Restriction Enzymes Restriction Enzymes scan the DNA sequence






















- Slides: 42

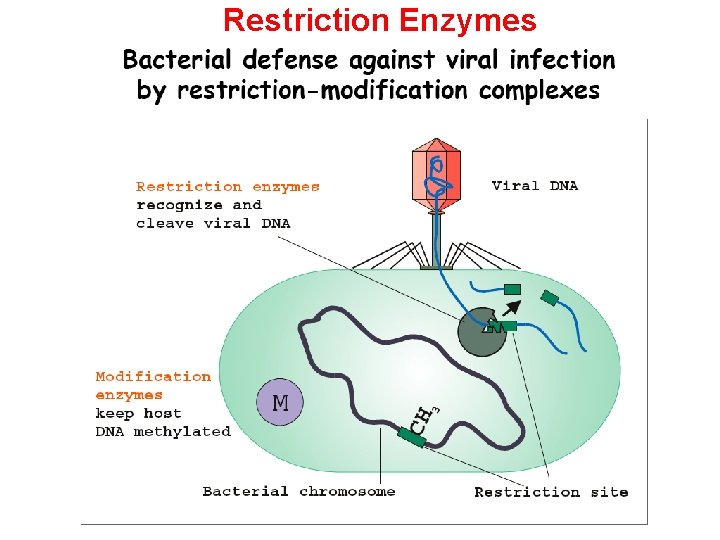
Restriction Enzymes

• Restriction Enzymes scan the DNA sequence • Find a very specific set of nucleotides • Make a specific cut

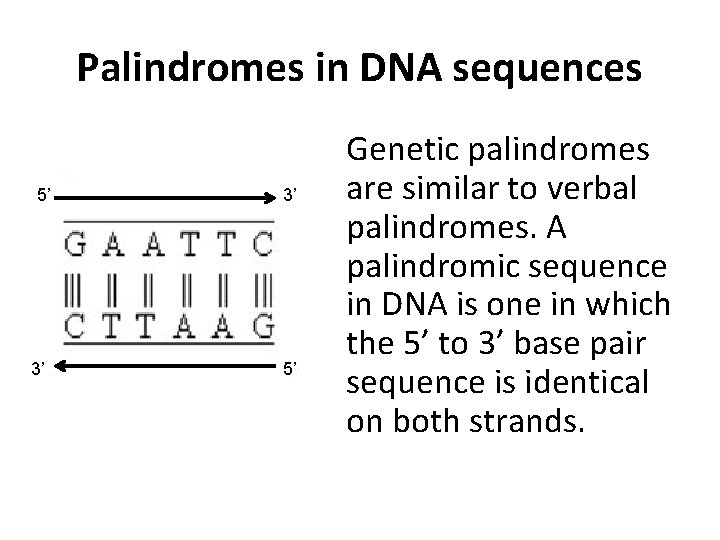
Palindromes in DNA sequences 5’ 3’ 3’ 5’ Genetic palindromes are similar to verbal palindromes. A palindromic sequence in DNA is one in which the 5’ to 3’ base pair sequence is identical on both strands.

Restriction enzymes recognize and make a cut within specific palindromic sequences, known as restriction sites, in the DNA. This is usually a 4 - or 6 base pair sequence.

Restriction Endonuclease Types Type I- multi-subunit, both endonuclease and methylase activities, cleave at random up to 1000 bp from recognition sequence Type II- most single subunit, cleave DNA within recognition sequence Type III- multi-subunit, endonuclease and methylase about 25 bp from recognition sequence

Hae III Hae. III is a restriction enzyme that searches the DNA molecule until it finds this sequence of four nitrogen bases. 5’ TGACGGGTTCGAGGCCAG 3’ 3’ ACTGCCCAAGGTCCGGTC 5’

Once the recognition site is found Hae III will cleave the DNA at that site 5’ TGACGGGTTCGAGGCCAG 3’ 3’ ACTGCCCAAGGTCCGGTC 5’

These cuts produce “blunt ends” 5’ TGACGGGTTCGAGG 3’ ACTGCCCAAGGTCC CCAG 3’ GGTC 5’

The names for restriction enzymes come from: • the type of bacteria in which the enzyme is found • the order in which the restriction enzyme was identified and isolated. Eco. RI for example R strain of E. coli bacteria I as it is was the first E. coli restriction enzyme to be discovered.

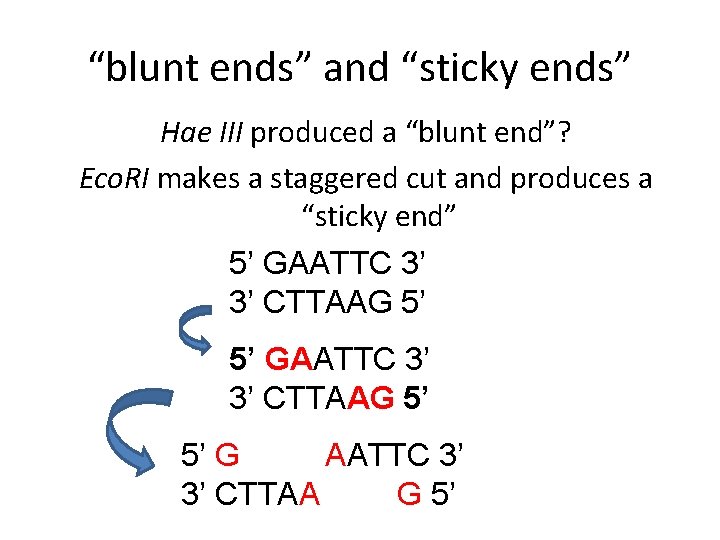
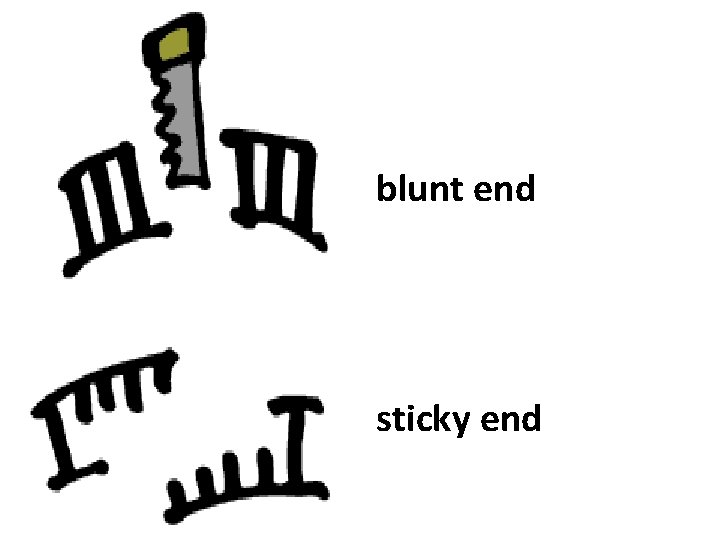
“blunt ends” and “sticky ends” Hae III produced a “blunt end”? Eco. RI makes a staggered cut and produces a “sticky end” 5’ GAATTC 3’ 3’ CTTAAG 5’ 5’ G AATTC 3’ 3’ CTTAA G 5’

blunt end sticky end

More examples of restriction sites of restriction enzymes with their cut sites Hind III: 5’ AAGCTT 3’ 3’ TTCGAA 5’ Bam HI: 5’ GGATCC 3’ 3’ CCTAGG 5’ Alu I: 5’ AGCT 3’ 3’ TCGA 5’

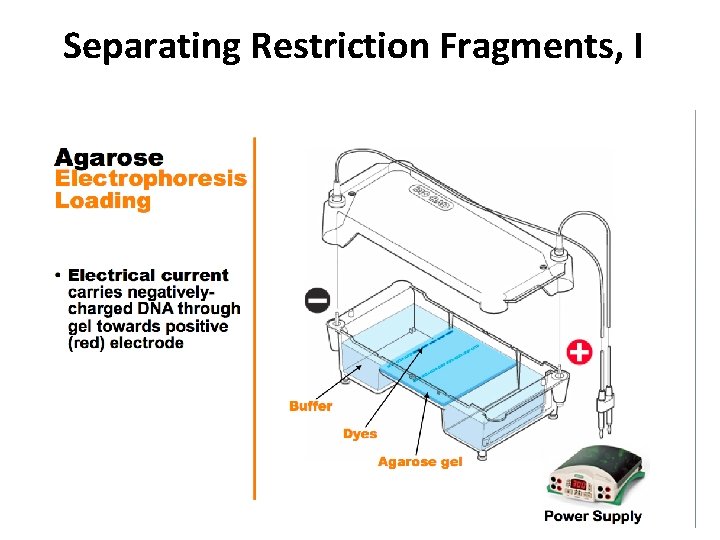
Separating Restriction Fragments, I

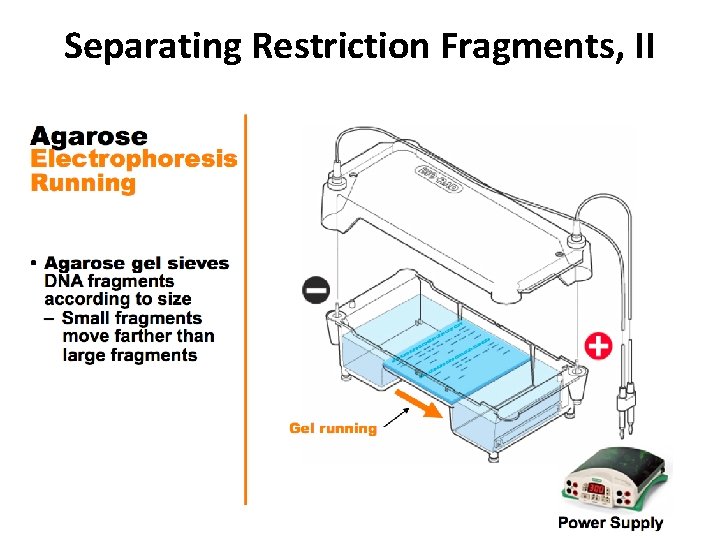
Separating Restriction Fragments, II

Gene Cloning • What is gene cloning? How does it differ from cloning an entire organism? • Why is gene cloning done? • How is gene cloning accomplished ? • What is a DNA ‘Library’?

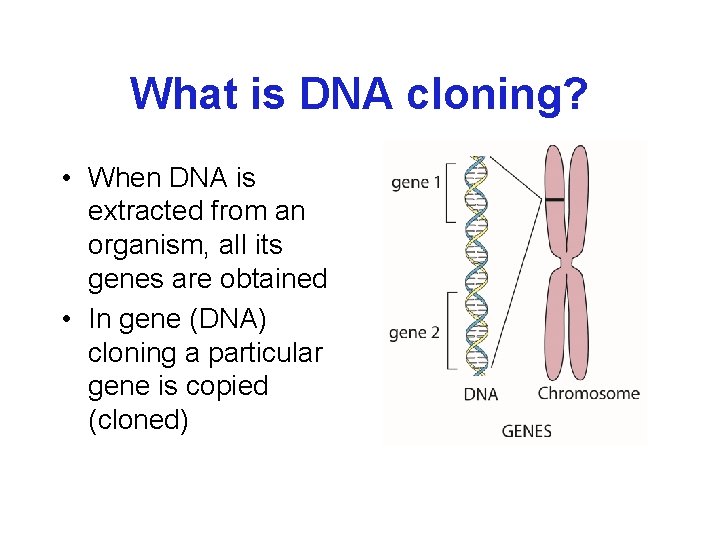
What is DNA cloning? • When DNA is extracted from an organism, all its genes are obtained • In gene (DNA) cloning a particular gene is copied (cloned)

Why Clone DNA? • A particular gene can be isolated and its nucleotide sequence determined • Control sequences of DNA can be identified & analyzed • Protein/enzyme/RNA function can be investigated • Mutations can be identified, e. g. gene defects related to specific diseases • Organisms can be ‘engineered’ for specific purposes, e. g. insulin production, insect resistance, etc.

How is DNA cloned? , I Blood sample • DNA is extractedhere from blood • Restriction enzymes, e. g. Eco. R I, Hind III, etc. , cut the DNA into small pieces • Different DNA pieces cut with the same enzyme can join, or recombine. DNA Restriction enzymes

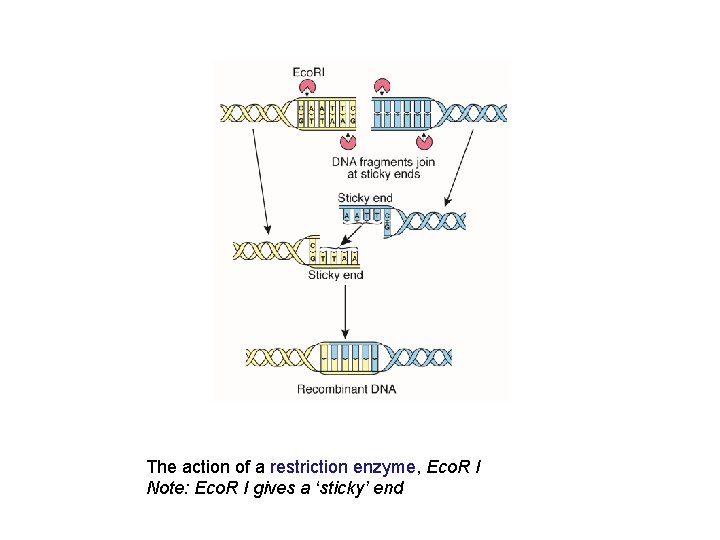
The action of a restriction enzyme, Eco. R I Note: Eco. R I gives a ‘sticky’ end

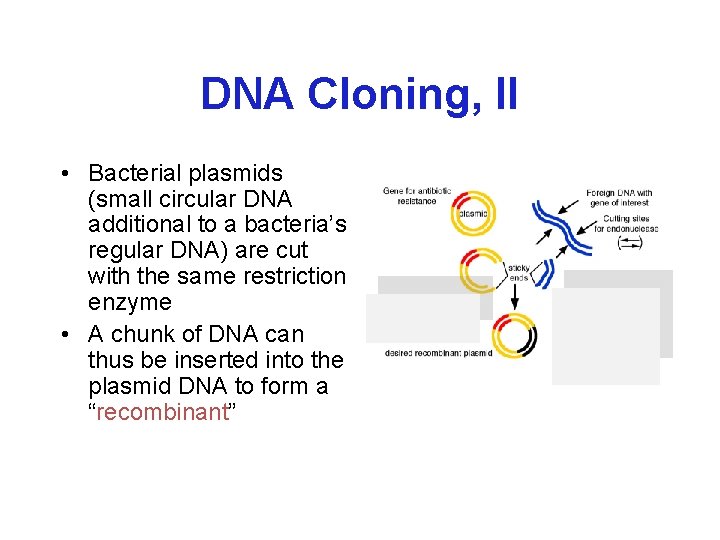
DNA Cloning, II • Bacterial plasmids (small circular DNA additional to a bacteria’s regular DNA) are cut with the same restriction enzyme • A chunk of DNA can thus be inserted into the plasmid DNA to form a “recombinant”

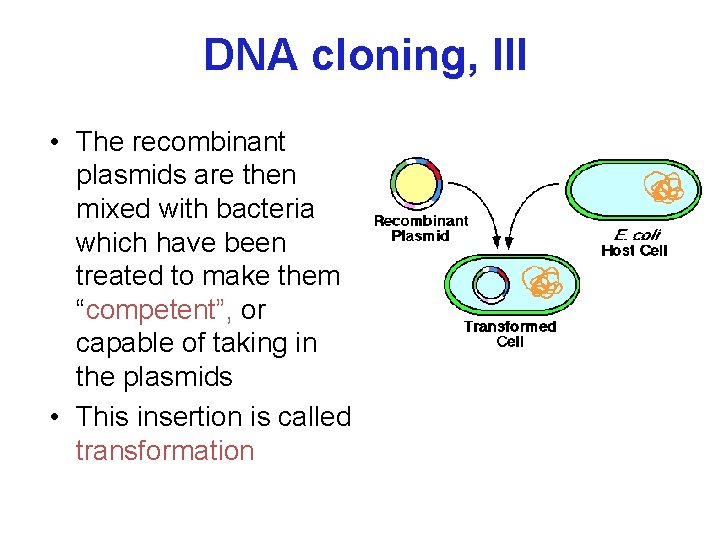
DNA cloning, III • The recombinant plasmids are then mixed with bacteria which have been treated to make them “competent”, or capable of taking in the plasmids • This insertion is called transformation

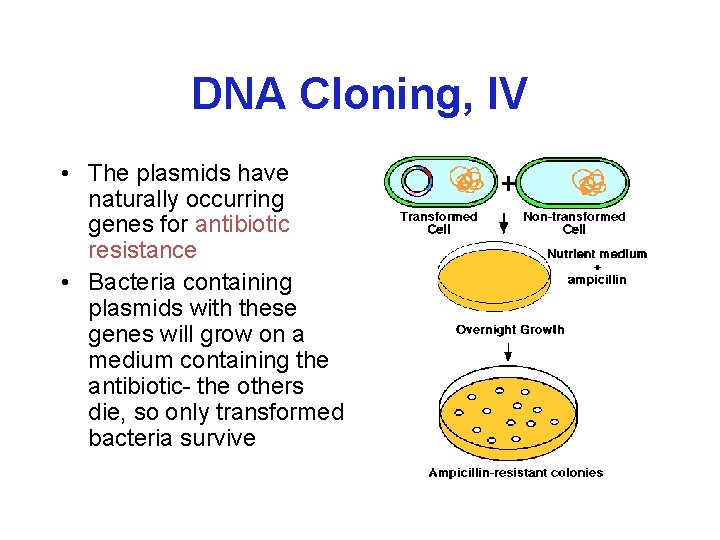
DNA Cloning, IV • The plasmids have naturally occurring genes for antibiotic resistance • Bacteria containing plasmids with these genes will grow on a medium containing the antibiotic- the others die, so only transformed bacteria survive

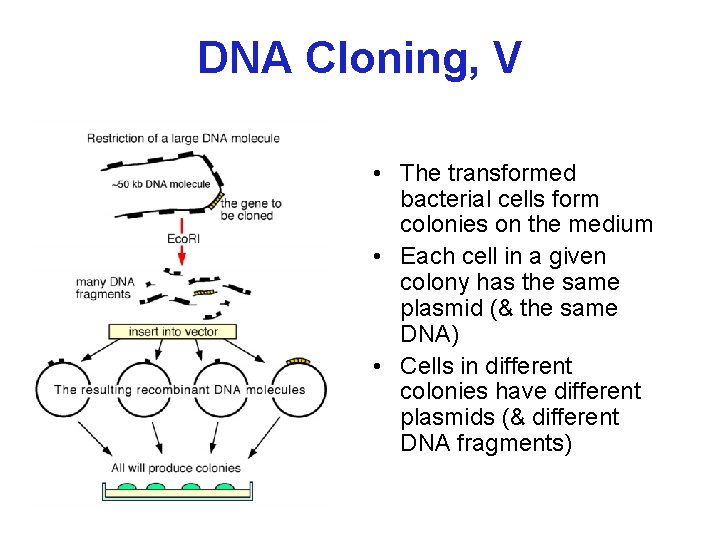
DNA Cloning, V • The transformed bacterial cells form colonies on the medium • Each cell in a given colony has the same plasmid (& the same DNA) • Cells in different colonies have different plasmids (& different DNA fragments)

Screening, I Screening can involve: 1. Phenotypic screeningthe protein encoded by the gene changes the color of the colony 2. Using antibodies that recognize the protein produced by a particular gene

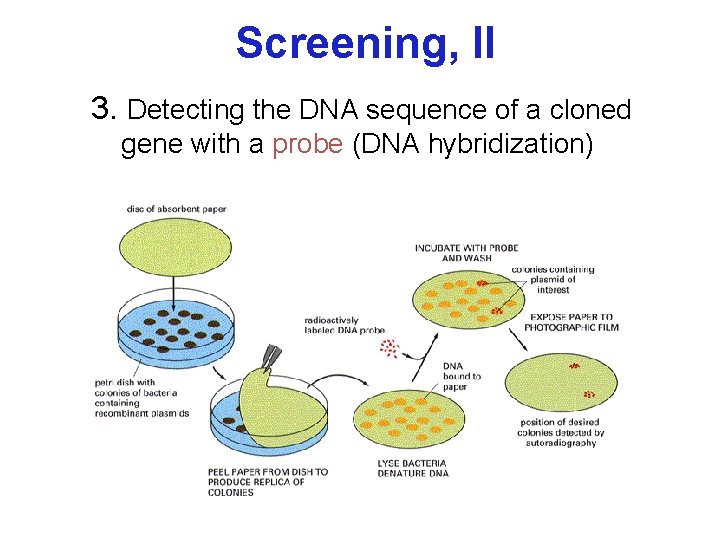
Screening, II 3. Detecting the DNA sequence of a cloned gene with a probe (DNA hybridization)

Polymerase Chain Reaction PCR

PCR • invented by Karry B. Mullis (1983, Nobel Prize 1993) • patent sold by Cetus corp. to La Roche for $300 million • depends on thermoresistant DNA polymerase (e. g. Taq polymerase) and a thermal cycler

Heat-stable DNA polymerase • Taq DNA polymerase was isolated from the bacterium Thermus aquaticus. • Taq polymerase is stable at the high temperatures (~95 o. C) used for denaturing DNA. Hot springs at Yellowstone National Park, Wyoming.

DNA polymerase requirements • • template primer nucleotides regulated p. H, salt concentration, cofactors

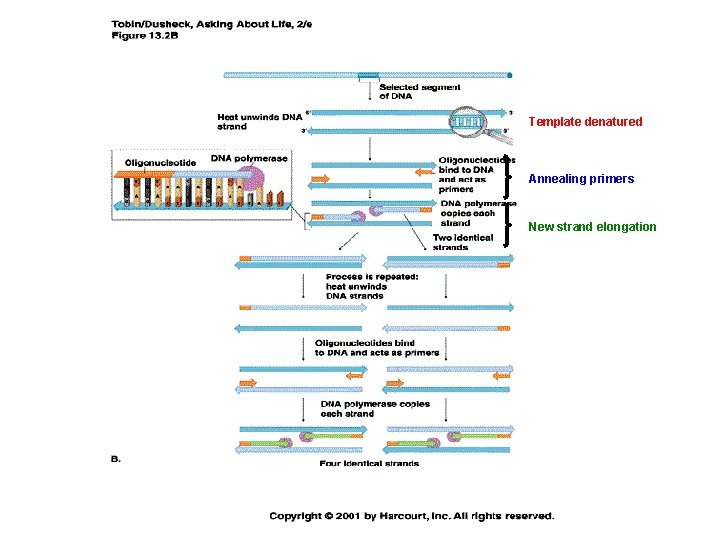
Steps in DNA replication 1) template denatured 2) primers anneal 3) new strand elongation

Steps in a PCR cycle 1) template denatured: 94 C, 30 sec 2) primers anneal 45 -72 C, depending on primer sequence 30 sec – 1 min 3) new strand elongation 72 C depending on the type of polymerase 1 min for 1000 nucleotides of amplified sequence Number of specific DNA molecule copies grows exponentially with each PCR cycle. Usually run 20 -40 cycles to get enough DNA for most applications (If you start with 2 molecules, after 30 cycles you will have more than a billion)

PCR Process • 25 -30 cycles • 2 minute cycles • DNA thermal cycler

Template denatured Annealing primers New strand elongation

Uses for PCR • Research – Gene cloning – Real-time PCR – DNA sequencing • Clinical – DNA fingerprinting • Crime scene analysis • Paternity testing • Archeological finds – Genetically inherited diseases

DNA Sequencing

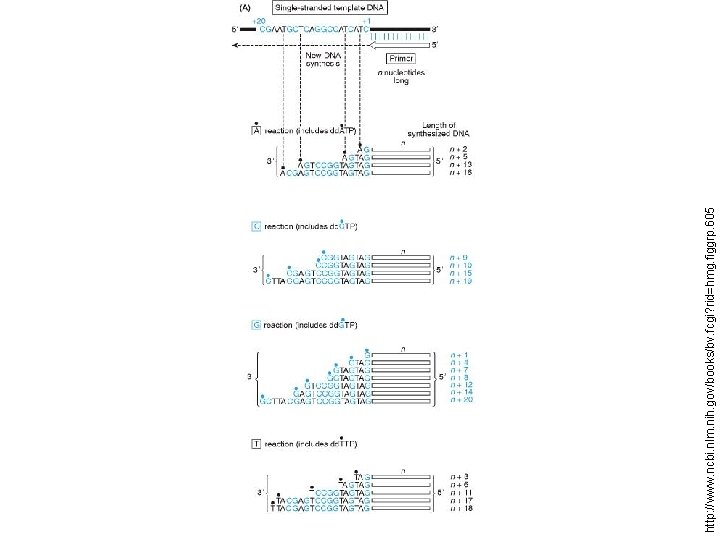
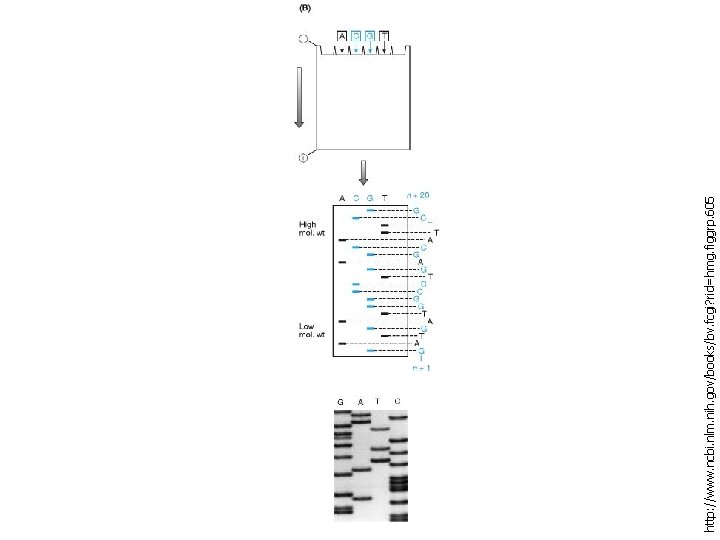
Chain termination method (Sanger Method), sequence of single stranded DNA is determined by enzymatic synthesis of complementary strands which terminate at specific nucleotide positions Chemical degradation method (Maxam-Gilbert Method), sequence of a double stranded DNA molecule is determined by chemical treatment that cuts at specific nucleotide positions

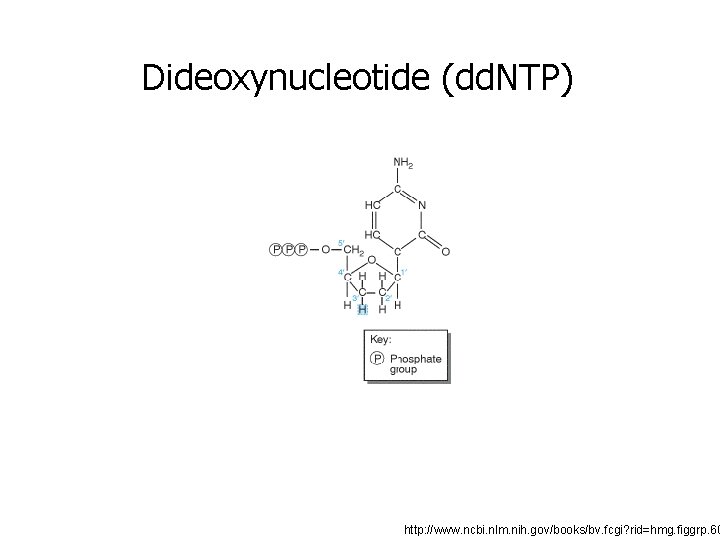
Dideoxynucleotide (dd. NTP) http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hmg. figgrp. 60

http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hmg. figgrp. 605

http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hmg. figgrp. 605

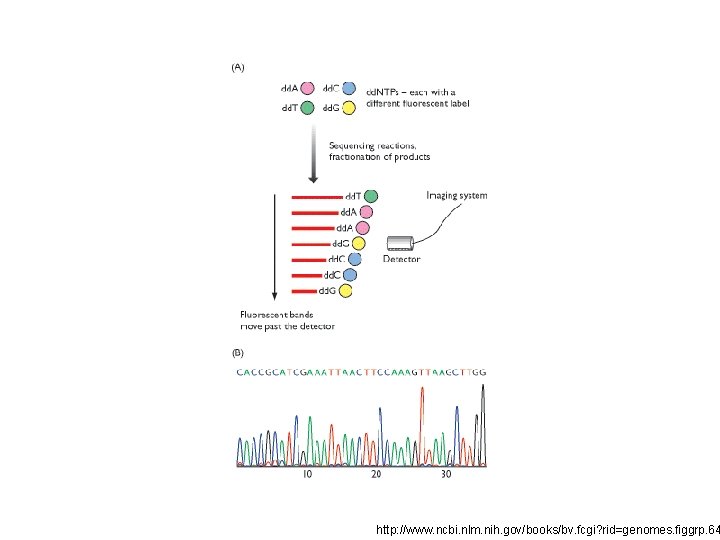
http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=genomes. figgrp. 64

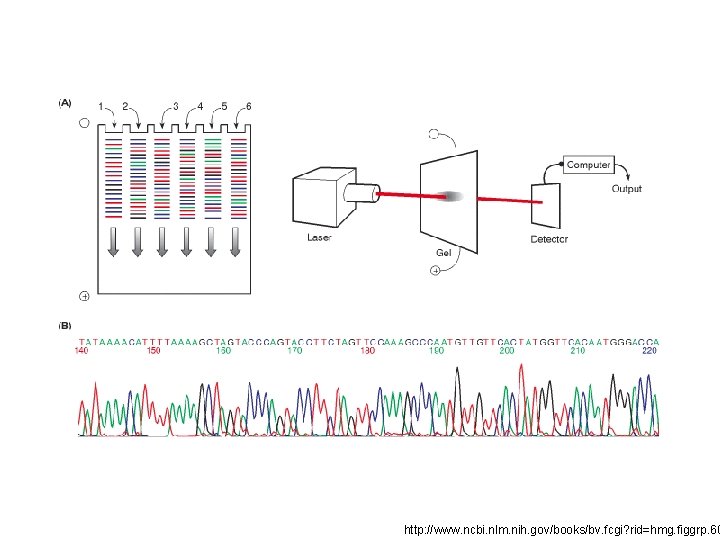
http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hmg. figgrp. 60

Costs and time for sequencing a human genome (3. 2 billion bp)