Sequence Alignment KunMao Chao Department of Computer Science






































- Slides: 56

Sequence Alignment Kun-Mao Chao (趙坤茂) Department of Computer Science and Information Engineering National Taiwan University, Taiwan E-mail: kmchao@csie. ntu. edu. tw WWW: http: //www. csie. ntu. edu. tw/~kmchao

Bioinformatics 2

Bioinformatics and Computational Biology. Related Journals: • • • Bioinformatics (previously called CABIOS) Bulletin of Mathematical Biology Computers and Biomedical Research Genome Research Genomics Journal of Bioinformatics and Computational Biology Journal of Molecular Biology Nature Nucleic Acid Research Science 3

Bioinformatics and Computational Biology. Related Conferences: • Intelligent Systems for Molecular Biology (ISMB) • Pacific Symposium on Biocomputing (PSB) • The Annual International Conference on Research in Computational Molecular Biology (RECOMB) • The IEEE Computer Society Bioinformatics Conference (CSB) • . . . 4

Bioinformatics and Computational Biology-Related Books: • Calculating the Secrets of Life: Applications of the Mathematical Sciences in Molecular Biology, by Eric S. Lander and Michael S. Waterman (1995) • Introduction to Computational Biology: Maps, Sequences, and Genomes, by Michael S. Waterman (1995) • Introduction to Computational Molecular Biology, by Joao Carlos Setubal and Joao Meidanis (1996) • Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology, by Dan Gusfield (1997) • Computational Molecular Biology: An Algorithmic Approach, by Pavel Pevzner (2000) • Introduction to Bioinformatics, by Arthur M. Lesk (2002) 5

Useful Websites • MIT Biology Hypertextbook – http: //www. mit. edu: 8001/afs/athena/course/other/esgbio/www/ 7001 main. html • The International Society for Computational Biology: – http: //www. iscb. org/ • National Center for Biotechnology Information (NCBI, NIH): – http: //www. ncbi. nlm. nih. gov/ • European Bioinformatics Institute (EBI): – http: //www. ebi. ac. uk/ • DNA Data Bank of Japan (DDBJ): – http: //www. ddbj. nig. ac. jp/ 6

Sequence Alignment 7

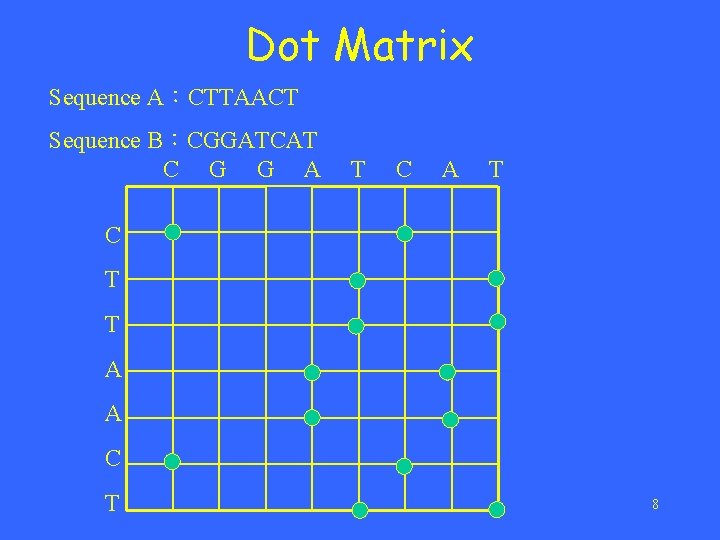
Dot Matrix Sequence A:CTTAACT Sequence B:CGGATCAT C G G A T C T T A A C T 8

Pairwise Alignment Sequence A: CTTAACT Sequence B: CGGATCAT An alignment of A and B: C---TTAACT CGGATCA--T Sequence A Sequence B 9

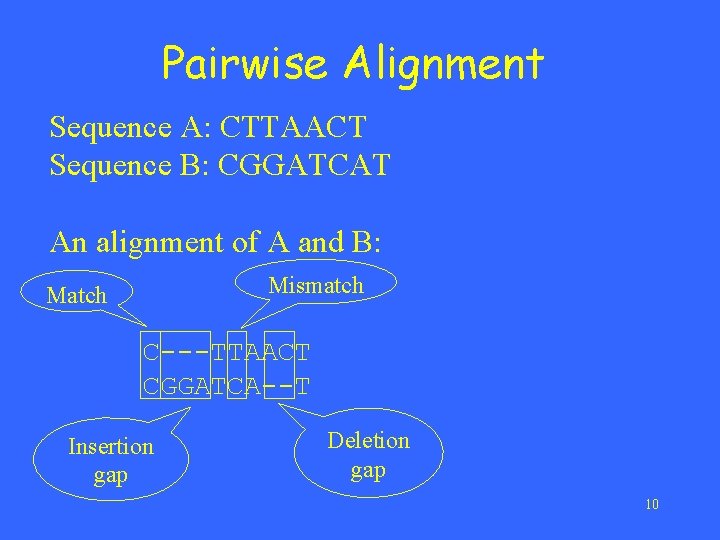
Pairwise Alignment Sequence A: CTTAACT Sequence B: CGGATCAT An alignment of A and B: Mismatch Match C---TTAACT CGGATCA--T Insertion gap Deletion gap 10

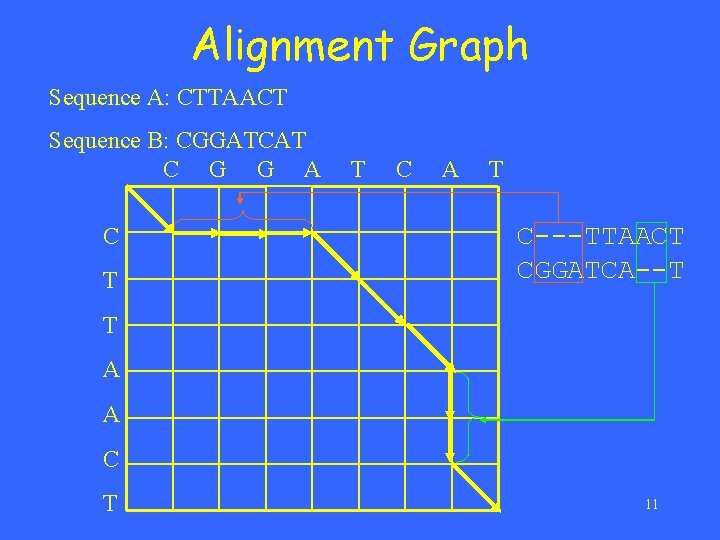
Alignment Graph Sequence A: CTTAACT Sequence B: CGGATCAT C G G A C T T C A T C---TTAACT CGGATCA--T T A A C T 11

A simple scoring scheme • Match: +8 (w(x, y) = 8, if x = y) • Mismatch: -5 (w(x, y) = -5, if x ≠ y) • Each gap symbol: -3 (w(-, x)=w(x, -)=-3) C - - - T T A A C T C G G A T C A - - T +8 -3 -3 -3 +8 -5 +8 -3 -3 Alignment score +8 = +12 12

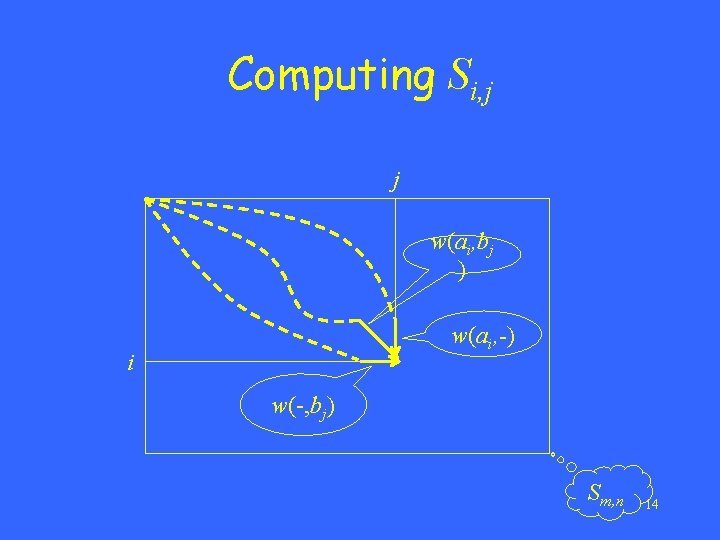
An optimal alignment -- the alignment of maximum score • Let A=a 1 a 2…am and B=b 1 b 2…bn. • Si, j: the score of an optimal alignment between a 1 a 2…ai and b 1 b 2…bj • With proper initializations, Si, j can be computed as follows. 13

Computing Si, j j w(ai, bj ) w(ai, -) i w(-, bj) Sm, n 14

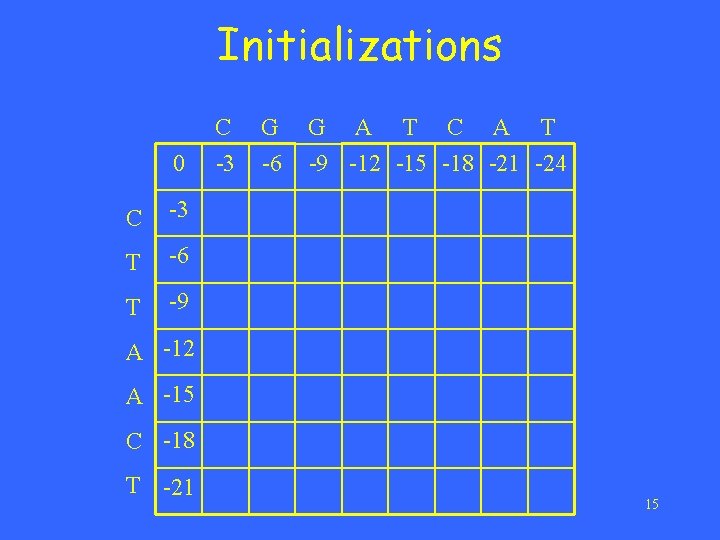
Initializations 0 C -3 T -6 T -9 C -3 G -6 G A T C A T -9 -12 -15 -18 -21 -24 A -12 A -15 C -18 T -21 15

S 3, 5 = ? 0 C -3 G -6 G A T C A T -9 -12 -15 -18 -21 -24 C -3 8 5 2 -1 -4 -7 -10 -13 T -6 5 3 0 -3 7 4 T -9 2 0 -2 -5 ? 1 -2 A -15 C -18 T -21 16

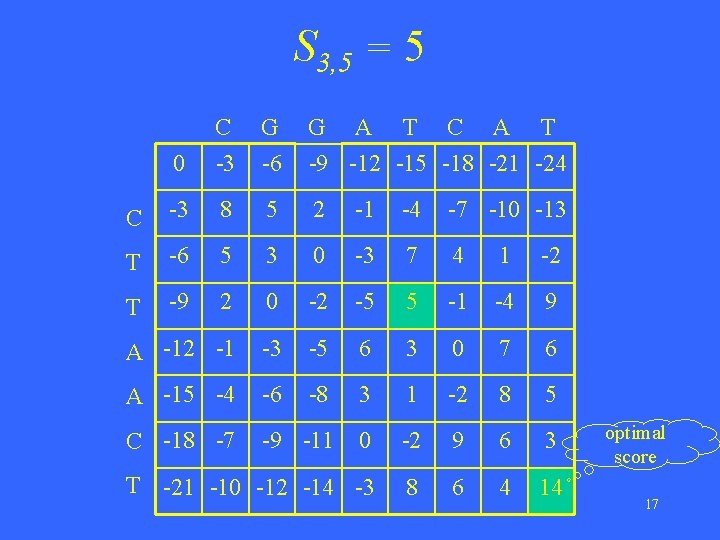
S 3, 5 = 5 0 C -3 G -6 G A T C A T -9 -12 -15 -18 -21 -24 C -3 8 5 2 -1 -4 -7 -10 -13 T -6 5 3 0 -3 7 4 1 -2 T -9 2 0 -2 -5 5 -1 -4 9 A -12 -1 -3 -5 6 3 0 7 6 A -15 -4 -6 -8 3 1 -2 8 5 C -18 -7 -9 -11 0 -2 9 6 3 T -21 -10 -12 -14 -3 8 6 4 14 optimal score 17

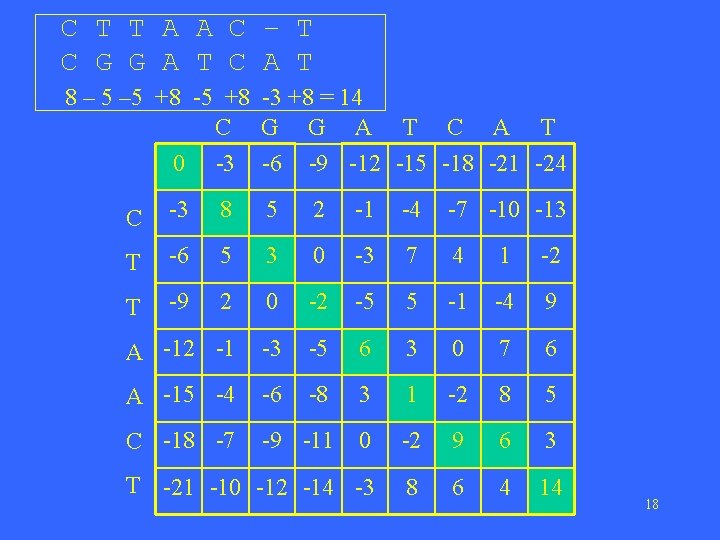
C T T A A C – T C G G A T C A T 8 – 5 +8 -5 +8 -3 +8 = 14 C G G A T C A T 0 -3 -6 -9 -12 -15 -18 -21 -24 C -3 8 5 2 -1 -4 -7 -10 -13 T -6 5 3 0 -3 7 4 1 -2 T -9 2 0 -2 -5 5 -1 -4 9 A -12 -1 -3 -5 6 3 0 7 6 A -15 -4 -6 -8 3 1 -2 8 5 C -18 -7 -9 -11 0 -2 9 6 3 T -21 -10 -12 -14 -3 8 6 4 14 18

Now try this example in class Sequence A: CAATTGA Sequence B: GAATCTGC Their optimal alignment? 19

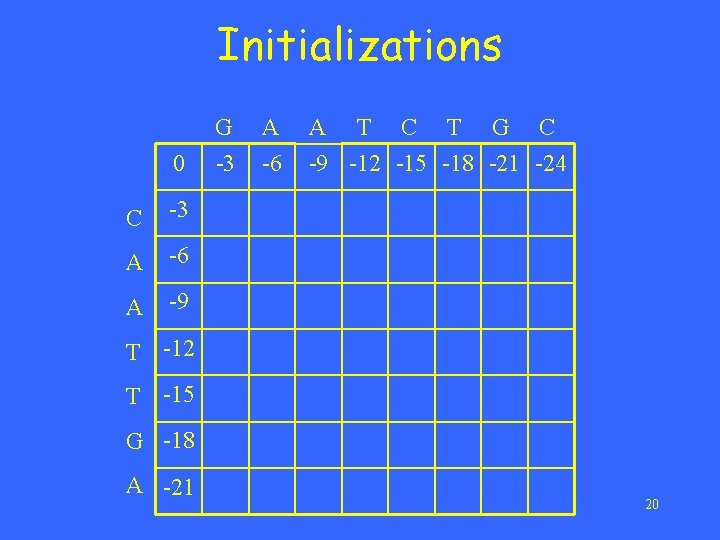
Initializations 0 C -3 A -6 A -9 G -3 A -6 A T C T G C -9 -12 -15 -18 -21 -24 T -12 T -15 G -18 A -21 20

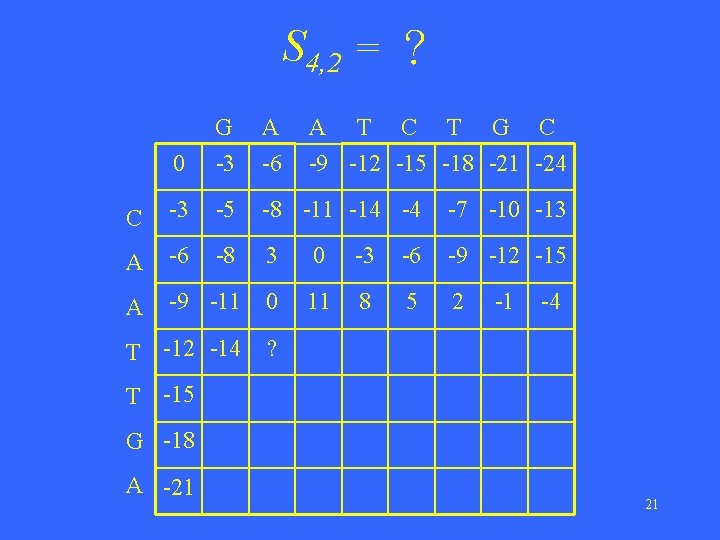
S 4, 2 = ? 0 G -3 A -6 A T C T G C -9 -12 -15 -18 -21 -24 C -3 -5 -8 -11 -14 -4 -7 -10 -13 A -6 -8 3 0 -3 -6 -9 -12 -15 A -9 -11 0 11 8 5 2 T -12 -14 ? -1 -4 T -15 G -18 A -21 21

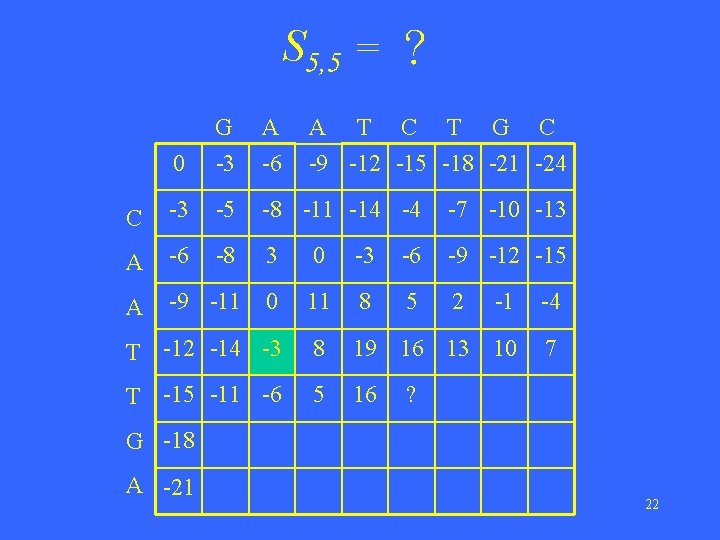
S 5, 5 = ? 0 G -3 A -6 A T C T G C -9 -12 -15 -18 -21 -24 C -3 -5 -8 -11 -14 -4 -7 -10 -13 A -6 -8 3 0 -3 -6 -9 -12 -15 A -9 -11 0 11 8 5 2 -1 -4 T -12 -14 -3 8 19 16 13 10 7 T -15 -11 -6 5 16 ? G -18 A -21 22

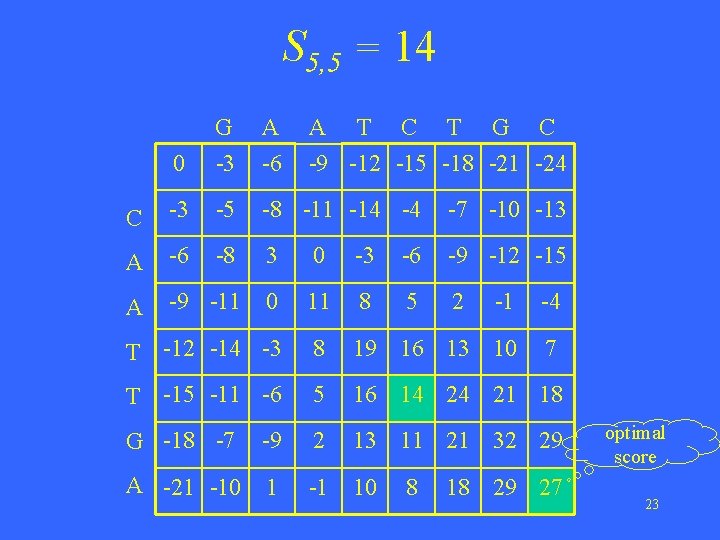
S 5, 5 = 14 0 G -3 A -6 A T C T G C -9 -12 -15 -18 -21 -24 C -3 -5 -8 -11 -14 -4 -7 -10 -13 A -6 -8 3 0 -3 -6 -9 -12 -15 A -9 -11 0 11 8 5 2 -1 -4 T -12 -14 -3 8 19 16 13 10 7 T -15 -11 -6 5 16 14 24 21 18 G -18 -7 -9 2 13 11 21 32 29 A -21 -10 1 -1 10 8 18 29 27 optimal score 23

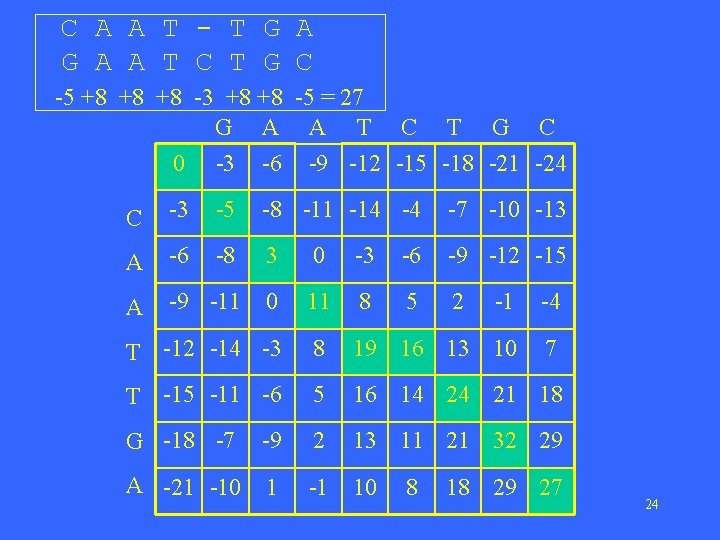
C A A T - T G A A T C T G C -5 +8 +8 +8 -3 +8 +8 -5 = 27 G A A T C T G C 0 -3 -6 -9 -12 -15 -18 -21 -24 C -3 -5 -8 -11 -14 -4 -7 -10 -13 A -6 -8 3 0 -3 -6 -9 -12 -15 A -9 -11 0 11 8 5 2 -1 -4 T -12 -14 -3 8 19 16 13 10 7 T -15 -11 -6 5 16 14 24 21 18 G -18 -7 -9 2 13 11 21 32 29 A -21 -10 1 -1 10 8 18 29 27 24

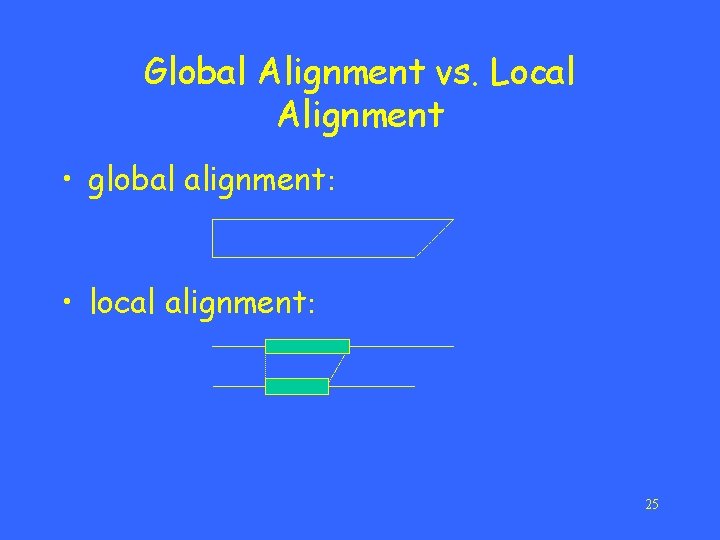
Global Alignment vs. Local Alignment • global alignment: • local alignment: 25

An optimal local alignment • Si, j: the score of an optimal local alignment ending at ai and bj • With proper initializations, Si, j can be computed as follows. 26

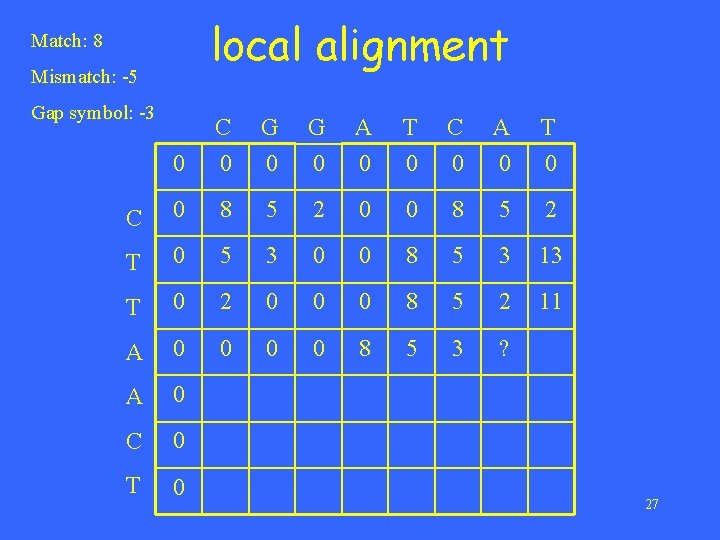
local alignment Match: 8 Mismatch: -5 Gap symbol: -3 0 C 0 G 0 A 0 T 0 C 0 8 5 2 0 0 8 5 2 T 0 5 3 0 0 8 5 3 13 T 0 2 0 0 0 8 5 2 11 A 0 0 8 5 3 ? A 0 C 0 T 0 27

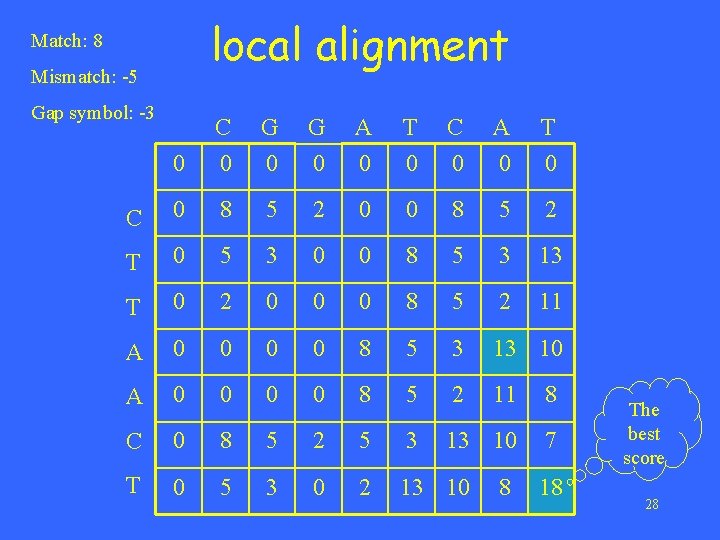
local alignment Match: 8 Mismatch: -5 Gap symbol: -3 0 C 0 G 0 A 0 T 0 C 0 8 5 2 0 0 8 5 2 T 0 5 3 0 0 8 5 3 13 T 0 2 0 0 0 8 5 2 11 A 0 0 8 5 3 13 10 A 0 0 8 5 2 11 8 C 0 8 5 2 5 3 13 10 7 T 0 5 3 0 2 13 10 8 18 The best score 28

A – C - T A T C A T 8 -3+8 = 18 C G 0 0 0 G 0 A 0 T 0 C 0 8 5 2 0 0 8 5 2 T 0 5 3 0 0 8 5 3 13 T 0 2 0 0 0 8 5 2 11 A 0 0 8 5 3 13 10 A 0 0 8 5 2 11 8 C 0 8 5 2 5 3 13 10 7 T 0 5 3 0 2 13 10 8 18 The best score 29

Now try this example in class Sequence A: CAATTGA Sequence B: GAATCTGC Their optimal local alignment? 30

Did you get it right? 0 G 0 A 0 T 0 C 0 T 0 G 0 C 0 0 0 8 5 2 8 A 0 0 8 8 5 5 3 0 5 A 0 0 8 16 13 10 7 4 1 T 0 0 5 13 24 21 18 15 12 T 0 0 2 10 21 19 29 26 23 G 0 8 5 7 A 0 5 16 13 15 13 23 34 32 18 16 26 37 34 31

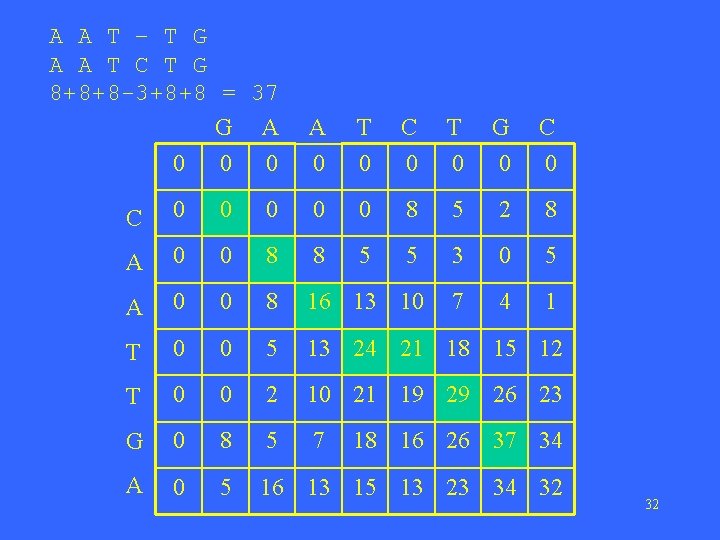
A A T – T G A A T C T G 8+8+8 -3+8+8 = 37 G A 0 0 0 A 0 T 0 C 0 T 0 G 0 C 0 0 0 8 5 2 8 A 0 0 8 8 5 5 3 0 5 A 0 0 8 16 13 10 7 4 1 T 0 0 5 13 24 21 18 15 12 T 0 0 2 10 21 19 29 26 23 G 0 8 5 7 A 0 5 16 13 15 13 23 34 32 18 16 26 37 34 32

Affine gap penalties • • Match: +8 (w(x, y) = 8, if x = y) Mismatch: -5 (w(x, y) = -5, if x ≠ y) Each gap symbol: -3 (w(-, x)=w(x, -)=-3) Each gap is charged an extra gap-open penalty: -4. -4 -4 C - - - T T A A C T C G G A T C A - - T +8 -3 -3 -3 +8 -5 +8 -3 -3 +8 = +12 Alignment score: 12 – 4 = 4 33

Affine gap panalties • A gap of length k is penalized x + k·y. gap-open penalty Three cases for alignment endings: gap-symbol penalty 1. . x an aligned pair 2. . x. . . - a deletion 3. . . . x an insertion 34

Affine gap penalties • Let D(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj ending with a deletion. • Let I(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj ending with an insertion. • Let S(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj. 35

Affine gap penalties (A gap of length k is penalized x + k·y. ) 36

Affine gap penalties D I D S I -y w(ai, bj) -x-y D I S S D -x-y -y I S 37

Constant gap penalties • • Match: +8 (w(x, y) = 8, if x = y) Mismatch: -5 (w(x, y) = -5, if x ≠ y) Each gap symbol: 0 (w(-, x)=w(x, -)=0) Each gap is charged a constant penalty: -4. -4 -4 C - - - T T A A C T C G G A T C A - - T +8 0 0 0 +8 -5 +8 0 0 +8 = +27 Alignment score: 27 – 4 = 19 38

Constant gap penalties • Let D(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj ending with a deletion. • Let I(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj ending with an insertion. • Let S(i, j) denote the maximum score of any alignment between a 1 a 2…ai and b 1 b 2…bj. 39

Constant gap penalties 40

Restricted affine gap panalties • A gap of length k is penalized x + f(k)·y. where f(k) = k for k <= c and f(k) = c for k > c Five cases for alignment endings: 1. . x an aligned pair 2. . x. . . - a deletion 3. . . . x an insertion 4. and 5. for long gaps 41

Restricted affine gap penalties 42

D(i, j) vs. D’(i, j) • Case 1: the best alignment ending at (i, j) with a deletion at the end has the last deletion gap of length <= c D(i, j) >= D’(i, j) • Case 2: the best alignment ending at (i, j) with a deletion at the end has the last deletion gap of length >= c D(i, j) <= D’(i, j) 43

k best local alignments • Smith-Waterman (Smith and Waterman, 1981; Waterman and Eggert, 1987) • FASTA (Wilbur and Lipman, 1983; Lipman and Pearson, 1985) • BLAST (Altschul et al. , 1990; Altschul et al. , 1997) 44

FASTA 1) Find runs of identities, and identify regions with the highest density of identities. 2) Re-score using PAM matrix, and keep top scoring segments. 3) Eliminate segments that are unlikely to be part of the alignment. 4) Optimize the alignment in a band. 45

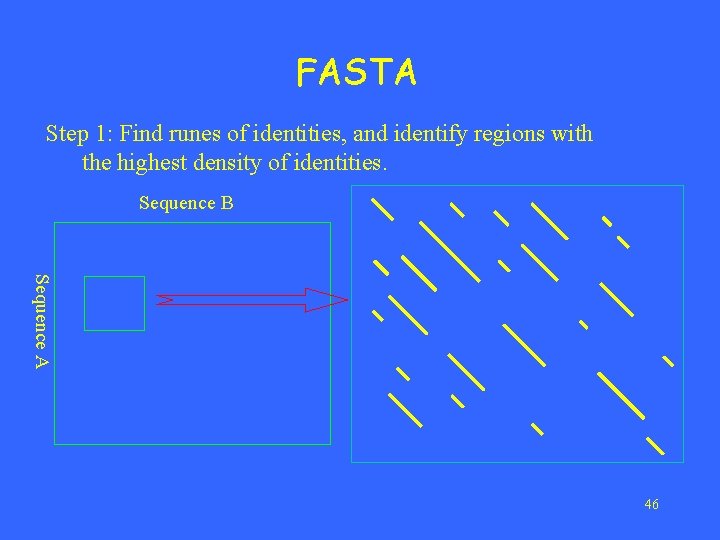
FASTA Step 1: Find runes of identities, and identify regions with the highest density of identities. Sequence B Sequence A 46

FASTA Step 2: Re-score using PAM matrix, and keep top scoring segments. 47

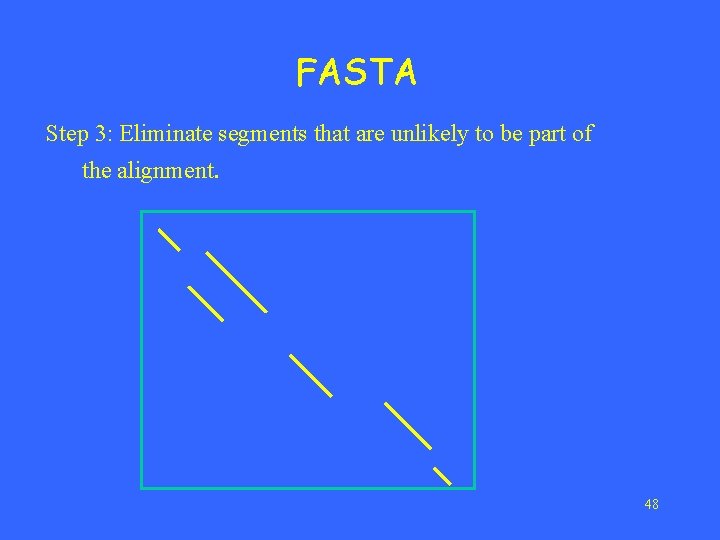
FASTA Step 3: Eliminate segments that are unlikely to be part of the alignment. 48

FASTA Step 4: Optimize the alignment in a band. 49

BLAST ü Basic Local Alignment Search Tool (by Altschul, Gish, Miller, Myers and Lipman) ü The central idea of the BLAST algorithm is that a statistically significant alignment is likely to contain a high-scoring pair of aligned words. 50

The maximal segment pair measure ü A maximal segment pair (MSP) is defined to be the highest scoring pair of identical length segments chosen from 2 sequences. (for DNA: Identities: +5; Mismatches: -4) the highest scoring pair • The MSP score may be computed in time proportional to the product of their lengths. (How? ) An exact procedure is too time consuming. • BLAST heuristically attempts to calculate the MSP score. 51

BLAST 1) Build the hash table for Sequence A. 2) Scan Sequence B for hits. 3) Extend hits. 52

BLAST Step 1: Build the hash table for Sequence A. (3 -tuple example) For protein sequences: For DNA sequences: Seq. A = AGATCGAT 12345678 AAA AAC. . AGA. . ATC. . CGA. . GAT. . TCG. . TTT 1 3 5 2 Seq. A = ELVIS Add xyz to the hash table if Score(xyz, ELV) ≧ T; Add xyz to the hash table if Score(xyz, LVI) ≧ T; Add xyz to the hash table if Score(xyz, VIS) ≧ T; 6 4 53

BLAST Step 2: Scan sequence B for hits. 54

BLAST Step 2: Scan sequence B for hits. Step 3: Extend hits. BLAST 2. 0 saves the time spent in extension, and hit considers gapped alignments. Terminate if the score of the sxtension fades away. (That is, when we reach a segment pair whose score falls a certain distance below the 55 best score found for shorter extensions. )

Remarks • Filtering is based on the observation that a good alignment usually includes short identical or very similar fragments. • The idea of filtration was used in both FASTA and BLAST. 56