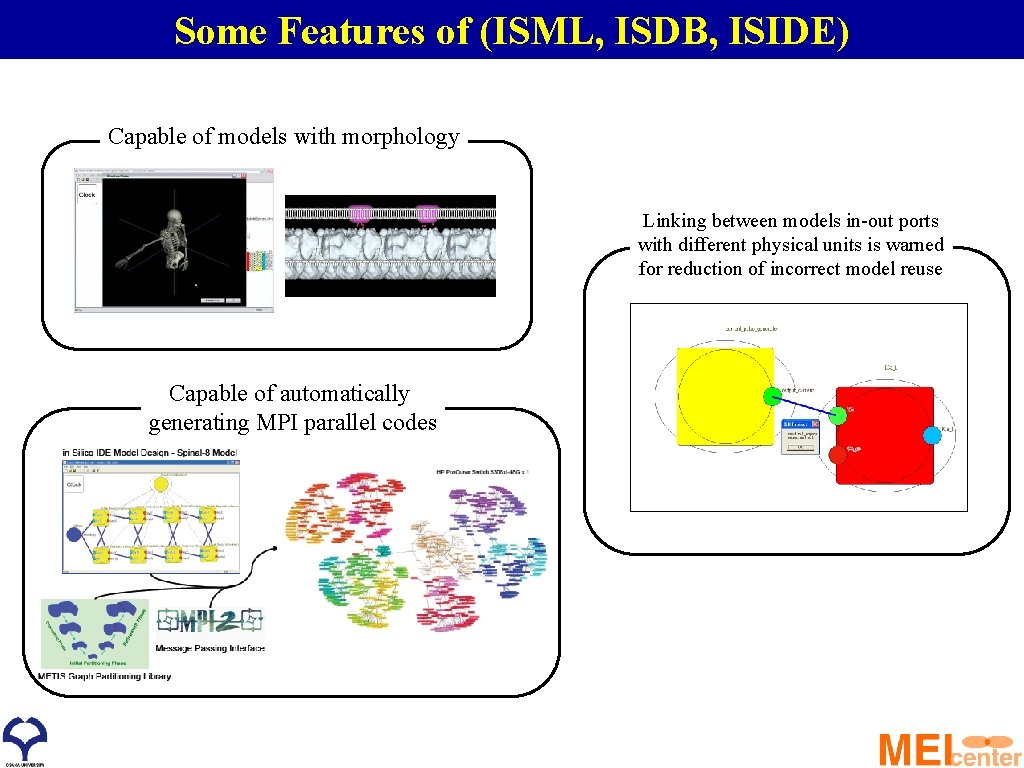
Some Features of (ISML, ISDB, ISIDE) Capable of models with morphology Linking between models in-out ports with different physical units is warned for reduction of incorrect model reuse Capable of automatically generating MPI parallel codes

A Questionnaire Survey on the Model Sharing in the Model-Related Life Science Fields in Japan Physiome and Systems Biology Initiative Japan F. Kajiya (Kawasaki) H. Kitano (Systems Biology Institute) Y. Kurachi (Osaka) M. Suematsu (Keio) T. Nomura (Osaka) R. Himeno (RIKEN) Y. Matsumoto (Tokyo)
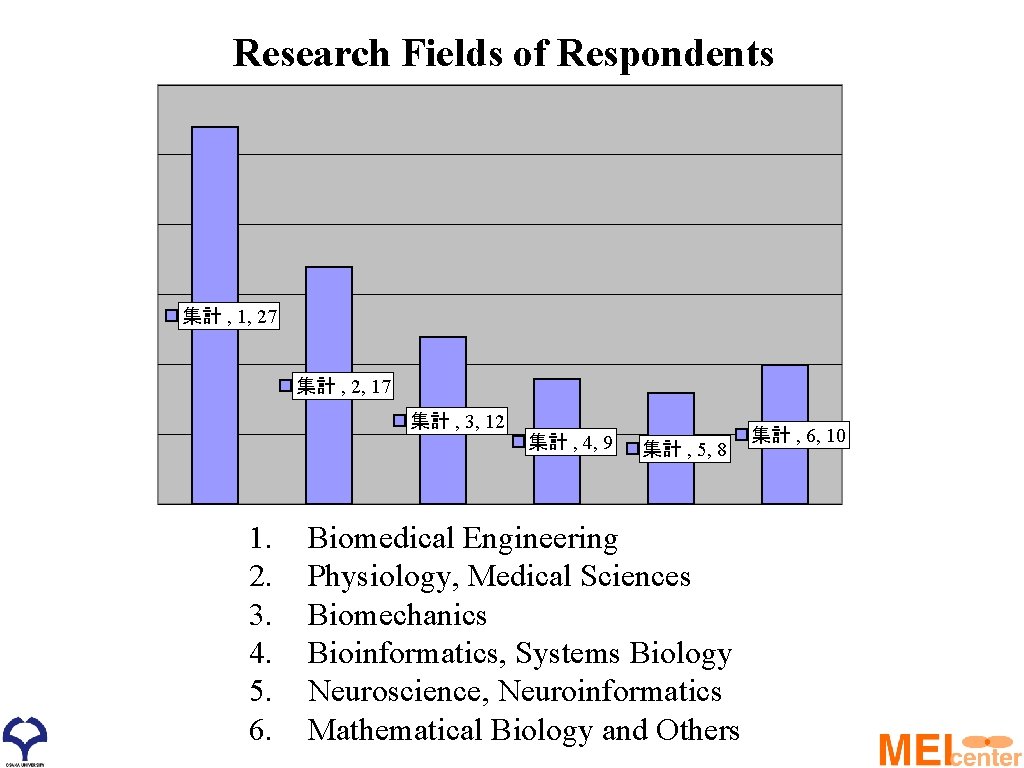
Research Fields of Respondents 集計 , 1, 27 集計 , 2, 17 集計 , 3, 12 1. 2. 3. 4. 5. 6. 集計 , 4, 9 集計 , 5, 8 Biomedical Engineering Physiology, Medical Sciences Biomechanics Bioinformatics, Systems Biology Neuroscience, Neuroinformatics Mathematical Biology and Others 集計 , 6, 10
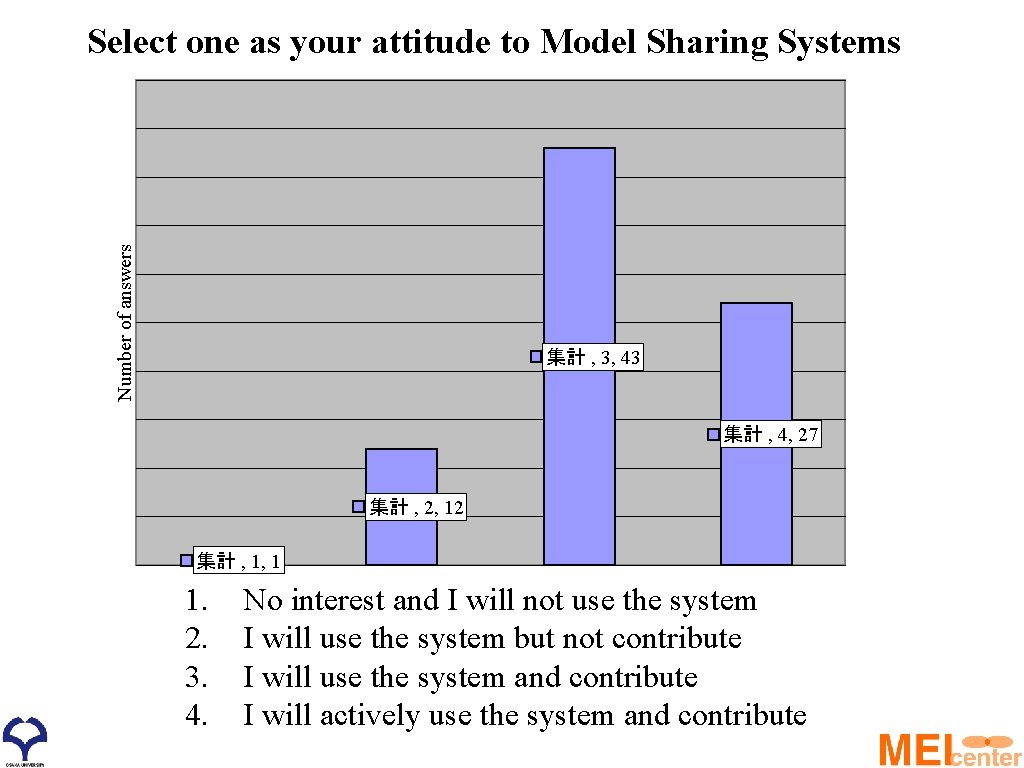
Number of answers Select one as your attitude to Model Sharing Systems 集計 , 3, 43 集計 , 4, 27 集計 , 2, 12 集計 , 1, 1 1. 2. 3. 4. No interest and I will not use the system I will use the system but not contribute I will use the system and contribute I will actively use the system and contribute
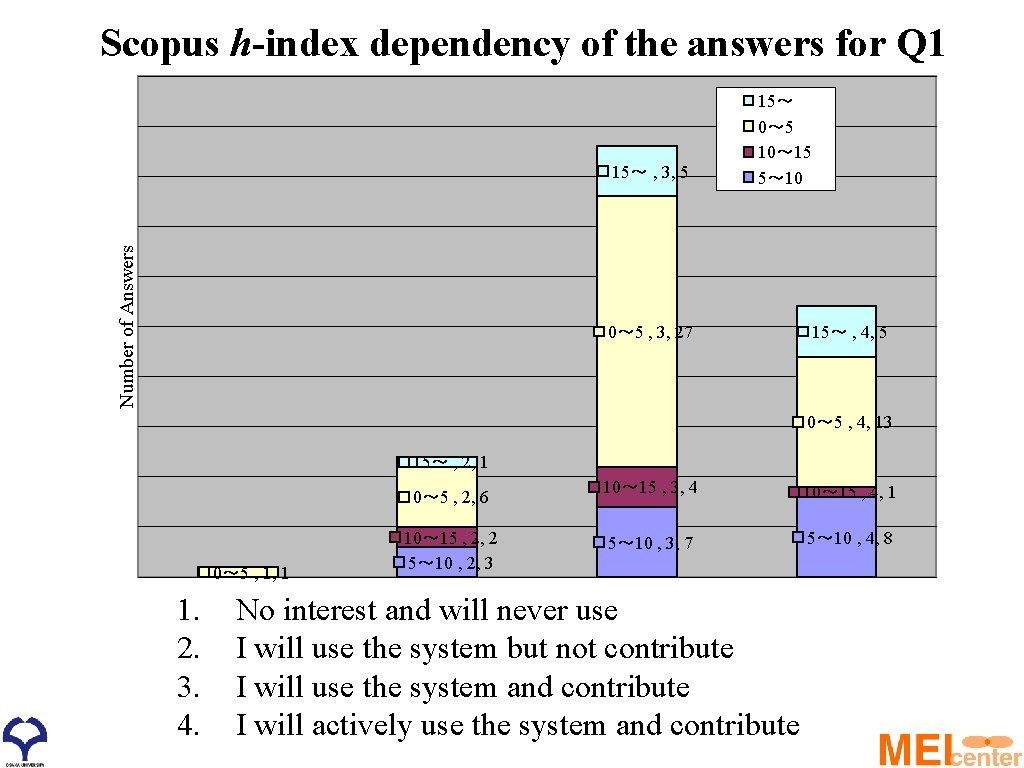
Scopus h-index dependency of the answers for Q 1 Number of Answers 15~ , 3, 5 15~ 0~ 5 10~ 15 5~ 10 0~ 5 , 3, 27 15~ , 4, 5 0~ 5 , 4, 13 15~ , 2, 1 0~ 5 , 2, 6 0~ 5 , 1, 1 1. 2. 3. 4. 10~ 15 , 2, 2 5~ 10 , 2, 3 10~ 15 , 3, 4 10~ 15 , 4, 1 5~ 10 , 3, 7 5~ 10 , 4, 8 No interest and will never use I will use the system but not contribute I will use the system and contribute I will actively use the system and contribute
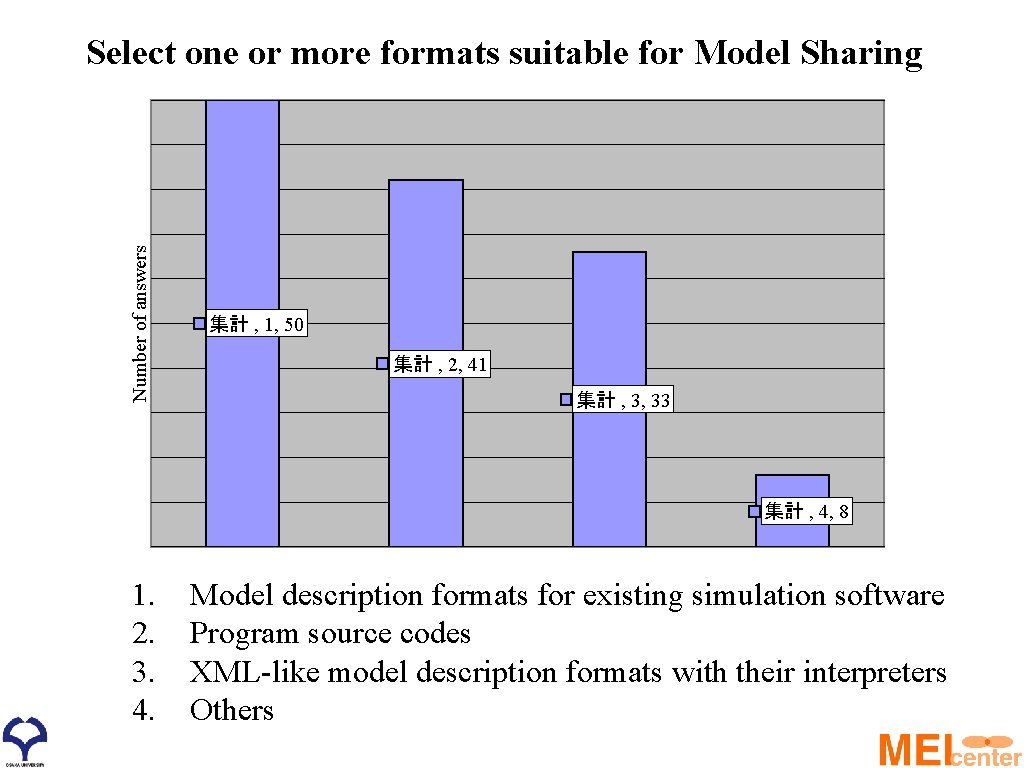
Number of answers Select one or more formats suitable for Model Sharing 集計 , 1, 50 集計 , 2, 41 集計 , 3, 33 集計 , 4, 8 1. 2. 3. 4. Model description formats for existing simulation software Program source codes XML-like model description formats with their interpreters Others
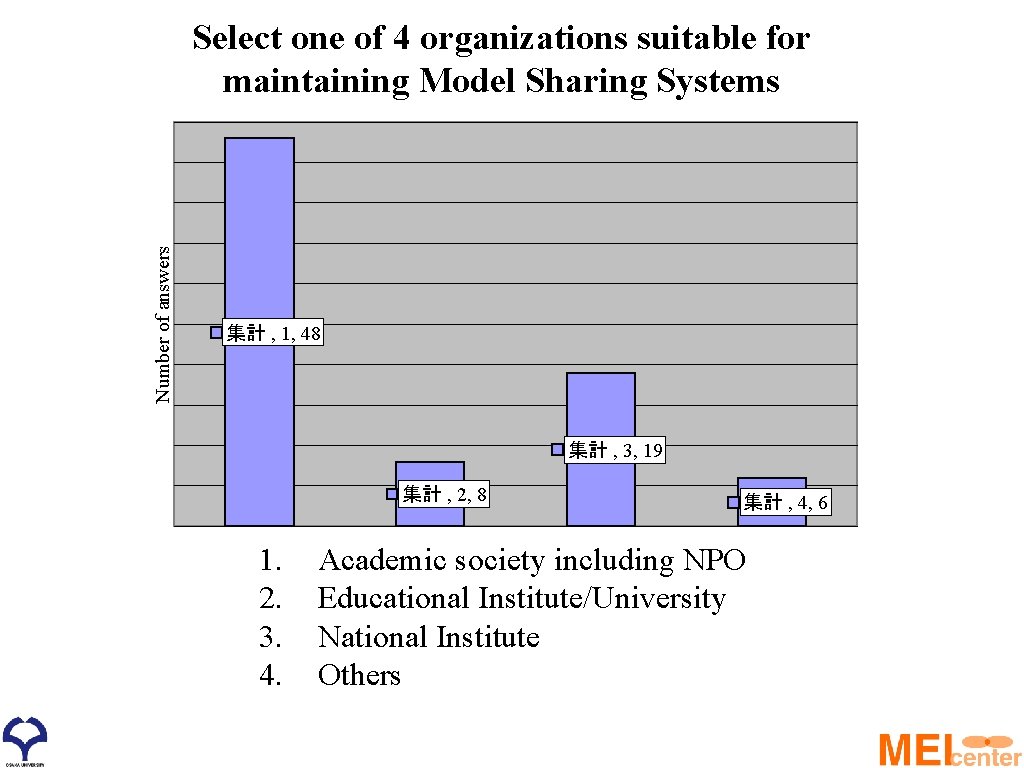
Number of answers Select one of 4 organizations suitable for maintaining Model Sharing Systems 集計 , 1, 48 集計 , 3, 19 集計 , 2, 8 1. 2. 3. 4. 集計 , 4, 6 Academic society including NPO Educational Institute/University National Institute Others
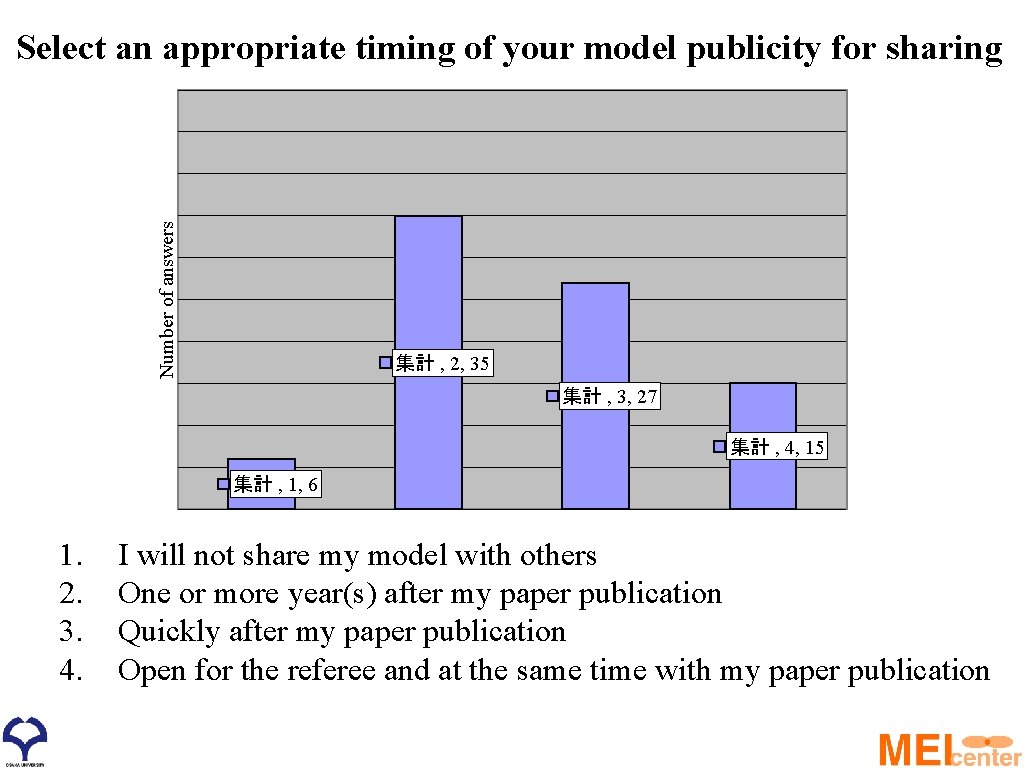
Number of answers Select an appropriate timing of your model publicity for sharing 集計 , 2, 35 集計 , 3, 27 集計 , 4, 15 集計 , 1, 6 1. 2. 3. 4. I will not share my model with others One or more year(s) after my paper publication Quickly after my paper publication Open for the referee and at the same time with my paper publication
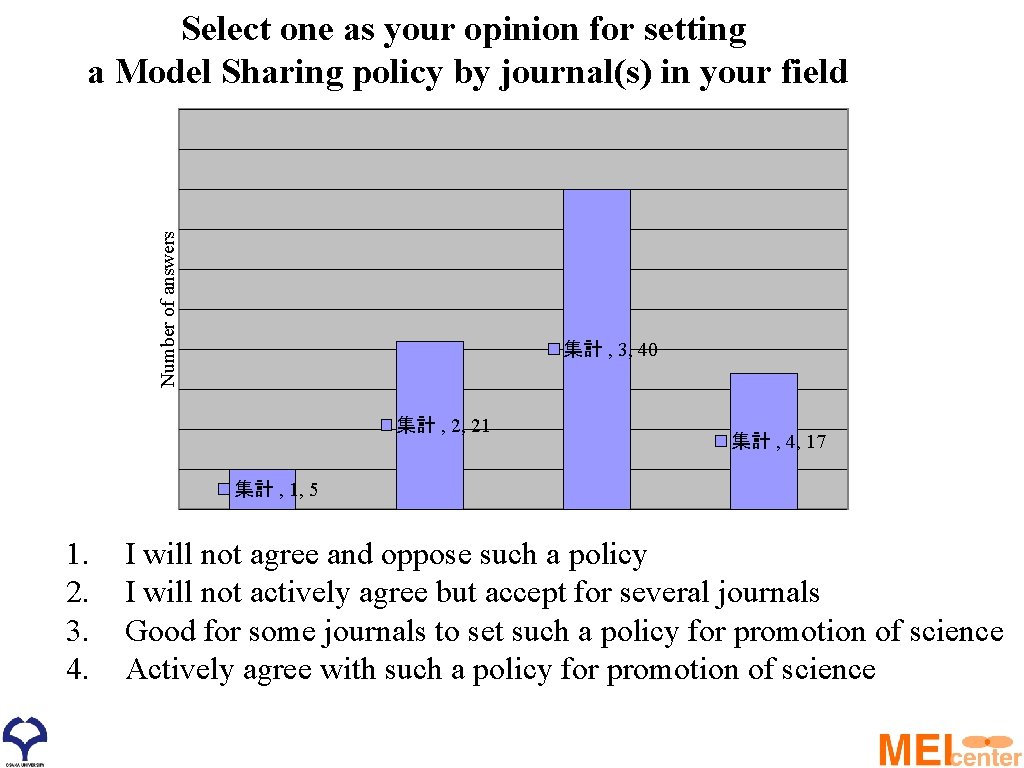
Number of answers Select one as your opinion for setting a Model Sharing policy by journal(s) in your field 集計 , 3, 40 集計 , 2, 21 集計 , 4, 17 集計 , 1, 5 1. 2. 3. 4. I will not agree and oppose such a policy I will not actively agree but accept for several journals Good for some journals to set such a policy for promotion of science Actively agree with such a policy for promotion of science
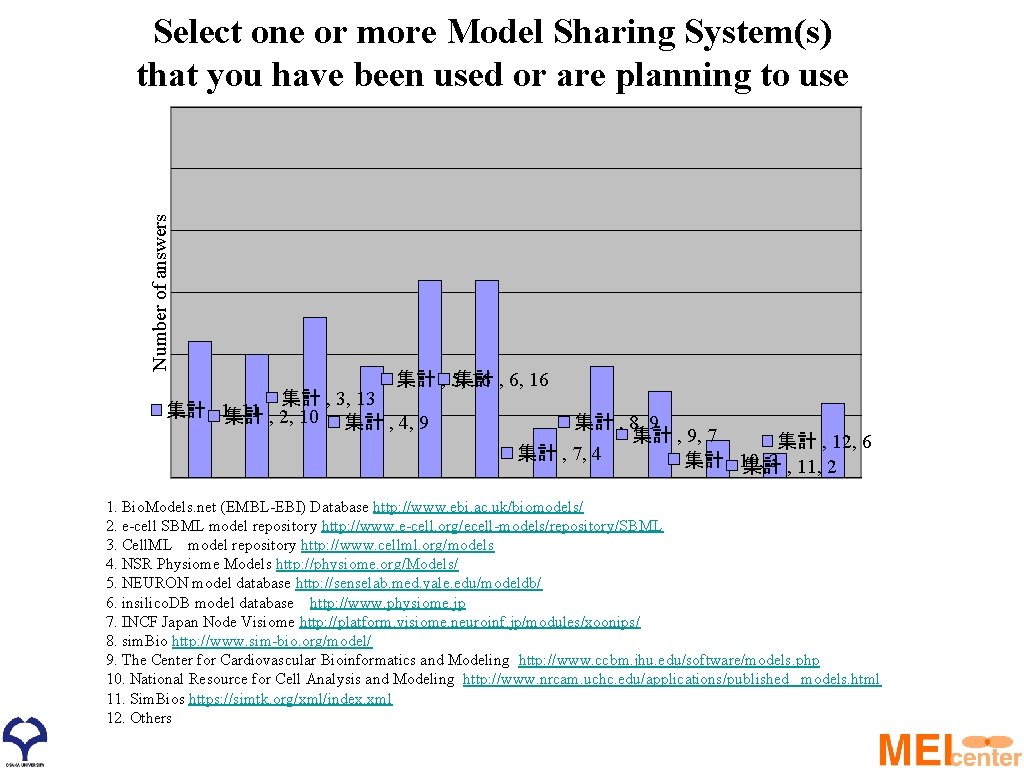
Number of answers Select one or more Model Sharing System(s) that you have been used or are planning to use 集計 , 5, 集計 16 , 6, 16 集計 , 3, 13 集計 , 1, 11 集計 , 2, 10 集計 , 4, 9 集計 , 8, 9 集計 , 9, 7 集計 , 12, 6 集計 , 7, 4 集計 , 10, 3 , 11, 2 集計 1. Bio. Models. net (EMBL-EBI) Database http: //www. ebi. ac. uk/biomodels/ 2. e-cell SBML model repository http: //www. e-cell. org/ecell-models/repository/SBML 3. Cell. ML model repository http: //www. cellml. org/models 4. NSR Physiome Models http: //physiome. org/Models/ 5. NEURON model database http: //senselab. med. yale. edu/modeldb/ 6. insilico. DB model database http: //www. physiome. jp 7. INCF Japan Node Visiome http: //platform. visiome. neuroinf. jp/modules/xoonips/ 8. sim. Bio http: //www. sim-bio. org/model/ 9. The Center for Cardiovascular Bioinformatics and Modeling http: //www. ccbm. jhu. edu/software/models. php 10. National Resource for Cell Analysis and Modeling http: //www. nrcam. uchc. edu/applications/published _models. html 11. Sim. Bios https: //simtk. org/xml/index. xml 12. Others

Comments Made by the Respondents Taking every detail of a published model non-public is like publishing a scientific paper in molecular biology without taking gene sequences public. Thus the model sharing by means of numerically verifiable manner is not surprising, and it will lead to a new spiral of intellectual scientific activity such as impacted by Science 2. 0. If the model sharing is going to be made by means of Open-Source like policy, it is preferable that the systems are run not by policies of scientific journals and societies but by certain organizations such as OBF (Open Bio Foundation) based on voluntary movement of the researchers. Otherwise, without sufficient incentive mechanisms, this field will loose top-class personnel for developing software. Necessity of: - international agreements to establish the system-managing organization - long range sufficient financial and human-resource supports to run the system - (automatic) model-curation mechanisms development of quantitative ontology providing for explosion of the number of models peer review for a submitted model in addition to journal review - education system compensating easy-use of shared models linking to the original paper, references, and related educational models to simulate - incentive mechanisms for contribution to the model sharing - incentive mechanisms for maintaining the systems

Collaborators Yoshiyuki Asai, Ph. D ISML-DB-IDEChief Developer Masakazu Yagi, Ph. D Hideki Oka, Ph. D (FEM and HPC) Face and Mouth Motion Takahito Urai (Programming) Kenji Yamada, Ph. D Tatsuhide Okamoto (Model Creator) Tissue Stiffness Eric Hien (HPC, Grid, Math-Editor) Rachid Ait-Haddou, Ph. D Morphological Modeling Ca 2+ dynamics Yoshiyuki Kido, Ph. D Masao Nakanishi (Large Scale Modeling) Yasuyuki Suzuki (Musculo-Skeletal System) Database and Ontology Yosuke Yumikura (Ca 2+ Dyanamics) Yoshiyuki Kagiyama, Ph. D Toshihiro Kawazu (Programming, Ca 2+ Dyanamics) Image Processing Daisuke Yamasaki, Ph. D Pharmacophore Kunichika Tumoto, Ph. D Cardiac Cell Dyanmics Masanori Ato (Pharmaco-Dynamics) Kazuyoshi Kawa (Constraint Dynamics)

Multi-Level Appendix
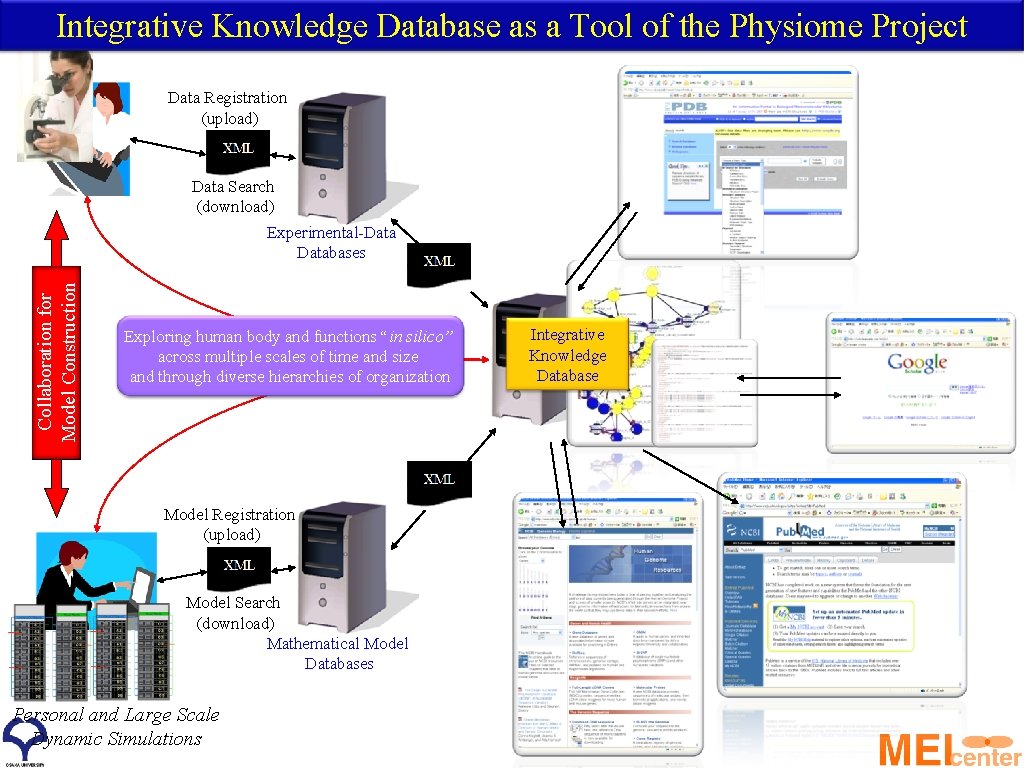
Integrative Knowledge Database as a Tool of the Physiome Project Data Registration (upload) Data Search (download) Collaboration for Model Construction Experimental-Databases Exploring human body and functions “in silico” across multiple scales of time and size and through diverse hierarchies of organization Model Registration (upload) Model Search (download) Mathematical Model Databases Personal and Large Scale Dynamic Simulations Integrative Knowledge Database
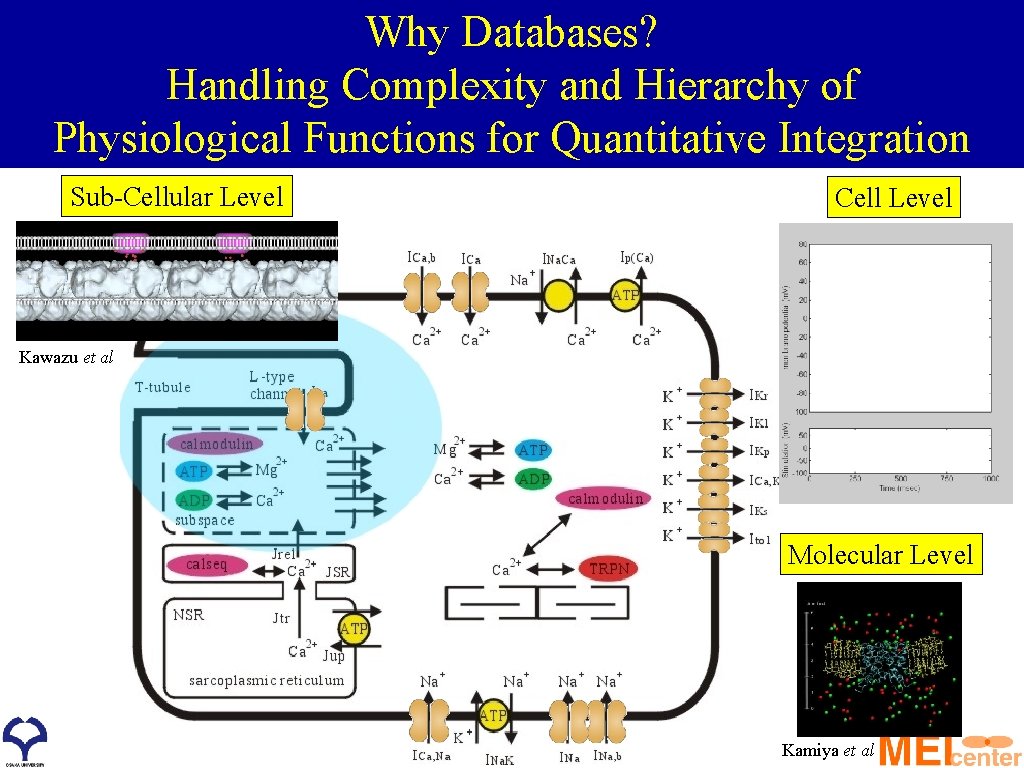
Why Databases? Handling Complexity and Hierarchy of Physiological Functions for Quantitative Integration Sub-Cellular Level Cell Level Kawazu et al Molecular Level Kamiya et al
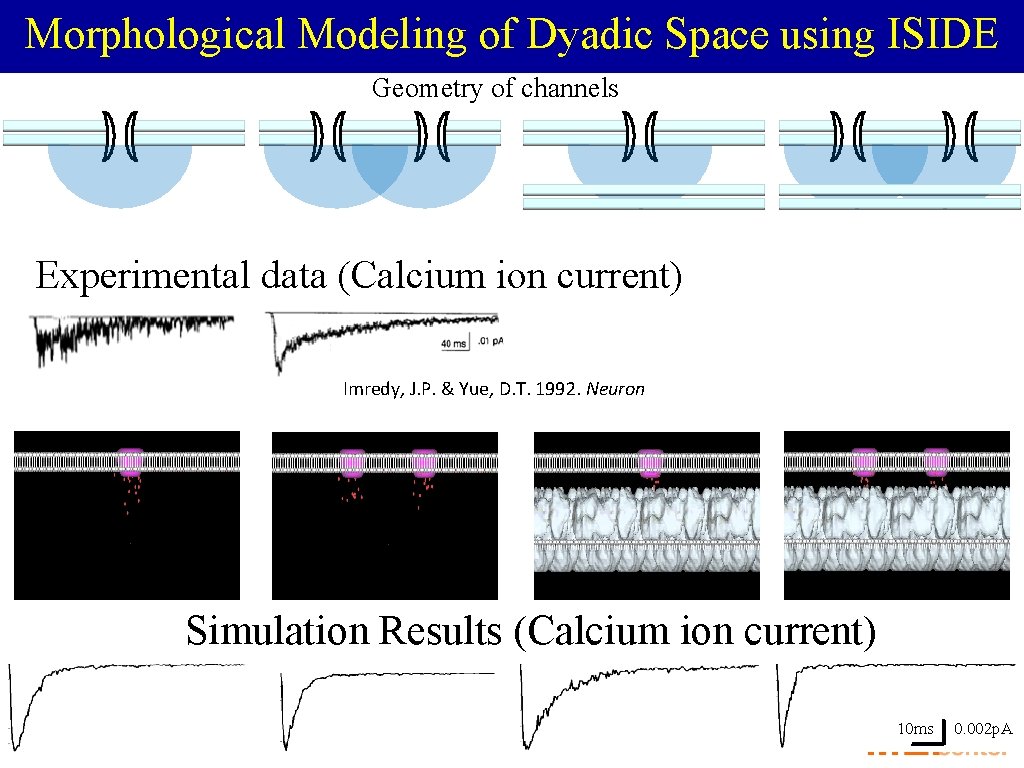
Morphological Modeling of Dyadic Space using ISIDE Geometry of channels Experimental data (Calcium ion current) Imredy, J. P. & Yue, D. T. 1992. Neuron Simulation Results (Calcium ion current) 10 ms 0. 002 p. A
- Slides: 22