Rich Probabilistic Methods for Gene Expression Eran Segal

Rich Probabilistic Methods for Gene Expression Eran Segal Ben Taskar Audrey Gasch Nir Friedman Daphne Koller

Outline • Motivation for richer models • PRMs for gene expression Modeling Learning Inference • Results Synthetic Stress Compendium
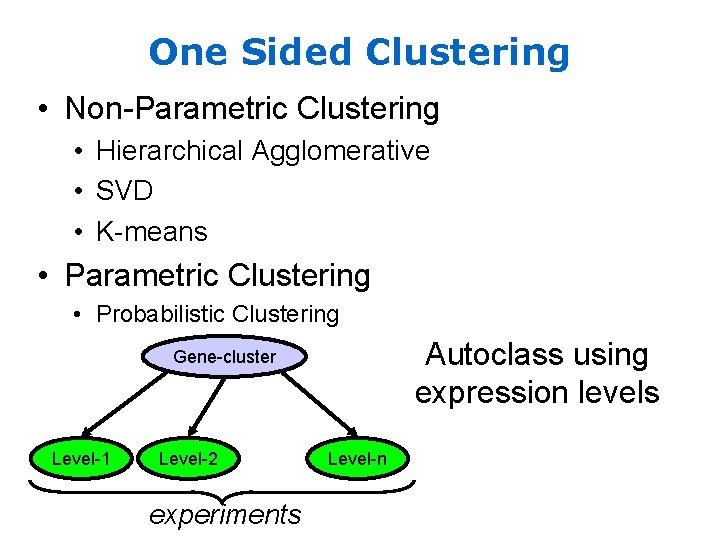
One Sided Clustering • Non-Parametric Clustering • Hierarchical Agglomerative • SVD • K-means • Parametric Clustering • Probabilistic Clustering Autoclass using expression levels Gene-cluster Level-1 Level-2 experiments Level-n
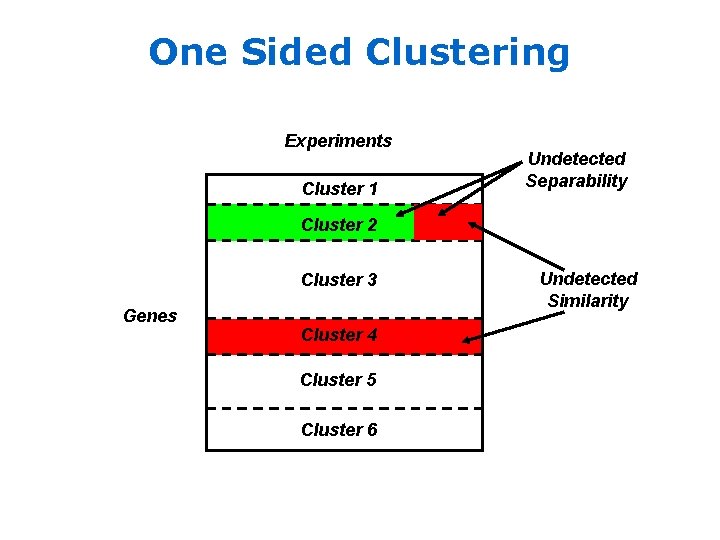
One Sided Clustering Experiments Cluster 1 Undetected Separability Cluster 2 Cluster 3 Genes Cluster 4 Cluster 5 Cluster 6 Undetected Similarity
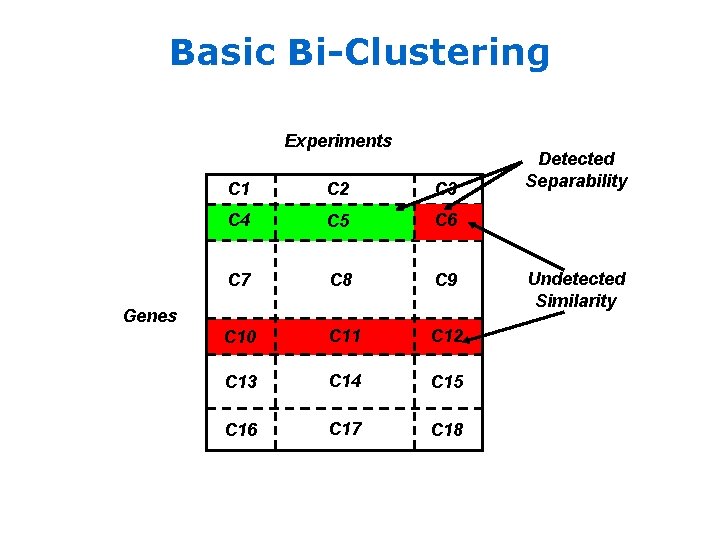
Basic Bi-Clustering Experiments C 1 C 2 C 3 C 4 C 5 C 6 C 7 C 8 C 9 C 10 C 11 C 12 C 13 C 14 C 15 C 16 C 17 C 18 Genes Detected Separability Undetected Similarity
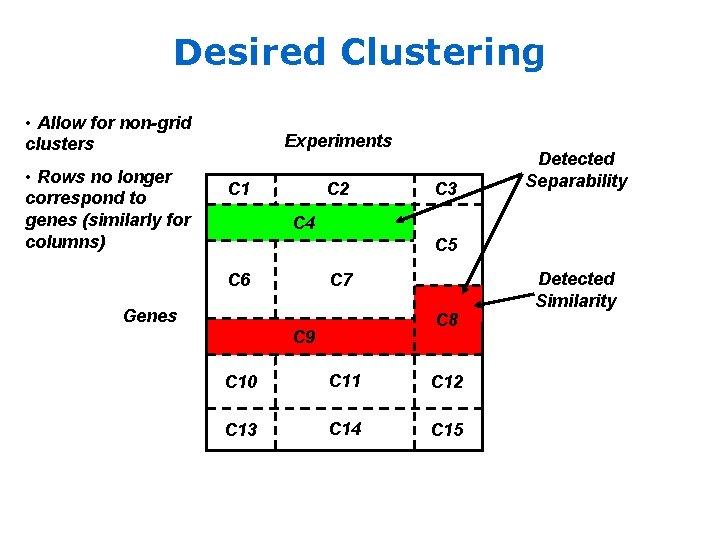
Desired Clustering • Allow for non-grid clusters • Rows no longer correspond to genes (similarly for columns) Experiments C 1 C 2 C 3 Detected Separability C 4 C 5 C 6 C 7 Genes C 8 C 9 C 10 C 11 C 12 C 13 C 14 C 15 Detected Similarity

Outline • Motivation for richer models • PRMs for gene expression Modeling Learning Inference • Results Synthetic Stress Compendium

PRMs: Relational Schema • Describes the types of objects and relations in the database Gene Mutation Cluster Binding Sites Functional Classes Experiment Cluster Exp. Attributes Expression Exp. Level
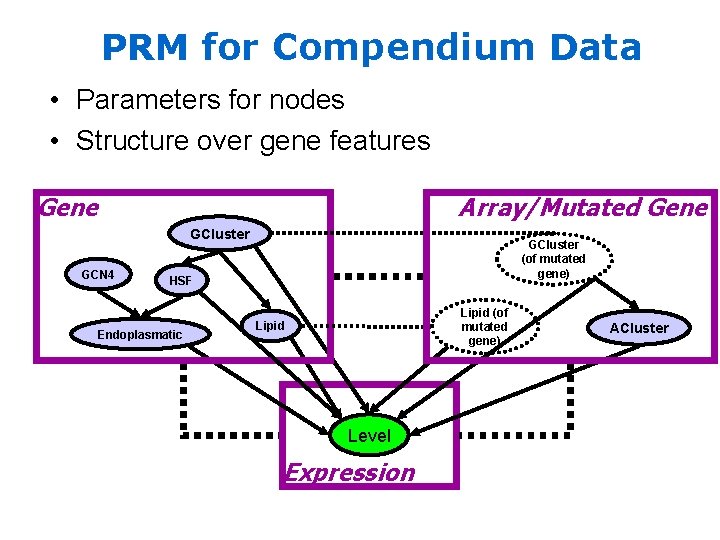
PRM for Compendium Data • Parameters for nodes • Structure over gene features Gene Array/Mutated Gene GCluster GCN 4 GCluster (of mutated gene) HSF Endoplasmatic Lipid (of mutated gene) Lipid Level Expression ACluster
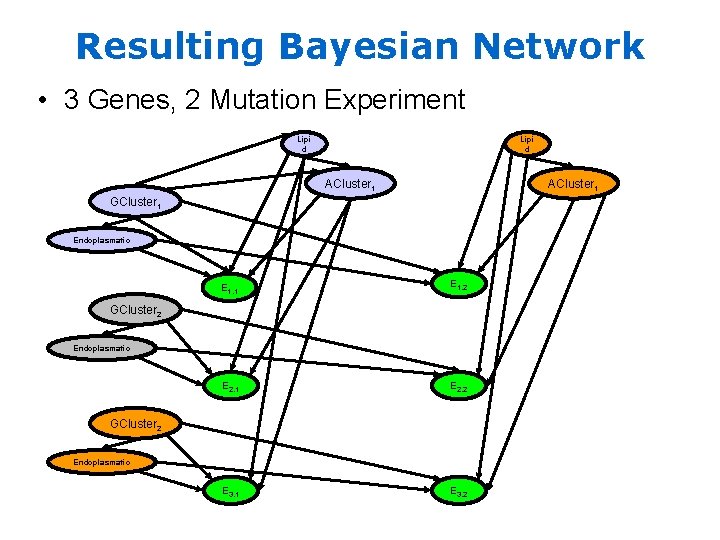
Resulting Bayesian Network • 3 Genes, 2 Mutation Experiment Lipi d ACluster 1 GCluster 1 Endoplasmatic E 1, 1 E 1, 2 E 2, 1 E 2, 2 E 3, 1 E 3, 2 GCluster 2 Endoplasmatic
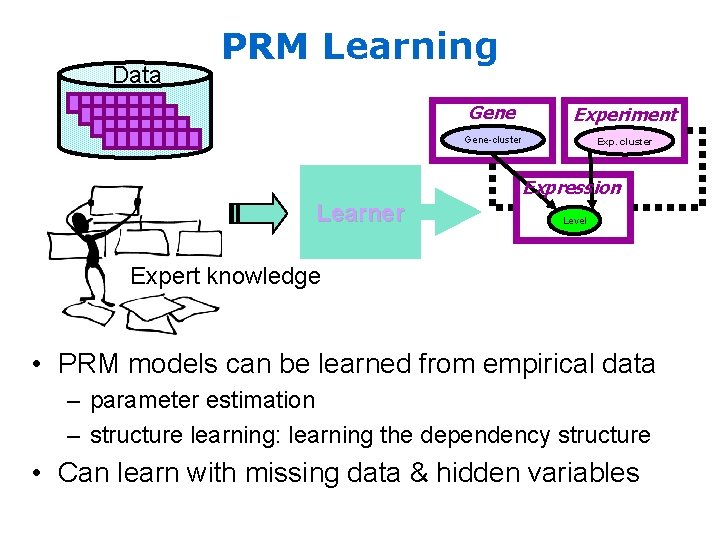
Data PRM Learning Gene Experiment Gene-cluster Expression Learner Level Expert knowledge • PRM models can be learned from empirical data – parameter estimation – structure learning: learning the dependency structure • Can learn with missing data & hidden variables

PRM Learning • Goal: Find PRM structure that explains the data well • Define scoring function to evaluate models – Bayesian Score works best Marginal likelihood Prior Score (S: D) = log [ P(D | S) P(S) ] – Automatically trades off fit to data (likelihood of data) with model complexity • Do heuristic search to find high-scoring structure – Structure found is not necessarily best one…
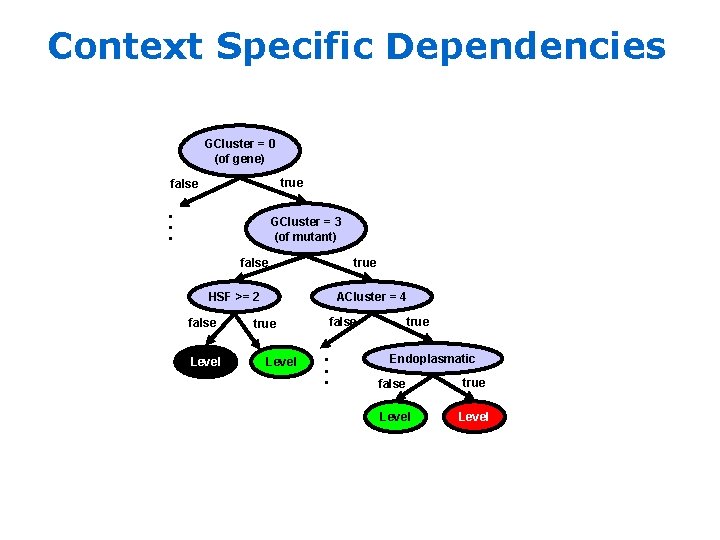
Context Specific Dependencies GCluster = 0 (of gene) true . . . false GCluster = 3 (of mutant) false true HSF >= 2 Level true false . . . false ACluster = 4 Endoplasmatic false true Level
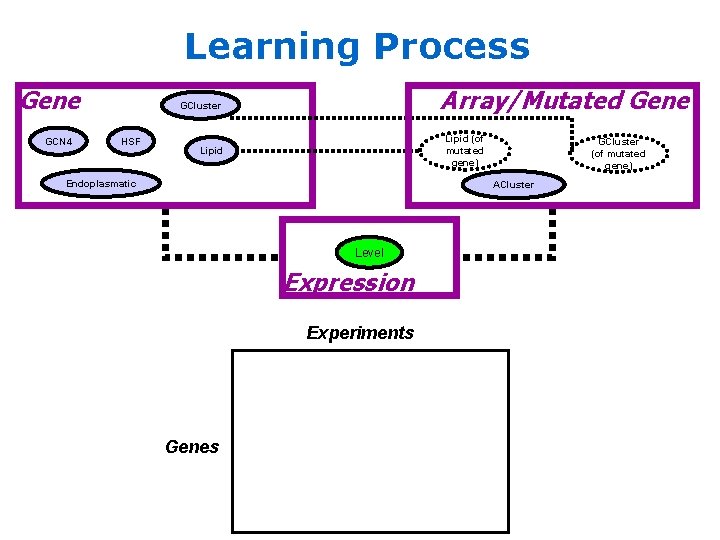
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic GCluster (of mutated gene) ACluster Level Expression Experiments Genes
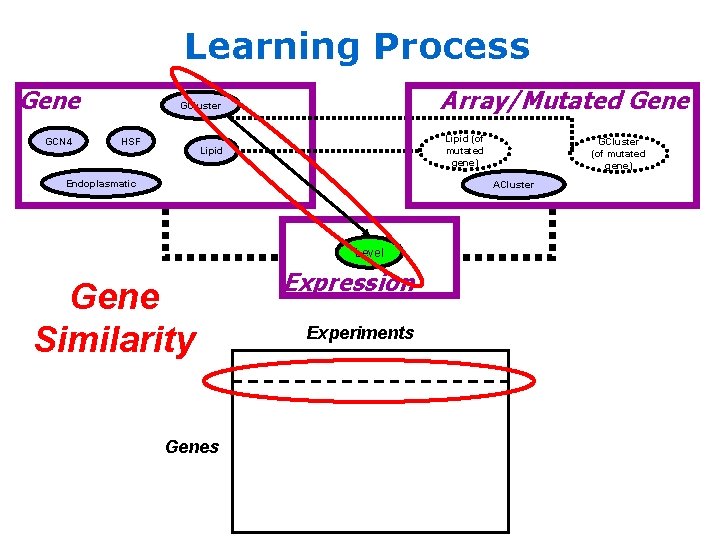
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic GCluster (of mutated gene) ACluster Level Gene Similarity Genes Expression Experiments
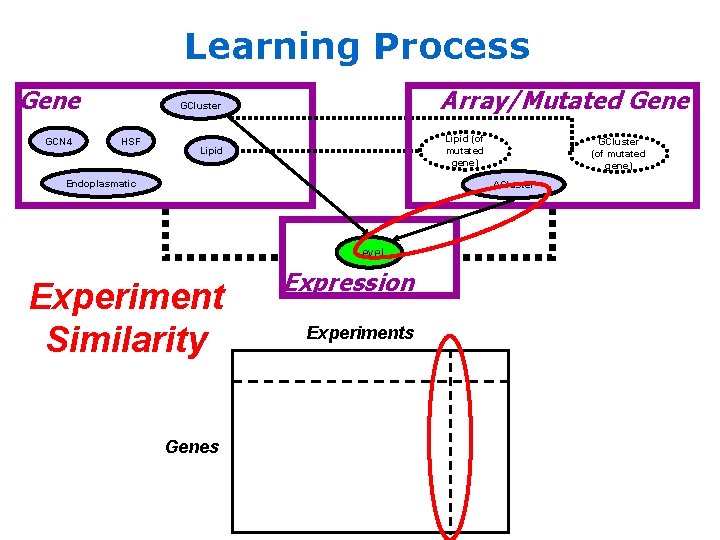
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic GCluster (of mutated gene) ACluster Level Experiment Similarity Genes Expression Experiments
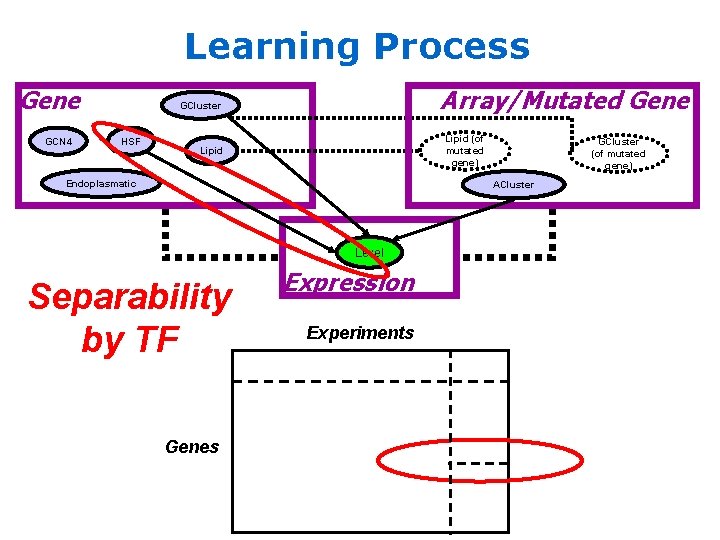
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic GCluster (of mutated gene) ACluster Level Separability by TF Genes Expression Experiments
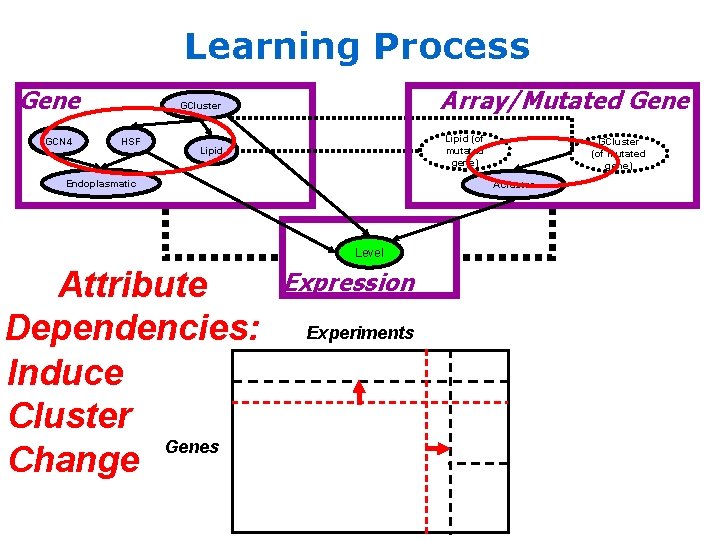
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic ACluster Level Attribute Dependencies: Induce Cluster Genes Change GCluster (of mutated gene) Expression Experiments
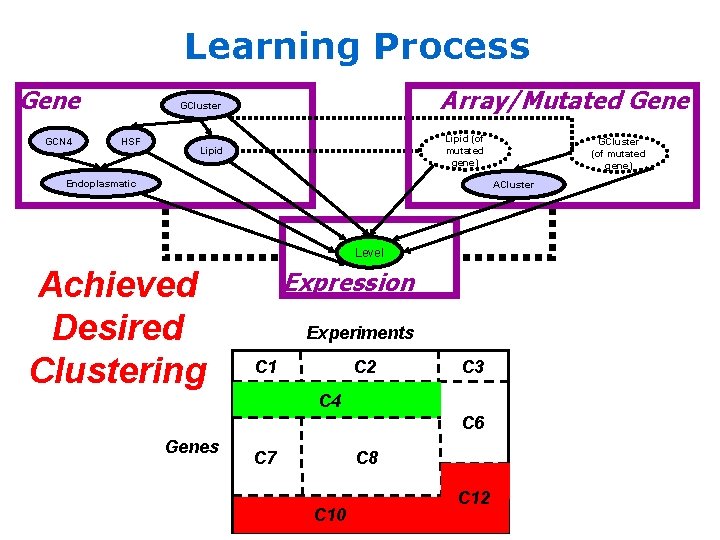
Learning Process Gene GCN 4 Array/Mutated Gene GCluster HSF Lipid (of mutated gene) Lipid Endoplasmatic GCluster (of mutated gene) ACluster Level Achieved Desired Clustering Expression Experiments C 1 C 2 C 3 C 4 C 6 Genes C 7 C 8 C 10 C 12

Outline • Motivation for richer models • PRMs for gene expression Modeling Learning Inference • Results Synthetic Stress Compendium

Synthetic Data: Recovering Structure • Synthetic data: 1000 genes, 90 arrays (12 types) • Parents recovered Simulated data: 84. 5% +/- 2. 5% Permuted data: 56% +/- 2. 5% • Cluster recovery Simulated data: PRMs: 98. 4% +/- 1. 07% Naïve Bayes: 90. 8% +/- 0. 42% Permuted data: PRMs: 88. 1% +/- 1. 52% Naïve Bayes: 76. 7% +/- 1. 42%

Stress Data • 954 genes, 88 arrays (12 types) • Structure learning 15 significant TFs + 7 significant function categories • Cluster coherence Average variance reduction 0. 69 -> 0. 61 in 3 iterations • Allowing annotation changes Average variance reduction 0. 69 -> 0. 56 in 3 iterations
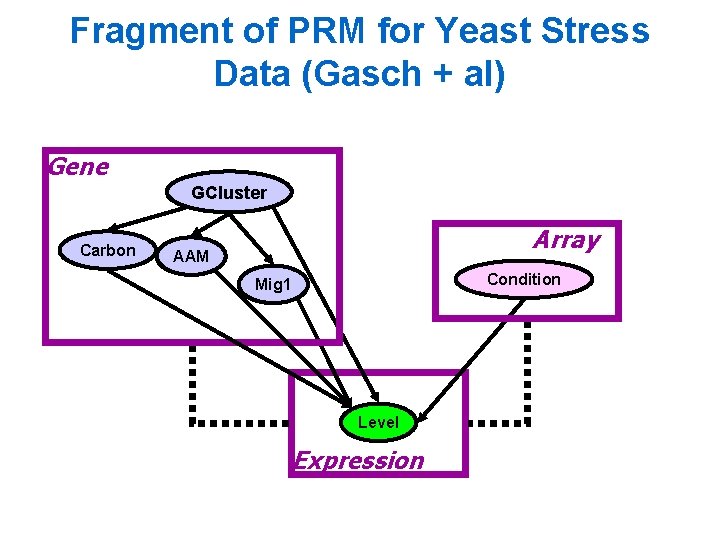
Fragment of PRM for Yeast Stress Data (Gasch + al) Gene GCluster Carbon Array AAM Condition Mig 1 Level Expression

Result: Context-Specific Groupings • A grouping is a set of genes that behave the same within a certain context — a condition or a set of conditions • Breakdown of genes into clusters is different in different contexts Yeast Stress Data (Gasch + al)

Example Biological Result • Discovered grouping of 17 genes – all induced in diauxic shift – all have 2 binding sites for Mig 1 transcription factor – many not known to have been regulated by Mig 1 • Context-sensitive groupings were key to identifying cluster
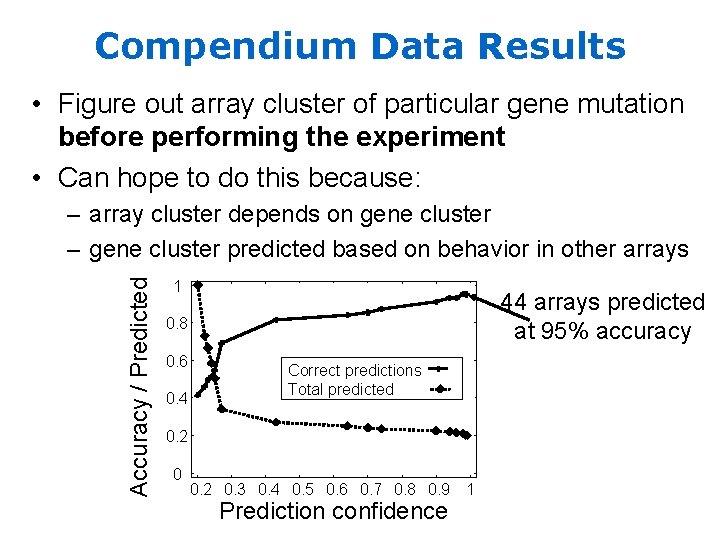
Compendium Data Results • Figure out array cluster of particular gene mutation before performing the experiment • Can hope to do this because: Accuracy / Predicted – array cluster depends on gene cluster – gene cluster predicted based on behavior in other arrays 1 44 arrays predicted at 95% accuracy 0. 8 0. 6 0. 4 Correct predictions Total predicted 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9 Prediction confidence 1

Future Directions • Handling time • Handling sequence data (TFs) • Incorporate structure information • Discovering pathways
- Slides: 27