Restriction Enzymes WHAT ARE RESTRICTION ENZYMES Restriction Enzymes















- Slides: 18

Restriction Enzymes

WHAT ARE RESTRICTION ENZYMES? • Restriction Enzymes: • are molecular scissors, or BETTER YET, molecular scalpels, that are able to cut DNA • are also called “restriction endonucleases”

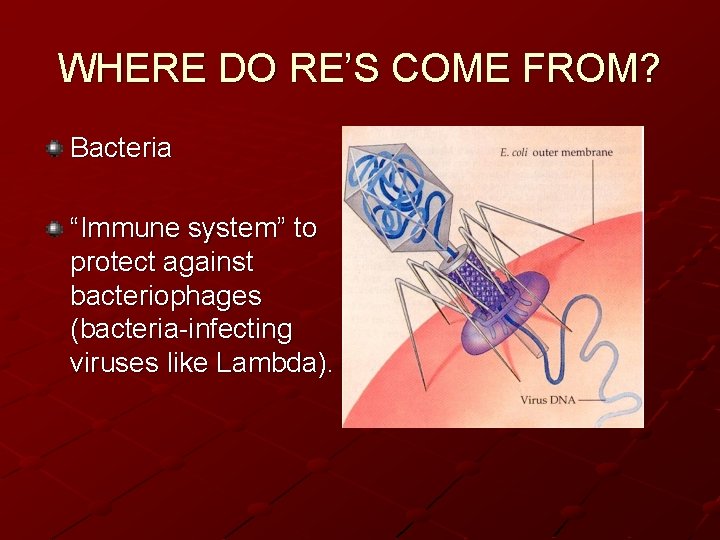
WHERE DO RE’S COME FROM? Bacteria “Immune system” to protect against bacteriophages (bacteria-infecting viruses like Lambda).

THOUGHT QUESTION Bacteria are prokaryotes. Prokaryotes do not have a nucleus. Both DNA and RE’s are in cytoplasm. Why isn’t bacterial DNA cut by RE’s?

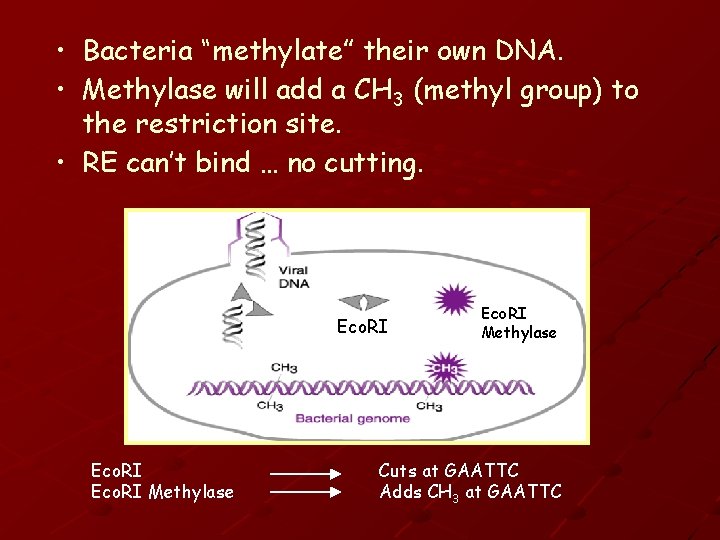
• Bacteria “methylate” their own DNA. • Methylase will add a CH 3 (methyl group) to the restriction site. • RE can’t bind … no cutting. Eco. RI Methylase Cuts at GAATTC Adds CH 3 at GAATTC

HOW DO RESTRICTION ENZYMES WORK? Usually cut DNA at a “palindrome” such as GAATTC. Palindrome – word or phrase when spelled backwords, spells the same word or phrase Ex. BOB MADAM I’M ADAM A Toyota! Race fast, safe car. A Toyota 5’ 3’ GAATTC | | | 3’ CTTAAG 5’ “Restriction site” or “Recognition Sequence”

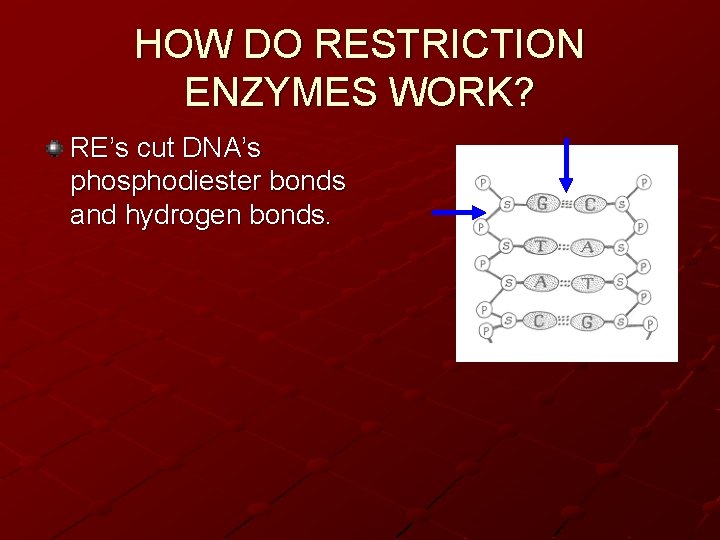
HOW DO RESTRICTION ENZYMES WORK? RE’s cut DNA’s phosphodiester bonds and hydrogen bonds.

HOW DO RESTRICTION ENZYMES WORK? • RE’s generate two different types of “cuts” • Blunt ends • Cut straight through the middle of the palindrome • Sticky ends • Uneven cut through the DNA leaves “overhang” of unattached bases

HOW DO RESTRICTION ENZYMES WORK? • Blunt vs. Sticky…which is more useful to genetic engineers? • RECOMBINANT DNA = DNA from 2+ sources

HOW IS DNA PASTED TOGETHER? Ligase – another enzyme which reconnects phosphodiester bonds.

In short… Enzymes BUILD or BREAK chemical bonds…

Enzymes - a brief reminder • Enzymes: • • Are proteins Usually end with “-ase” Speed up chemical reactions (catalysts) Have specific shapes that fit specific substrates Lower the activation energy of a reaction Can be used over, and over again Often require “cofactors” to work properly • metal ions - ex. iron, zinc, copper • organic molecules (COENZYMES) generally derived from vitamins

WHAT ARE RE’S USED FOR? • Genetic engineering – pasting together DNA from two different organisms. • Spider silk gene in goats • Glo-fish • GFP addition (to tag and track gene expression) • Golden Rice • Human Insulin

WHAT ELSE ARE RE’S USED FOR? Forensics – DNA Fingerprinting for crime scene investigation and paternity testing. Everyone’s DNA has a different sequence – even though only 0. 1% different. How frequently would Eco. RI cut DNA? 6 4 = once every 4096 bp Lambda (48, 514 bp) would expect about 12 Eco. RI sites

HOW ARE RE’S NAMED? • After bacteria which produces them. • Genus Recognition Site Eco. RI Escherichia • Species coli • Strain RY 13 • 1 st letter • 1 st two letters • 1 st letter • Order Isolated I G^AATTC

HOW ARE RE’S NAMED? After bacteria which produces them. Eco. RI Hind. III Genus Escherichia Haemophilus Bacillus Species coli influenzae amylo. Strain R d H Order Isolated I III I Recognition Site G^AATTC A^AGCTT Bam. HI G^GATGC

Quick review… • • • • 1. Monomer of DNA = 2. Three parts of the monomer = 3. A - T with ___ hydrogen bond 4. Which is bigger -- purines or pyrmidines? 5. What two bases are pyrimidines? 6. Charge on DNA = 7. DNA has a ____-wise and ____-hand twist. 8. Bond between sugars - phosphates = 9. Restriction enzymes are found only in… 10. Same forwards and backwards = 11. Bacteria add ____groups to their DNA to prevent cutting. 12. Overhangs of unpaired bases = 13. In Hind. III, the “in” comes from the _____ of bacteria. 14. Enzymes usually end in _____.

• 1. Monomer of DNA = Nucleotide • 2. Three parts of the monomer = Sugar (deoxyribose), phosphate, base • 3. A - T with 2 hydrogen bonds • 4. Which is bigger -- purines or pyrmidines? Purines • 5. What two bases are pyrimidines? Cytosine and thymine • 6. Charge on DNA = negative • 7. DNA has a Clock -wise and right -hand twist. • 8. Bond between sugars - phosphates = phosphodiester • 9. Restriction enzymes are found only in… bacteria • 10. Same forwards and backwards = palindrome • 11. Bacteria add CH 3 (methyl) groups to their DNA to prevent cutting. • 12. Overhangs of unpaired bases = sticky ends • 13. In Hind. III, the “in” comes from the species of bacteria. • 14. Enzymes usually end in -ase.