SISTEM ENDOMEMBRAN KOMPARTEMEN INTRASELULER REGULASI PROTEIN PASCATRANSLASI Concept

SISTEM ENDOMEMBRAN KOMPARTEMEN INTRASELULER REGULASI PROTEIN PASCATRANSLASI

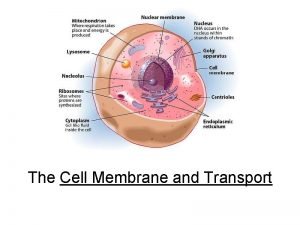
Concept 6. 4: The endomembrane system regulates protein traffic and performs metabolic functions in the cell • Components of the endomembrane system: – Nuclear envelope – Endoplasmic reticulum – Golgi apparatus – Lysosomes – Vacuoles – Plasma membrane • These components are either continuous or connected via transfer by vesicles Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings
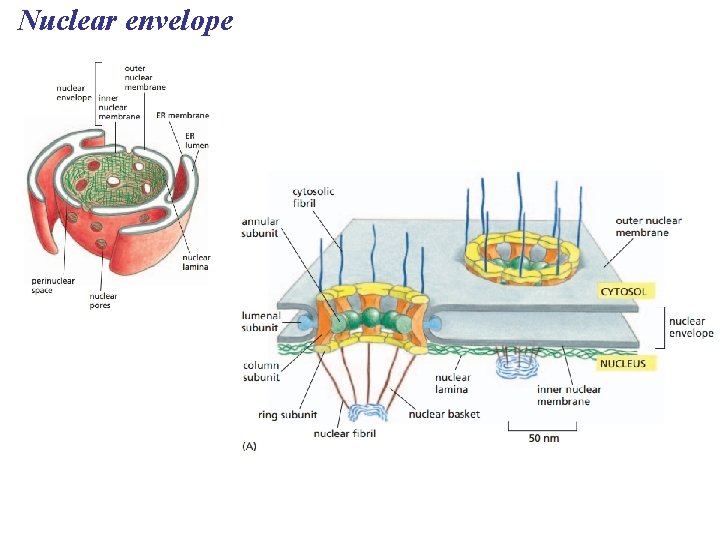
Nuclear envelope

The Endoplasmic Reticulum: Biosynthetic Factory • The endoplasmic reticulum (ER) accounts for more than half of the total membrane in many eukaryotic cells • The ER membrane is continuous with the nuclear envelope • There are two distinct regions of ER: – Smooth ER, which lacks ribosomes – Rough ER, with ribosomes studding its surface Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings
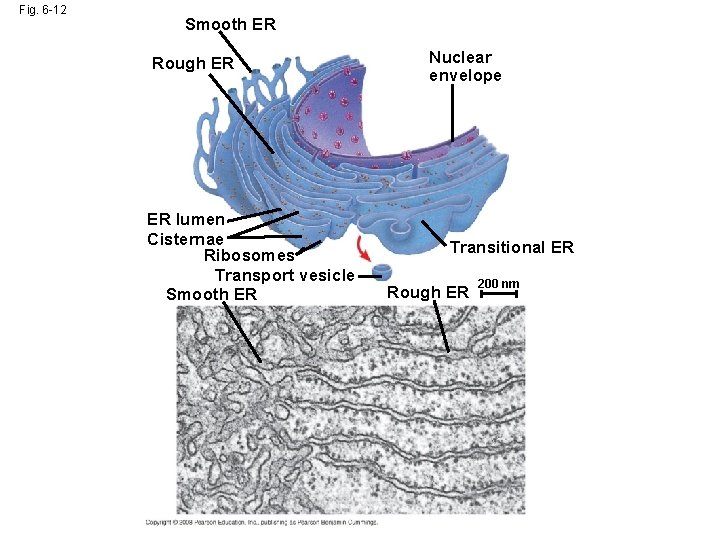
Fig. 6 -12 Smooth ER Rough ER ER lumen Cisternae Ribosomes Transport vesicle Smooth ER Nuclear envelope Transitional ER Rough ER 200 nm

Functions of Smooth ER • The smooth ER – Synthesizes lipids – Metabolizes carbohydrates – Detoxifies poison – Stores calcium Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings

Functions of Rough ER • The rough ER – Has bound ribosomes, which secrete glycoproteins (proteins covalently bonded to carbohydrates) – Distributes transport vesicles, proteins surrounded by membranes – Is a membrane factory for the cell Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings

The Golgi Apparatus: Shipping and Receiving Center • The Golgi apparatus consists of flattened membranous sacs called cisternae • Functions of the Golgi apparatus: – Modifies products of the ER – Manufactures certain macromolecules – Sorts and packages materials into transport vesicles Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings
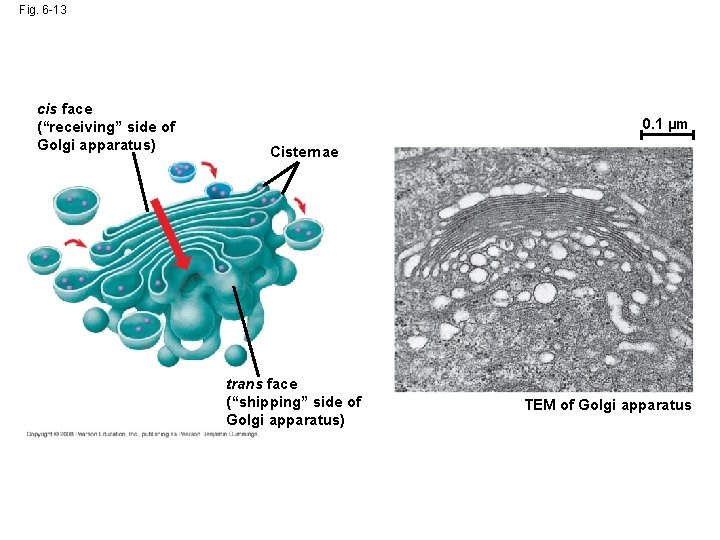
Fig. 6 -13 cis face (“receiving” side of Golgi apparatus) 0. 1 µm Cisternae trans face (“shipping” side of Golgi apparatus) TEM of Golgi apparatus

Lysosomes: Digestive Compartments • A lysosome is a membranous sac of hydrolytic enzymes that can digest macromolecules • Lysosomal enzymes can hydrolyze proteins, fats, polysaccharides, and nucleic acids Animation: Lysosome Formation Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings

• Some types of cell can engulf another cell by phagocytosis; this forms a food vacuole • A lysosome fuses with the food vacuole and digests the molecules • Lysosomes also use enzymes to recycle the cell’s own organelles and macromolecules, a process called autophagy Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings
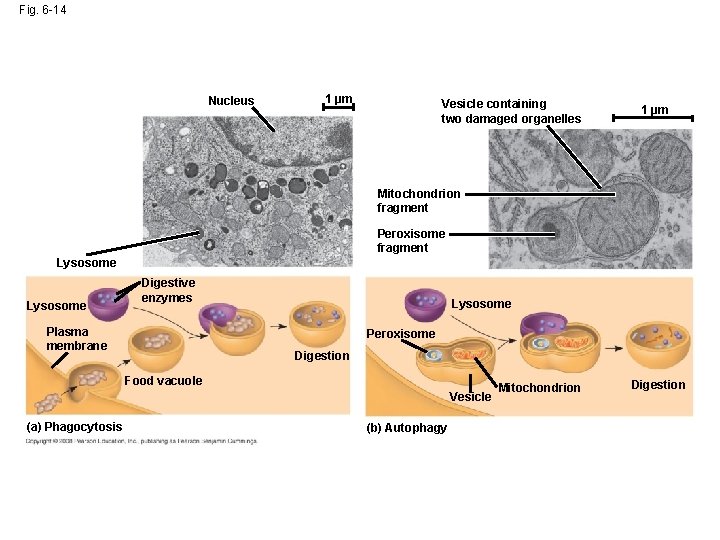
Fig. 6 -14 Nucleus 1 µm Vesicle containing two damaged organelles 1 µm Mitochondrion fragment Peroxisome fragment Lysosome Digestive enzymes Plasma membrane Lysosome Peroxisome Digestion Food vacuole Vesicle (a) Phagocytosis (b) Autophagy Mitochondrion Digestion

Vacuoles: Diverse Maintenance Compartments • A plant cell or fungal cell may have one or several vacuoles Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings

• Food vacuoles are formed by phagocytosis • Contractile vacuoles, found in many freshwater protists, pump excess water out of cells • Central vacuoles, found in many mature plant cells, hold organic compounds and water Video: Paramecium Vacuole Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings
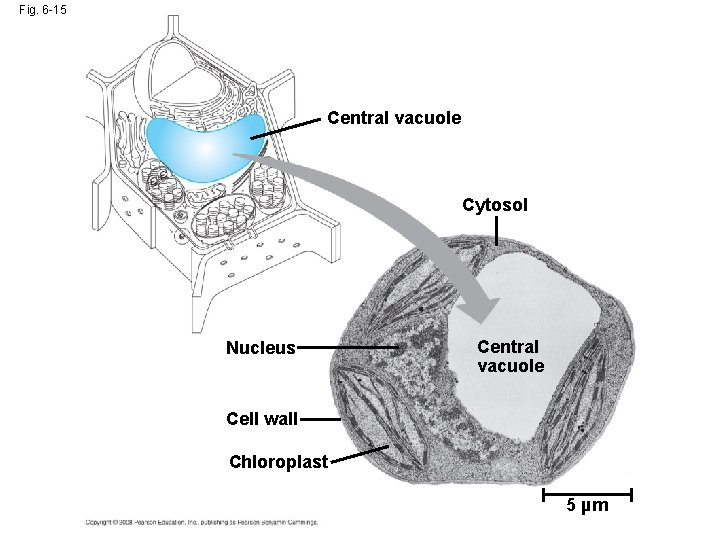
Fig. 6 -15 Central vacuole Cytosol Nucleus Central vacuole Cell wall Chloroplast 5 µm

SISTEM ENDOMEMBRAN KOMPARTEMEN INTRASELULER REGULASI PROTEIN PASCATRANSLASI

Polyribosomes • A number of ribosomes can translate a single m. RNA simultaneously, forming a polyribosome (or polysome) • Polyribosomes enable a cell to make many copies of a polypeptide very quickly Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
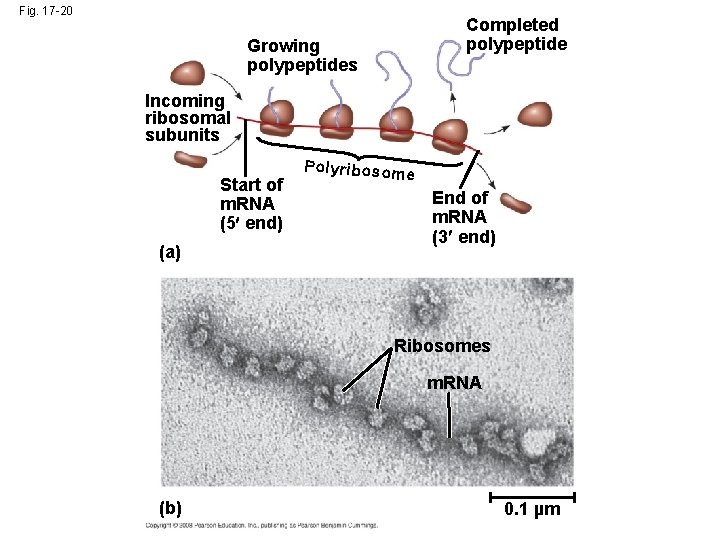
Fig. 17 -20 Completed polypeptide Growing polypeptides Incoming ribosomal subunits Start of m. RNA (5 end) (a) Polyribosom e End of m. RNA (3 end) Ribosomes m. RNA (b) 0. 1 µm

Completing and Targeting the Functional Protein • Often translation is not sufficient to make a functional protein • Polypeptide chains are modified after translation • Completed proteins are targeted to specific sites in the cell Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

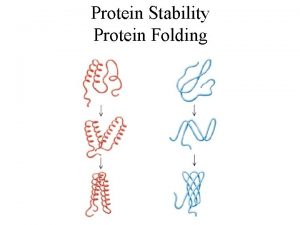
Protein Folding and Post-Translational Modifications • During and after synthesis, a polypeptide chain spontaneously coils and folds into its threedimensional (3 D) shape • Proteins may also require post-translational modifications before doing their job • Some polypeptides are activated by enzymes that cleave them • Other polypeptides come together to form the subunits of a protein Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
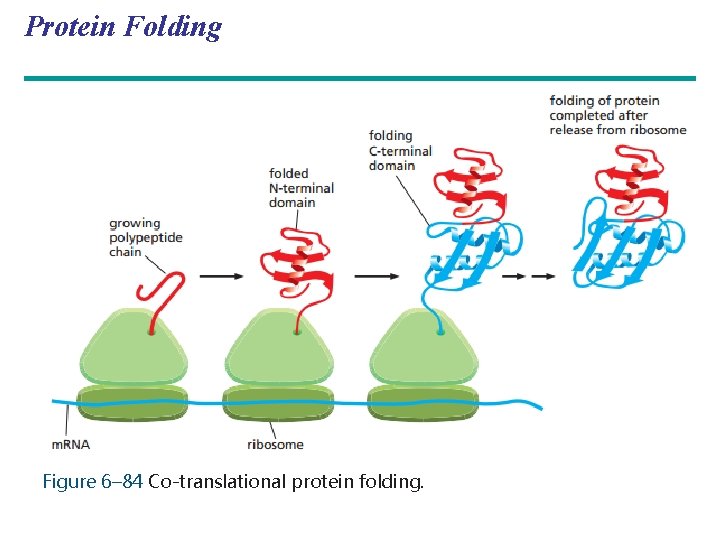
Protein Folding Figure 6– 84 Co-translational protein folding.
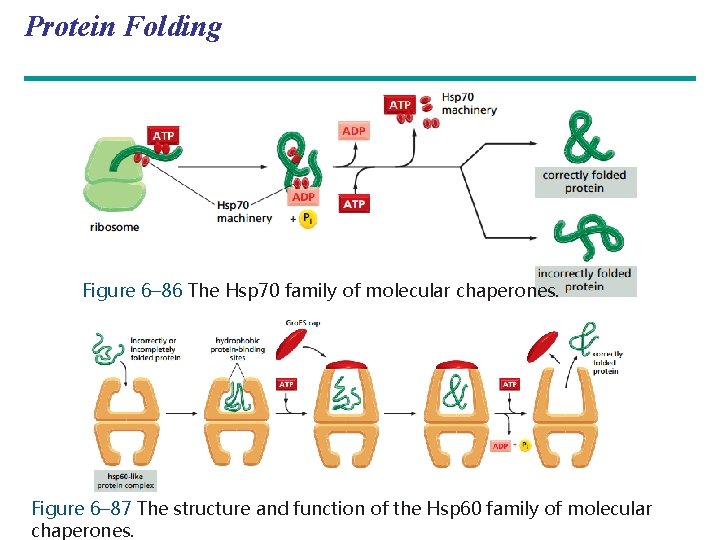
Protein Folding Figure 6– 86 The Hsp 70 family of molecular chaperones. Figure 6– 87 The structure and function of the Hsp 60 family of molecular chaperones.

Targeting Polypeptides to Specific Locations • Two populations of ribosomes are evident in cells: free ribsomes (in the cytosol) and bound ribosomes (attached to the ER) • Free ribosomes mostly synthesize proteins that function in the cytosol • Bound ribosomes make proteins of the endomembrane system and proteins that are secreted from the cell • Ribosomes are identical and can switch from free to bound Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

• Polypeptide synthesis always begins in the cytosol • Synthesis finishes in the cytosol unless the polypeptide signals the ribosome to attach to the ER • Polypeptides destined for the ER or for secretion are marked by a signal peptide • A signal-recognition particle (SRP) binds to the signal peptide • The SRP brings the signal peptide and its ribosome to the ER Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
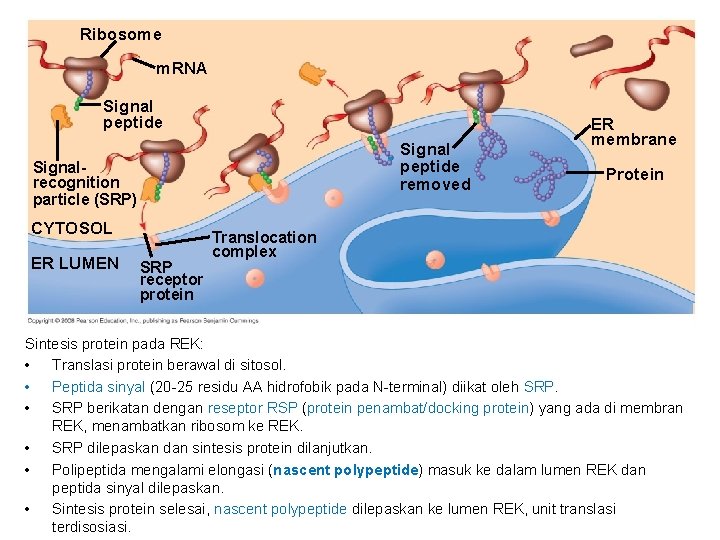
Ribosome m. RNA Signal peptide removed Signalrecognition particle (SRP) CYTOSOL ER LUMEN SRP receptor protein ER membrane Protein Translocation complex Sintesis protein pada REK: • Translasi protein berawal di sitosol. • Peptida sinyal (20 -25 residu AA hidrofobik pada N-terminal) diikat oleh SRP. • SRP berikatan dengan reseptor RSP (protein penambat/docking protein) yang ada di membran REK, menambatkan ribosom ke REK. • SRP dilepaskan dan sintesis protein dilanjutkan. • Polipeptida mengalami elongasi (nascent polypeptide) masuk ke dalam lumen REK dan peptida sinyal dilepaskan. • Sintesis protein selesai, nascent polypeptide dilepaskan ke lumen REK, unit translasi terdisosiasi.
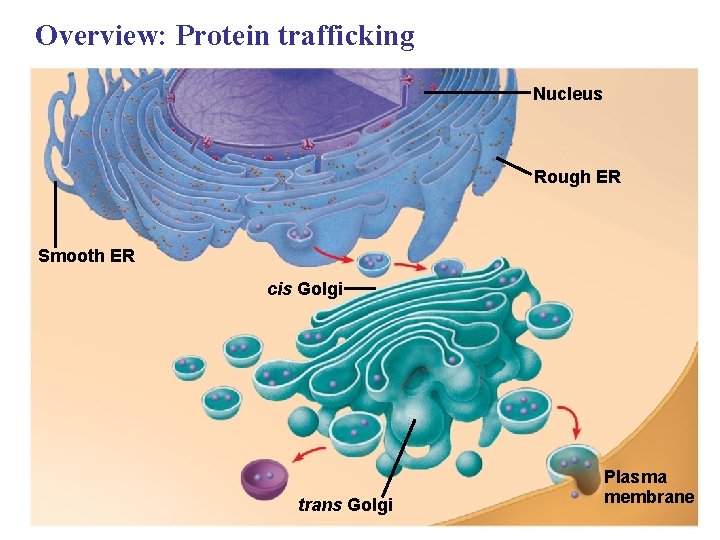
Overview: Protein trafficking Nucleus Rough ER Smooth ER cis Golgi trans Golgi Plasma membrane
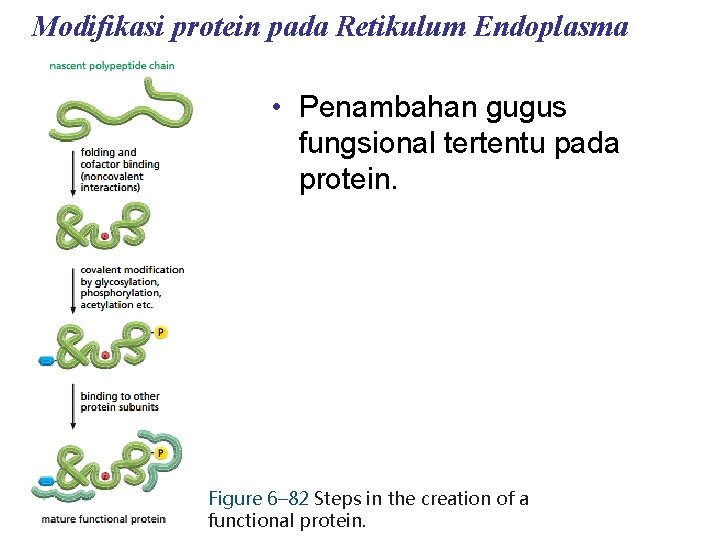
Modifikasi protein pada Retikulum Endoplasma • Penambahan gugus fungsional tertentu pada protein. Figure 6– 82 Steps in the creation of a functional protein.

To be continue… Take home activity 1. Modifikasi protein pada RE 2. Sortasi protein sekresi pada apparatus Golgi PPT print 6 slides/page; references
- Slides: 28