Rich Probabilistic Models for Gene Expression Eran Segal

Rich Probabilistic Models for Gene Expression Eran Segal (Stanford) Ben Taskar (Stanford) Audrey Gasch (Berkeley) Nir Friedman (Hebrew University) Daphne Koller (Stanford)

Our Goals u Find patterns in gene expression data
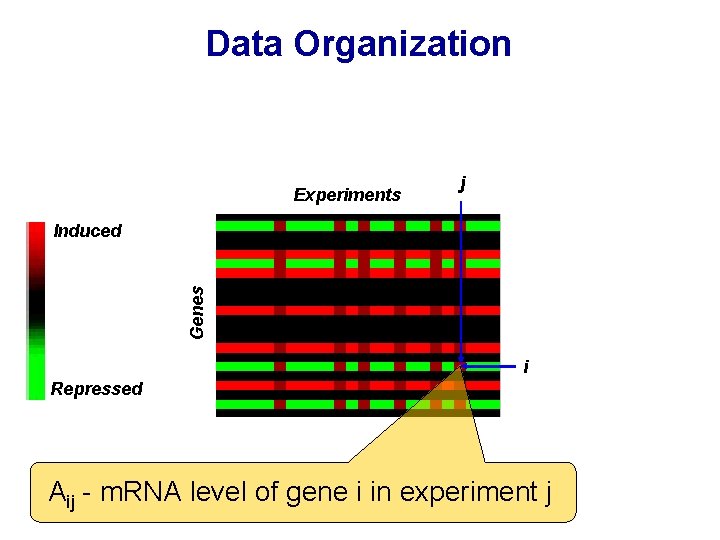
Data Organization Experiments j Genes Induced i Repressed Aij - m. RNA level of gene i in experiment j
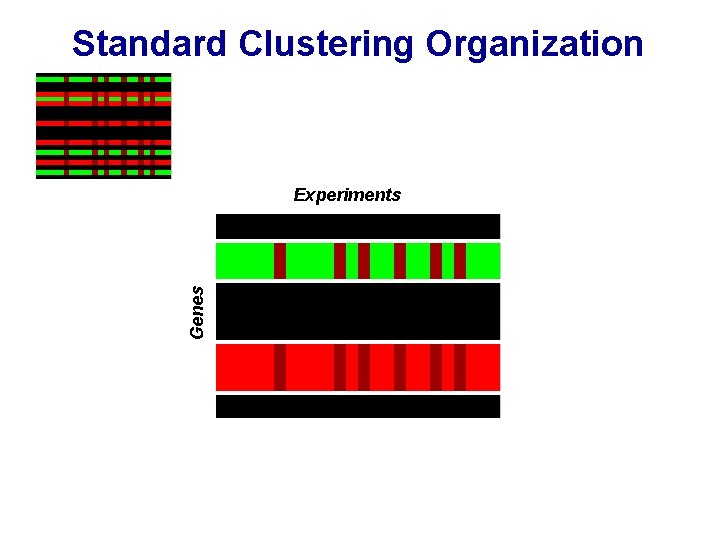
Standard Clustering Organization Genes Experiments
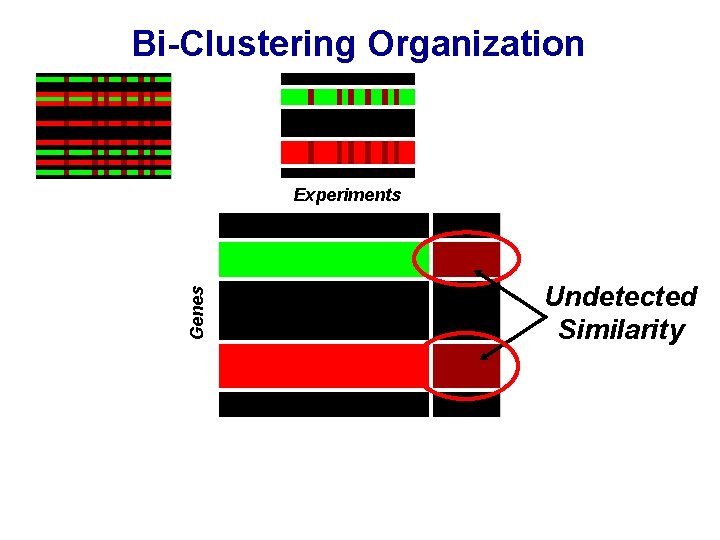
Bi-Clustering Organization Genes Experiments Undetected Similarity
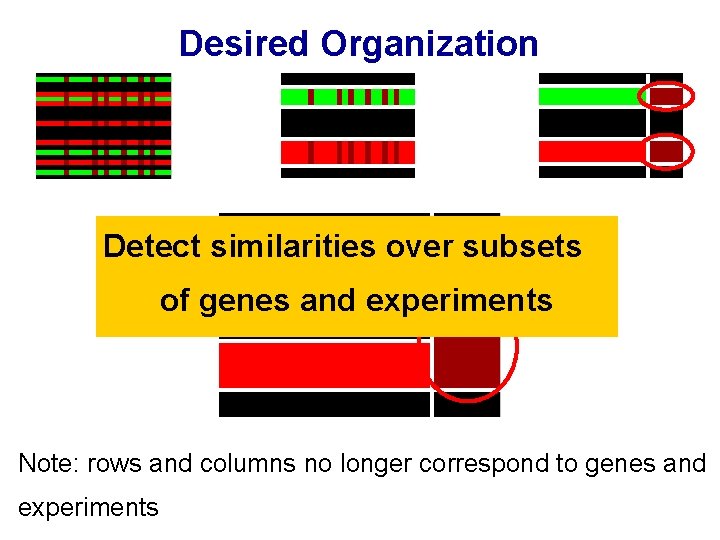
Desired Organization Detect similarities over subsets of genes and experiments Note: rows and columns no longer correspond to genes and experiments
Incorporate Heterogeneous Data A CG CT A Clinical information C u. Find correlations directly u. Focus Annotations (GO, MIPS, YPD) on novel discoveries Experimental Details
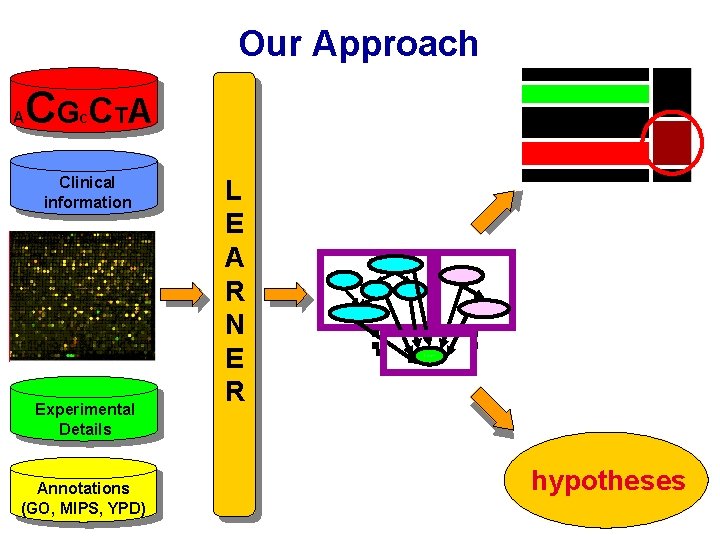
Our Approach A CG CT A C Clinical information Experimental Details Annotations (GO, MIPS, YPD) L E A R N E R Gene Cluster Exp. type GCN 4 HSF Lipid Exp. cluster Endoplasmatic Level hypotheses
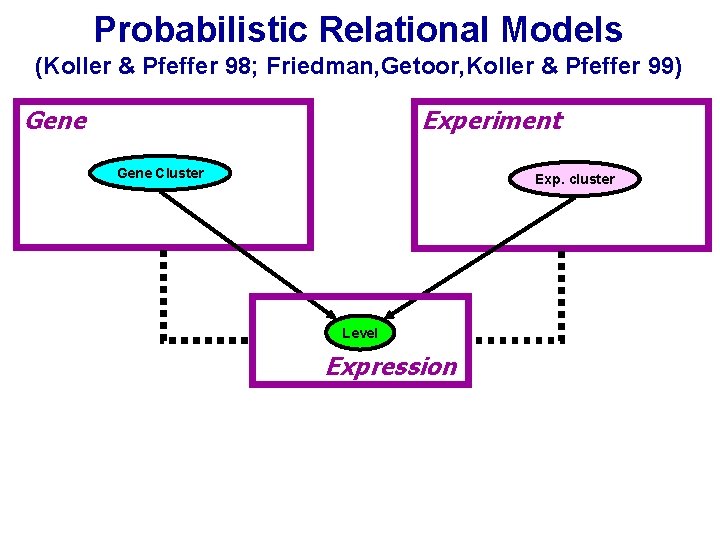
Probabilistic Relational Models (Koller & Pfeffer 98; Friedman, Getoor, Koller & Pfeffer 99) Gene Experiment Gene Cluster Exp. cluster Level Expression

Resulting Bayesian Network Gene Experiment Gene Cluster Exp. cluster Level Expression + Exp. Cluster 2 Exp. Cluster 1 Gene Cluster 1 Level 1, 2 Level 2, 1 Level 2, 2 Level 3, 1 Level 3, 2 Gene Cluster 3
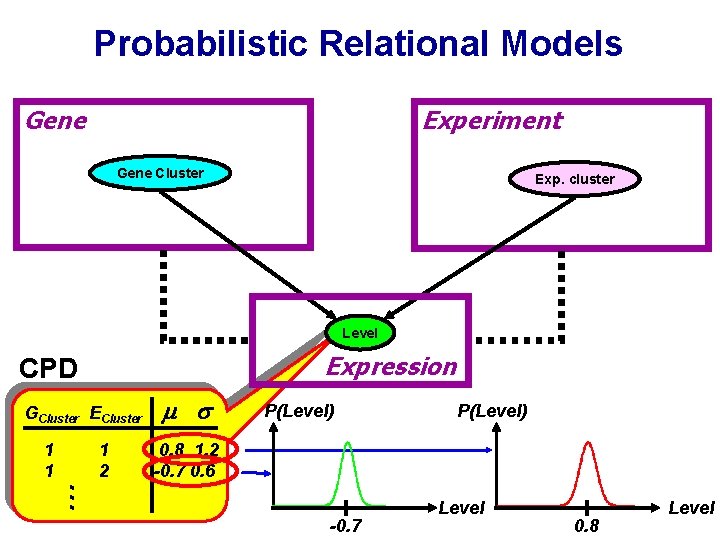
Probabilistic Relational Models Gene Experiment Gene Cluster Exp. cluster Level Expression CPD GCluster ECluster 1 2 P(Level) 0. 8 1. 2 -0. 7 0. 6 … 1 1 -0. 7 Level 0. 8 Level
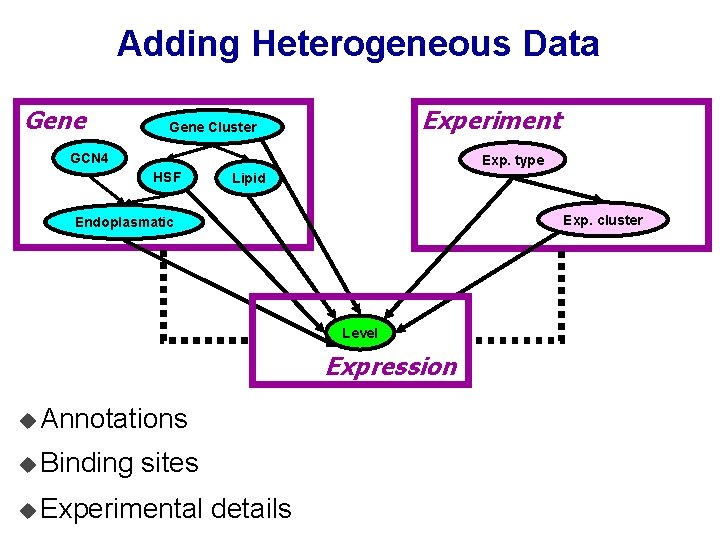
Adding Heterogeneous Data Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression u Annotations u Binding sites u Experimental details
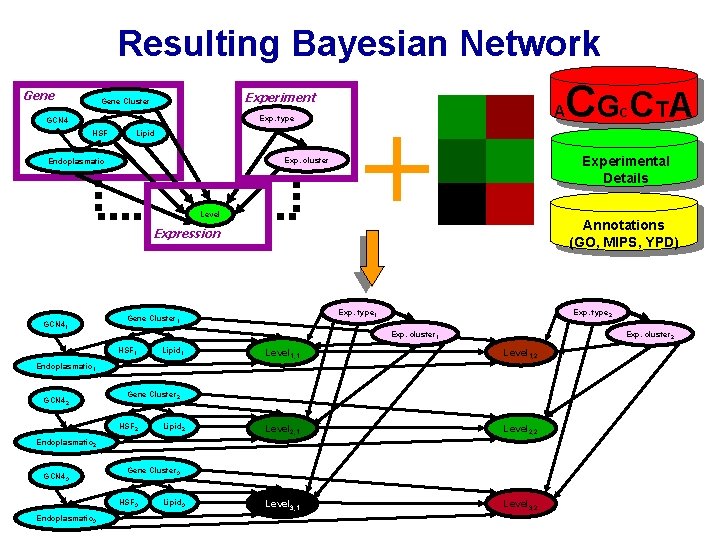
Resulting Bayesian Network Gene Experiment Gene Cluster Exp. type GCN 4 HSF Lipid Exp. cluster Endoplasmatic Level Expression GCN 41 + A Annotations (GO, MIPS, YPD) Exp. type 2 Exp. cluster 1 HSF 1 Lipid 1 Exp. cluster 2 Level 1, 1 Level 1, 2 Level 2, 1 Level 2, 2 Level 3, 1 Level 3, 2 Endoplasmatic 1 GCN 42 Gene Cluster 2 HSF 2 Lipid 2 Endoplasmatic 2 GCN 43 Gene Cluster 3 HSF 3 Endoplasmatic 3 Lipid 3 C Experimental Details Exp. type 1 Gene Cluster 1 CG CT A
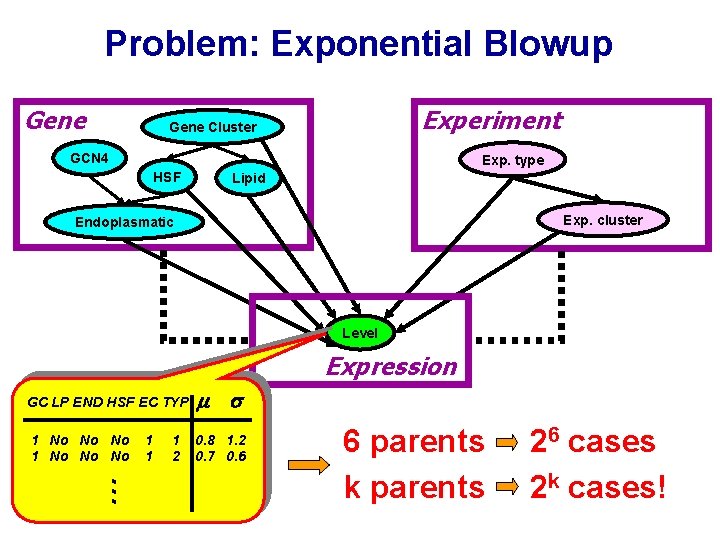
Problem: Exponential Blowup Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression 1 No No No 0. 8 1. 2 0. 7 0. 6 … GC LP END HSF EC TYP 1 1 1 2 6 parents k parents 26 cases 2 k cases!
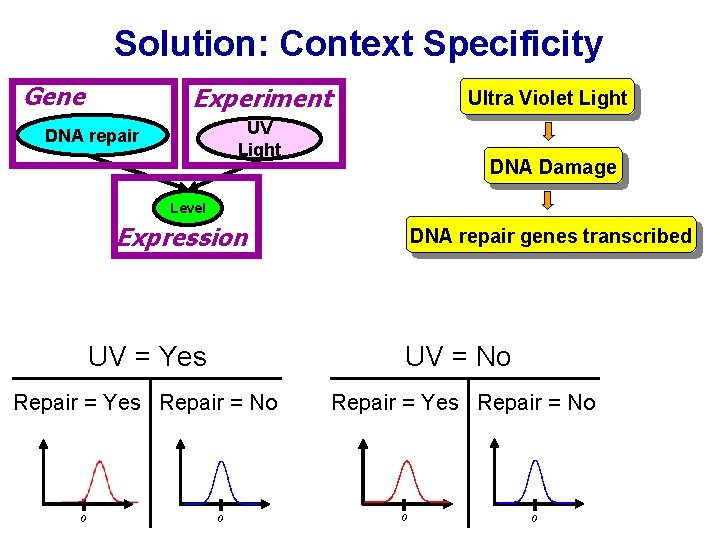
Solution: Context Specificity Gene Experiment Ultra Violet Light UV Light DNA repair DNA Damage Level Expression DNA repair genes transcribed UV = Yes UV = No Repair = Yes Repair = No 0 0
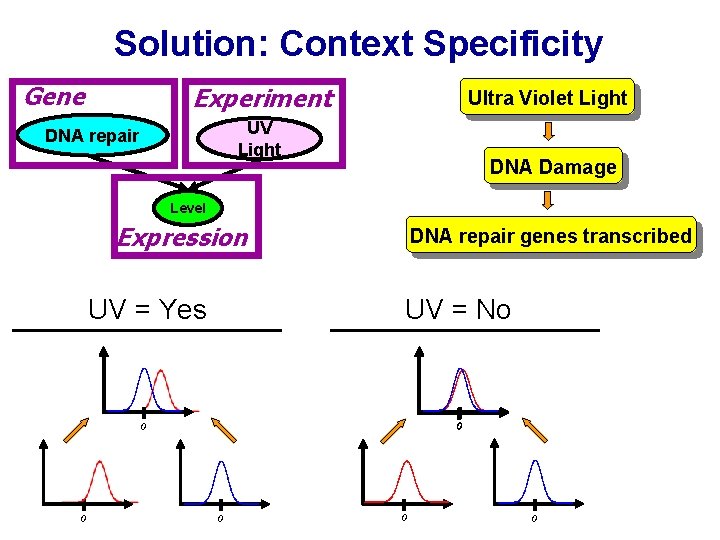
Solution: Context Specificity Gene Experiment Ultra Violet Light UV Light DNA repair DNA Damage Level Expression UV = Yes DNA repair genes transcribed UV = No 0 0 0
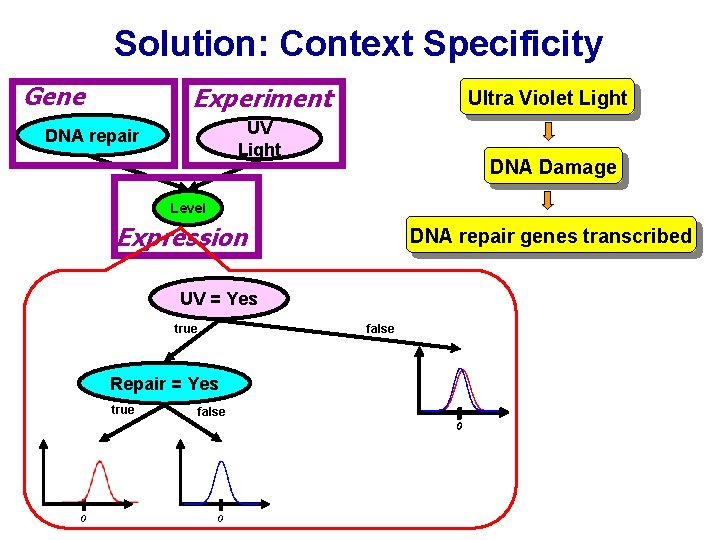
Solution: Context Specificity Gene Experiment Ultra Violet Light UV Light DNA repair DNA Damage Level Expression DNA repair genes transcribed UV = Yes true false Repair = Yes true false 0 0 0
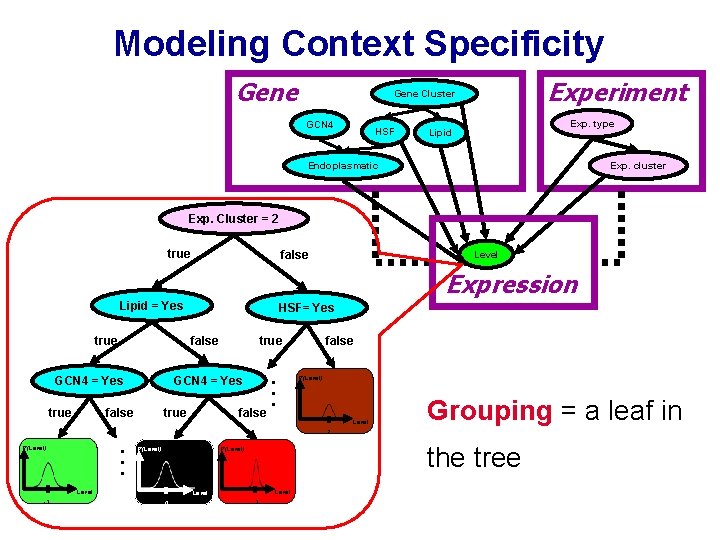
Modeling Context Specificity Gene Experiment Gene Cluster GCN 4 HSF Exp. type Lipid Endoplasmatic Exp. cluster Exp. Cluster = 2 true false Level Expression Lipid = Yes false GCN 4 = Yes true false true . . . true HSF= Yes false P(Level) false Level Grouping = a leaf in 2 . . . P(Level) Level -3 the tree P(Level) Level 0 3

How do I learn these models?
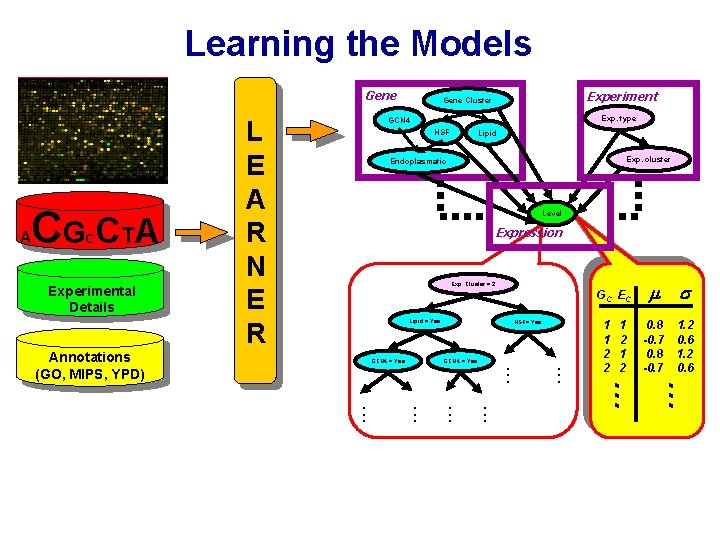
Learning the Models Gene Exp. cluster Endoplasmatic Level Expression G C EC GCN 4 = Yes . . . Annotations (GO, MIPS, YPD) HSF= Yes 1 1 2 2 1 2 … Lipid = Yes 0. 8 -0. 7 1. 2 0. 6 … Exp. Cluster = 2 . . . Experimental Details Lipid . . . C L E A R N E R HSF . . . CG CT A Exp. type GCN 4 . . . A Experiment Gene Cluster
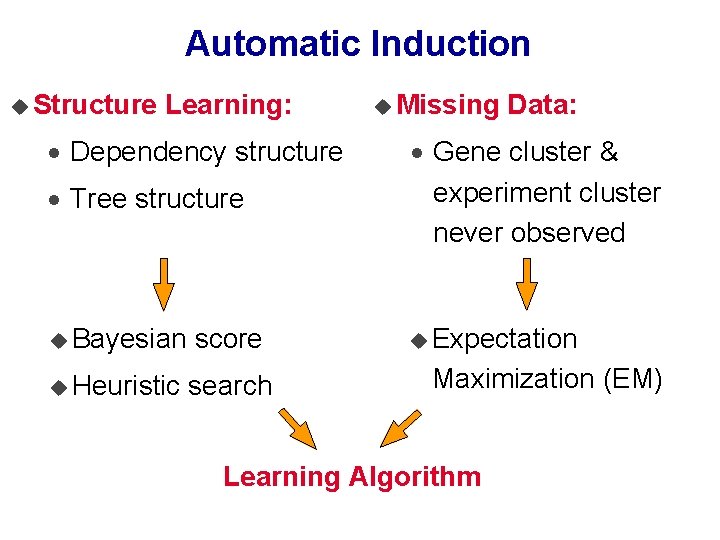
Automatic Induction u Structure Learning: · Dependency structure u Missing Data: · Tree structure · Gene cluster & experiment cluster never observed u Bayesian score u Expectation u Heuristic search Maximization (EM) Learning Algorithm
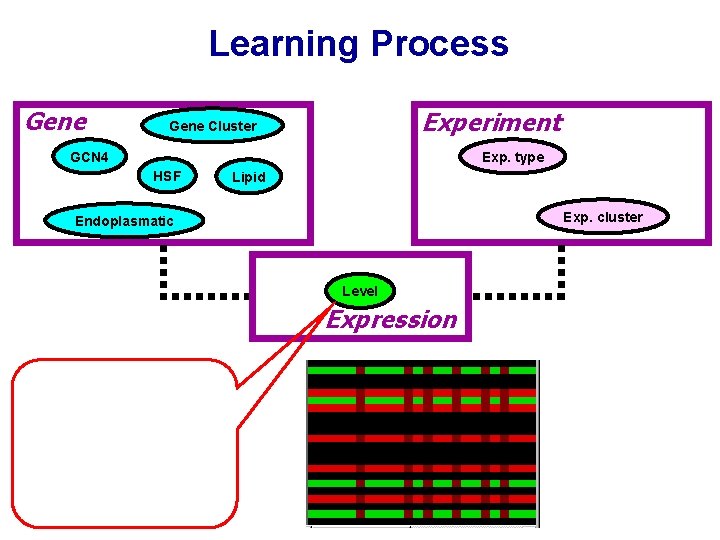
Learning Process Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression

Learning Process Experiment Similarity Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression Exp. Cluster = 2
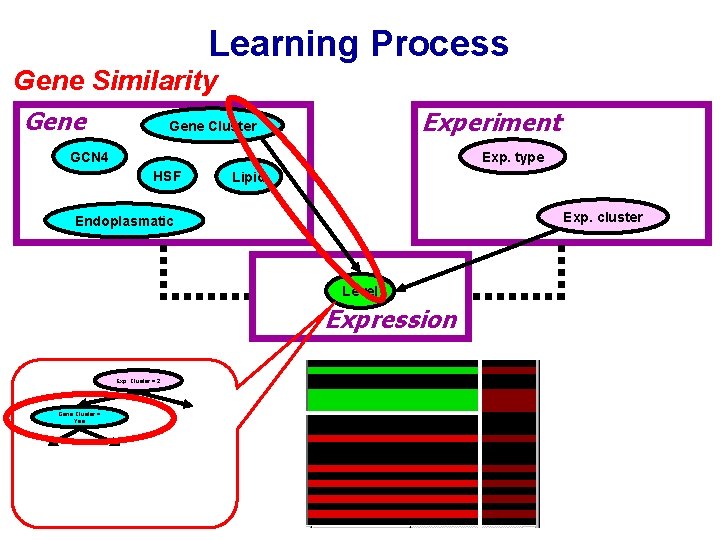
Learning Process Gene Similarity Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression Exp. Cluster = 2 Gene Cluster = Yes
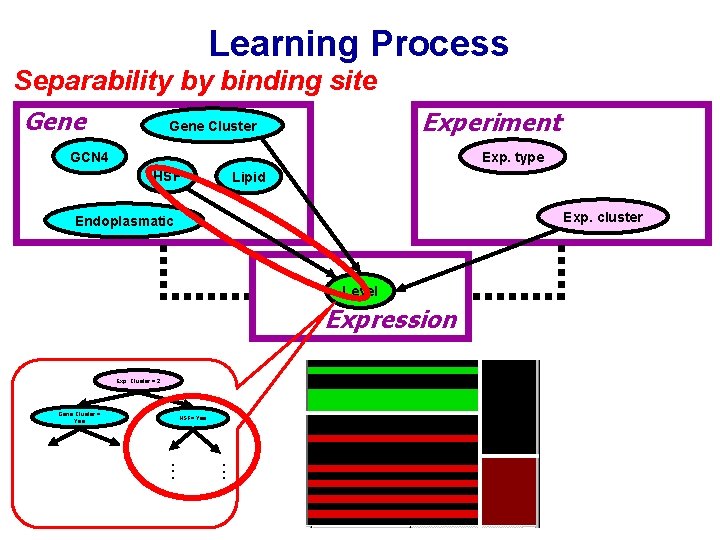
Learning Process Separability by binding site Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression Exp. Cluster = 2 Gene Cluster = Yes . . . HSF= Yes
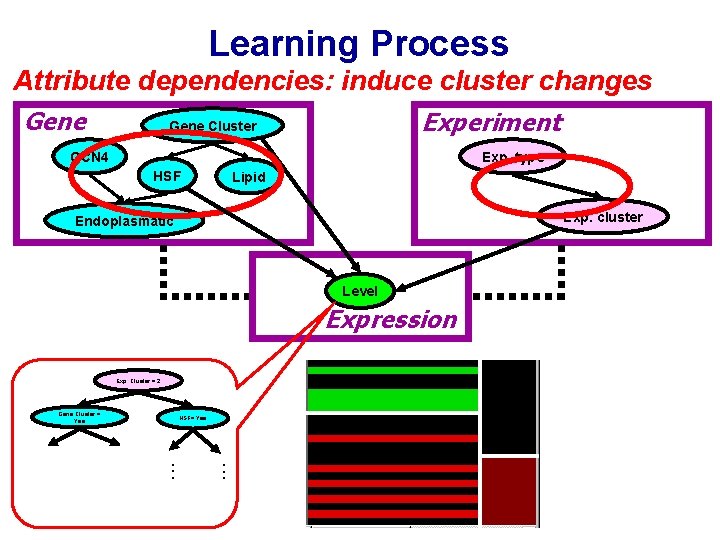
Learning Process Attribute dependencies: induce cluster changes Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression Exp. Cluster = 2 Gene Cluster = Yes . . . HSF= Yes
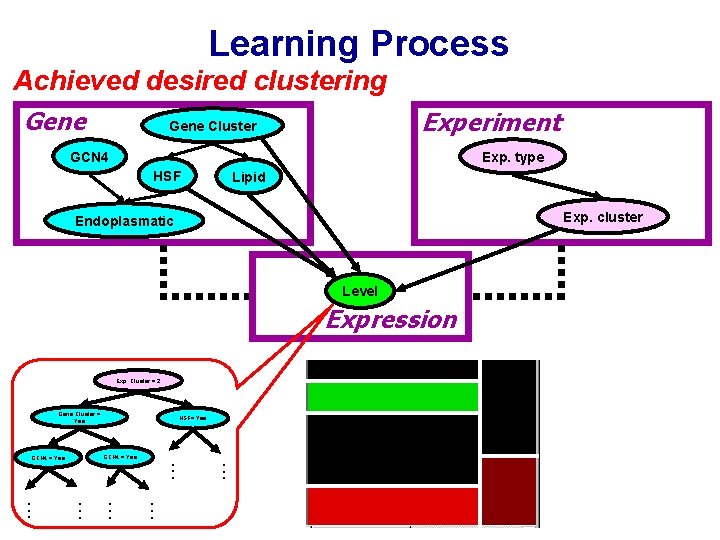
Learning Process Achieved desired clustering Gene Experiment Gene Cluster GCN 4 Exp. type HSF Lipid Exp. cluster Endoplasmatic Level Expression Exp. Cluster = 2 Gene Cluster = Yes HSF= Yes . . . . GCN 4 = Yes

Yeast Stress Data (Gasch et al 2001) u Measured u 92 response to stress cond. arrays u We selected ~900 genes u Added data: TRANSFAC, MIPS Results: u 15 u 7 significant TFs significant function categories u 793 Groupings

Context Specific Groupings u Down in nitrogen depletion u Transporter genes u Metabolism of amino acids

Context Specific Groupings u Up in Starvation, Nitrogen depletion & DTT u Transporter genes u Metabolism of nitrogen

Example Biological Finding u Discovered grouping of 17 genes · All induced in diauxic shift · All have 2 binding sites for MIG 1 transcription factor · Many not known to be regulated by MIG 1 u Context-sensitive groupings were key to finding cluster

Compendium Data (Hughes et al 2000) u 300 samples of yeast deletion mutants Gene Array/Mutated Gene GCluster GCN 4 GCluster (of mutated gene) HSF Endoplasmatic Lipid (of mutated gene) Lipid Level Expression ACluster

Resulting Bayesian Network Gene 1 mutant Gene 3 mutant Gene 1 Lipid 1 Gene Cluster 1 Lipid 3 Array. cluster 1 Array. cluster 3 HSF 1 Gene 2 Gene Cluster 2 Level 1, 1 Level 1, 2 Level 2, 1 Level 2, 2 Level 3, 1 Level 3, 2 HSF 2 Gene 3 Gene Cluster 3 HSF 3 Gene 4 Gene Cluster 4 HSF 4

Experimental Setup u Goal: predict the effect of mutating specific genes without performing the experiment (!) u Example: predicting the effect of mutating gene 4 Gene 4 mutant u Available information: · Attributes of gene 4 · Gene Cluster of gene 4 as a gene Lipid 4 ? Array. cluster ? Gene Cluster 4 HSF 4
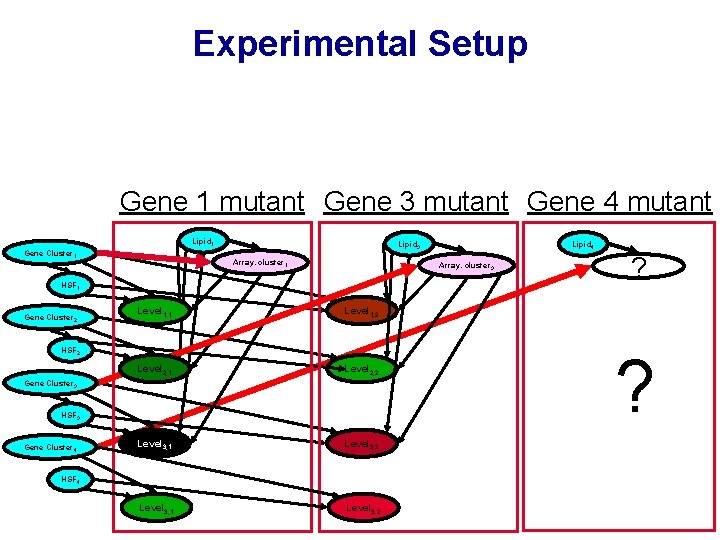
Experimental Setup Gene 1 mutant Gene 3 mutant Gene 4 mutant Lipid 1 Gene Cluster 1 Lipid 3 Array. cluster 1 Array. cluster 3 HSF 1 Gene Cluster 2 Level 1, 1 Level 1, 2 Level 2, 1 Level 2, 2 Level 3, 1 Level 3, 2 HSF 2 Gene Cluster 3 HSF 3 Gene Cluster 4 Lipid 4 HSF 4 ? Array. cluster ?
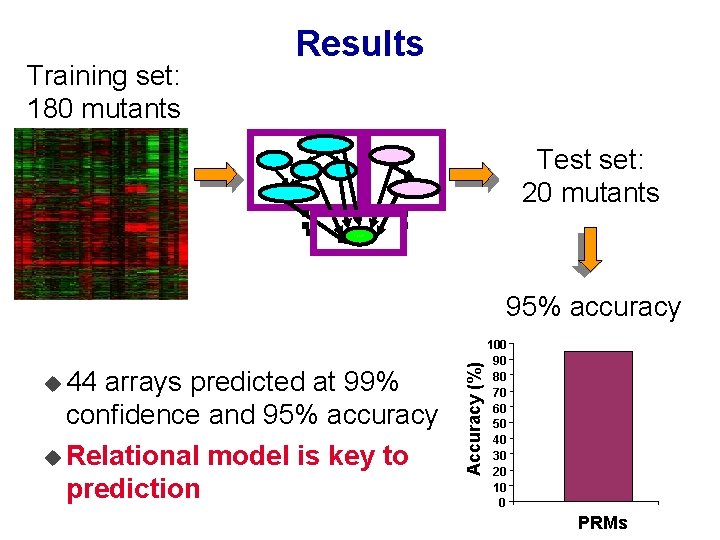
Results Training set: 180 mutants Gene Cluster Test set: 20 mutants Exp. type GCN 4 HSF Lipid Exp. cluster Endoplasmatic Level u 44 arrays predicted at 99% confidence and 95% accuracy u Relational model is key to prediction Accuracy (%) 95% accuracy 100 90 80 70 60 50 40 30 20 10 0 PRMs

Conclusions u Presented a unified probabilistic framework: · Models complex biological domains · Expressive data organization · Incorporates heterogeneous data u Future directions: · Incorporate DNA and protein sequence data · Discover regulatory networks Thank You! u Paper: http: //www. cs. stanford. edu/~eran u Software (soon): http: //dags. stanford. edu/bio u Contact: eran@cs. stanford. edu
- Slides: 37