Restriction Enzymes KAMAL KUMAR GUPTA Restriction Enzymes Nobel

Restriction Enzymes KAMAL KUMAR GUPTA

Restriction Enzymes Nobel prize in Physiology and Medicine (1978) Discovery of Restriction enzymes and their application in molecular genetics l Werner Arber l Deniel Nathans l Hamilton Smith

Restriction Enzymes 1962: “molecular scissors” discovered in bacteria An enzymatic immune system that recognizes and destroys foreign DNA

Restriction Enzymes Also called restriction endonclease 1962: “molecular scissors” discovered bacteria An enzymatic immune system recognizes and destroys foreign DNA in that

Restriction Endonucleases Restriction : restrict viruses Viral genome is destroyed upon entry Endonuclease Endo (inside), nuclease (cuts nucleic acid)

Restriction Endonucleases Why don’t bacteria destroy their own DNA with their restriction enzymes?

Restriction endonuclease recognizes a short and specific DNA sequence and cuts the DNA from inside. The specific DNA sequence is called restriction site (recognition sequence)

R-M System Restriction-modification (R-M) system Endonuclease activity: cuts foreign DNA Methyltransferase activity: protects host DNA from cleavage by the restriction enzyme. Methylate one of the bases in each strand Restriction enzyme and its cognate modification system constitute the R-M system
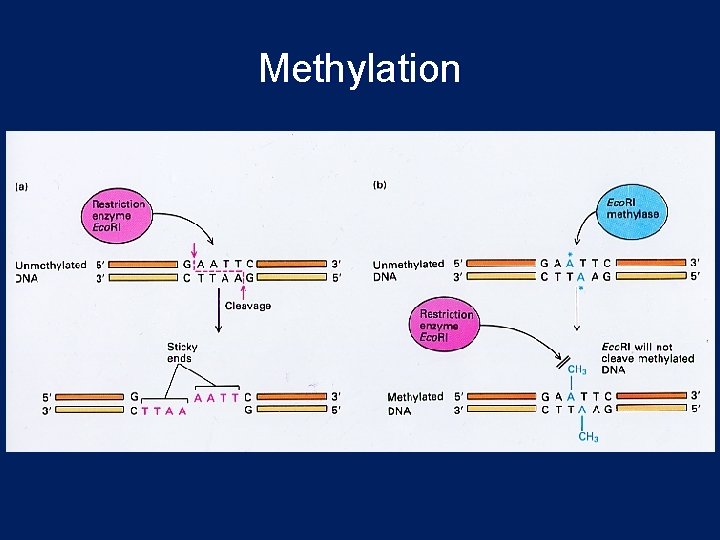
Methylation

Restriction Endonucleases Named for bacterial genus, species, strain, and type Example: Eco. R 1 Genus: Escherichia Species: coli Strain: R Order discovered: 1
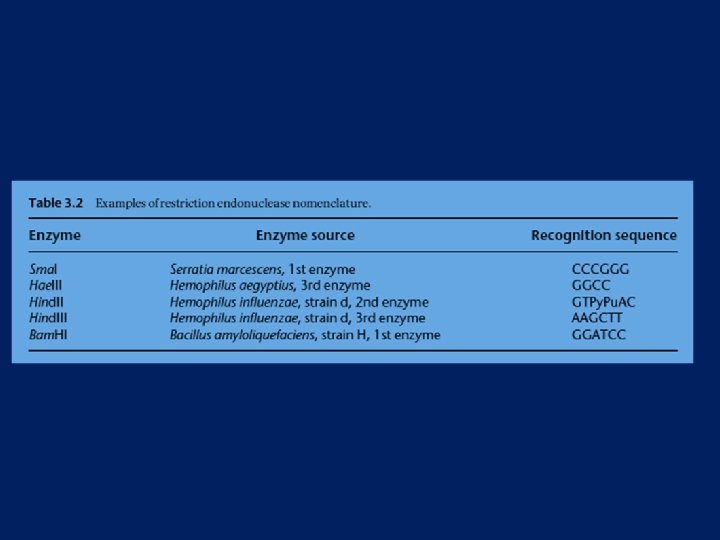

Categories of restriction enzymes NON SPECIFIC

Type II Two different enzymes which both recognize the same target sequence, which is symmetrical. The two enzymes either cleave or modify the recognition sequence

l l l Restriction and modification are mediated by separate enzymes so it is possible to cleave DNA in the absence of modification. Restriction activities do not require cofactors such as ATP or S-adenosylmethionine, making them easier to use. Recognize a defined, usually symmetrical, sequence and cut within it. Many of them also make a staggered break in the DNA.
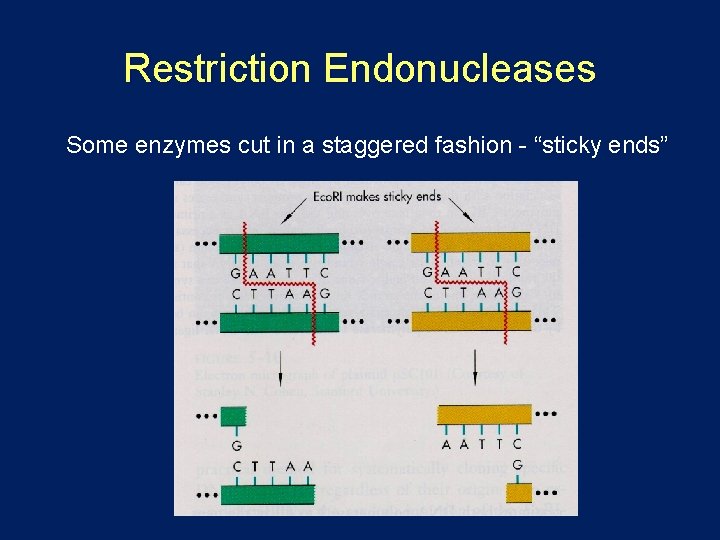
Restriction Endonucleases Some enzymes cut in a staggered fashion - “sticky ends”

Restriction Endonucleases Recognition sites have symmetry (palindromic) “Able was I, ere, I saw Elba” 5’-GGATCC-3’ Bam H 1 site: 3’-CCTAGG-5’

TYPES OF CUTS l BLUNT l STAGGERED

Restriction Endonucleases Enzymes recognize specific 4 -8 bp sequences Some enzymes cut in a staggered fashion - “sticky ends” Eco. RI 5’…GAATTC… 3’ 3’…CTTAAG… 5’
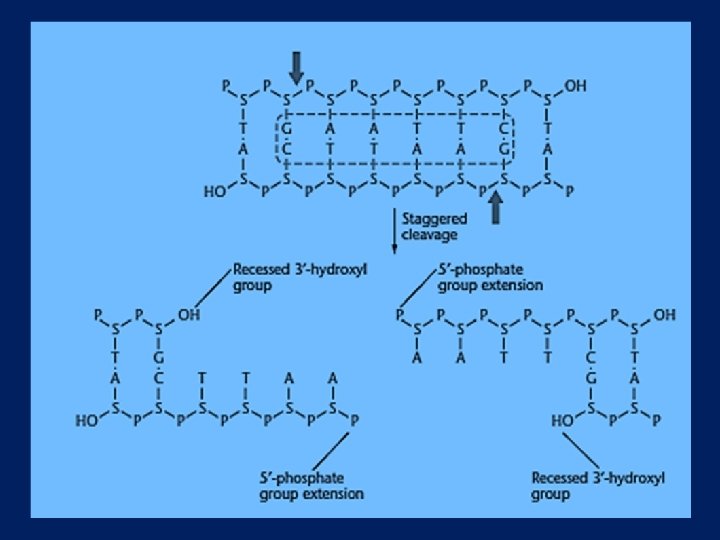

Restriction Endonucleases Some enzymes cut in a staggered fashion - “sticky ends” Pst. I 5’…CTGCAG… 3’ 3’…GACGTC… 5’ 5’…CTGCA-OH… 3’ 3’…G-P… 5’ 5’… P-G… 3’ 3’…OH-ACGTC… 5

Restriction Endonucleases Some enzymes cut in a direct fashion – “blunt ends” Pvu. II 5’…CAGCTG… 3’ 3’…GTCGAC… 5’
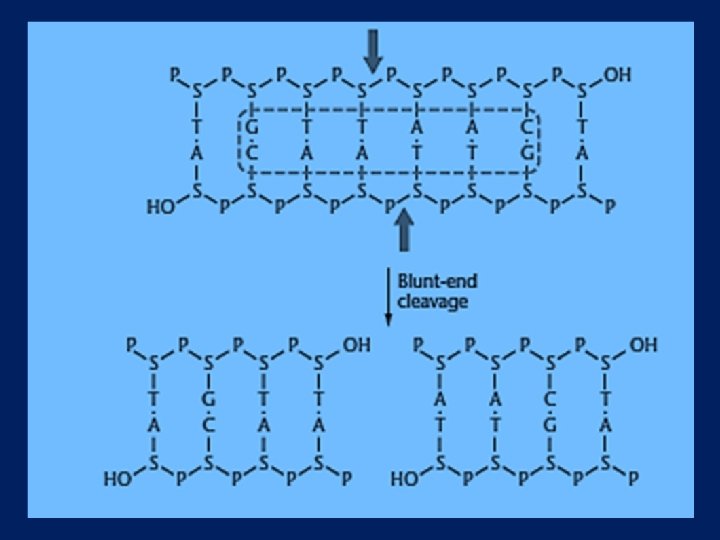
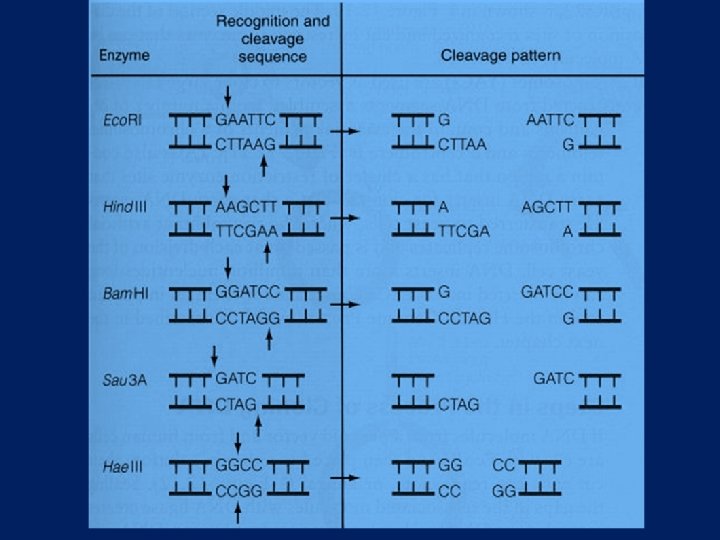

In a DNA molecule the fraction of Adenine is 20%. Calculate the number of restriction sites for Eco. R 1 in the DNA of size 500 kb

Few Restriction Enzymes Target sequence (cut at /) 5' -->3' Enzyme l l l l Bam HI Eco RI Hind III Mbo I Pst I Sma I Taq I Xma I l l l l G/ G A T C C G/ A A T T C A/ A G C T T /G A T C CTGCA/G CCC/GGG T/CGA C/CCGGG



Restriction endonucleases sequence are isoschizomers that recognize the same Isoschizomers are pairs of restriction enzymes specific to the same recognition sequence. For example, Sphl (CGTAC/G) and Bbu. I (CGTAC/G) are isoschizomers of each other. An enzyme that recognizes the same sequence but cuts it differently is a neoschizomer. Neoschizomers are a specific type (subset) of isoschizomer. For example, Sma. I (CCC/GGG) and Xma. I (C/CCGGG) are neoschizomers of each other.
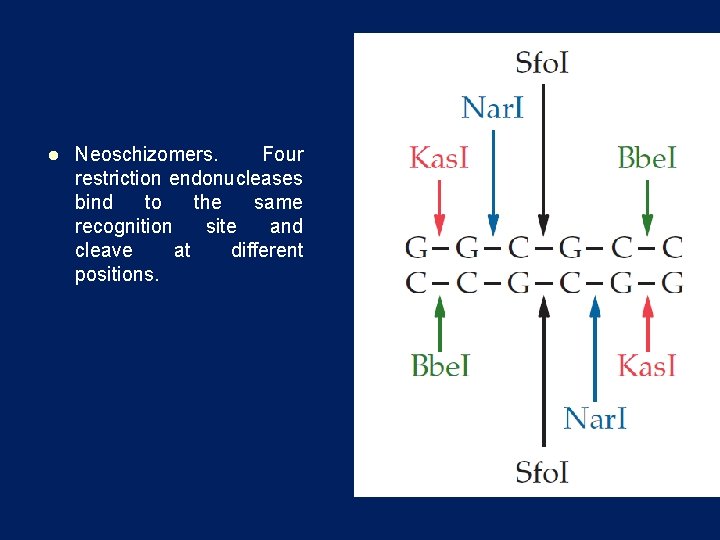
l Neoschizomers. Four restriction endonucleases bind to the same recognition site and cleave at different positions.

In some cases, a restriction endonuclease will cleave a sequence only if the cytosines of the recognition site are not methylated whereas another restriction endonuclease will cut the same sequence if these cytosines are methylated. Hpa. II cleaves only nonmethylated CCGG sites GGCC sites, Msp. I, an isoschizomer of Hpa. II, cuts this sequence regardless of cytosine methylation

Isocaudomers are pairs of restriction enzymes that have slightly different recognition sequences but upon cleavage generate identical termini. For example the enzymes Mbo I and Bam. H I are isocaudomers: Mbo I N*GATC N N CTAG*N Bam. H I G*GATC C C CTAG*G

Type I One enzyme with different subunits for recognition, cleavage and methylation. Recognizes and methylates a single sequence but cleaves DNA up to 1000 bp away l The type I systems were the first to be characterized and a typical example is that from E. coli K 12. l The active enzyme consists of two restriction subunits, two modification (methylation) subunits and one recognition subunit. These subunits are the products of the hsd. R, hsd. M and hsd. S genes.

l l l The methylation and cutting reactions both require ATP and S-adenosylmethionine as cofactors. The recognition sequences are quite long with no recognizable features such as symmetry. The enzyme also cuts unmodified DNA at some distance from the recognition sequence. However, because the methylation reaction is performed by the same enzyme which mediates cleavage, the target DNA may be modified before it is cut. Type I systems are of little value for gene manipulation

Type III One enzyme with two different subunits, one for recognition and modification and one for cleavage. Recognizes and methylates same sequence but cleaves 24– 26 bp away

Type IIs Two different enzymes but recognition sequence is asymmetric. Cleavage occurs on one side of recognition sequence up to 20 bp away

Type IIS restriction endonucleases A subgroup of the type II category of restriction enzymes and are occasionally used for cloning and other molecular studies These enzymes cut DNA, usually in both strands, a fixed number of nucleotides away from one end of the recognition site. Moreover, any particular sequence of nucleotides may be present between the binding sequence and the cut sites. The cleavages for most type IIS restriction enzymes are staggered.

Fok. I restriction endonuclease binds to GGATG CCTAC Cuts 9 nucleotides downstream on the upper strand 13 nucleotides downstream on the lower strand, producing a recessed 3′ hydroxyl end a 4 -nucleotide extension at the 5′ phosphate end. GGATGNNNNN CCTACNNNNNNN, where N denotes A, C, G, or T. Of course, with this single-letter code A simpler notation is GGATG(N)9 CCTAC(N)13 Or GGATG(9/13). Few type IIS restriction endonucleases cleave DNA both upstream and downstream from their recognition sites.

Alkaline phosphatase Removes 5’ phosphate groups of DNA molecules; Bacterial alkaline phosphatase (BAP) is more stable but less active than calf intestinal alkaline phosphatase (CIP) T 4 polynucleotide kinase Catalyzes the transfer of the terminal (g) phosphate from a kinase nucleoside 5’-triphosphate to a 5’-hydroxyl group of a polynucleotide DNase I Degrades double-stranded DNA by hydrolyzing internal phosphodiester linkages

Klenow fragment Proteolytic product of E. coli DNA polymerase I that has both polymerase and 3’-exonuclease activities and no 5’exonuclease activity because fractionation of the digestion products removes the fragment with the 5’ exonuclease activity a Klenow fragment with only DNA polymerase activity because of a mutation in the 3’-exonuclease sequence is also available

E. coli exonuclease III Sequentially removes nucleotides from 3’-OH ends of DNA molecules except from protruding 3’-OH termini Mung bean nuclease Single-strand DNA and RNA endonuclease Nuclease Bal-31 Degrades both 3’ and 5’ ends of DNA without internal cleavages RNase H Degrades the RNA strand of a DNA: RNA hybrid molecule S 1 nuclease Degrades single-stranded DNA

Poly(A) polymerase Adds AMP from ATP to the 3’ end of m. RNA T 7 DNA polymerase and 3’-exonuclease activities Taq DNA polymerase Heat-stable DNA polymerase from Thermus aquaticus β-Agarase I Digests agarose; used to retrieve separated DNA molecules from agarose gels

Reverse Transcriptase l Temin and Baltimore

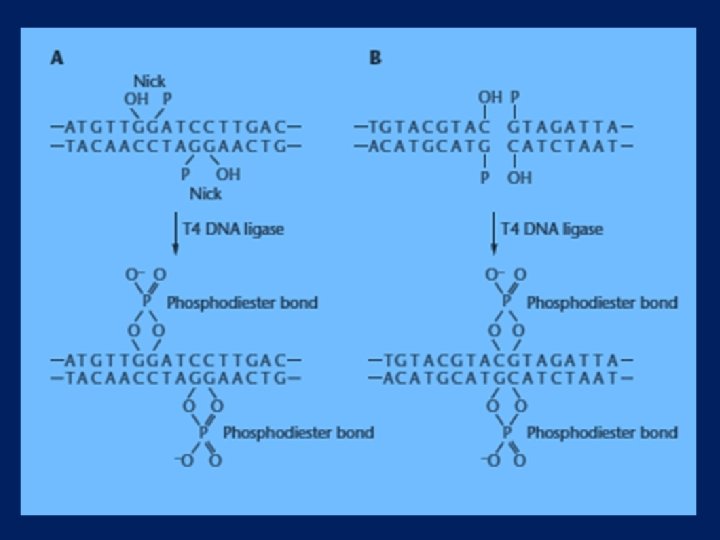
DNA Ligase l l This enzyme catalyzes the formation of phosphodiester bonds at the ends of DNA strands that are already held together by the base pairing of two extensions. DNA ligase also joins blunt ends that come in contact when they both bind to the enzyme. The reaction conditions for DNA ligations depend on whether the DNA molecules have extensions or blunt ends. With protruding ends, the reaction is often carried out at low temperatures for long periods to ensure that the extensions remain base paired. Blunt-end ligations require 10 to 100 times more T 4 DNA ligase than do ligations of DNA molecules with extensions and are conducted atroom temperature because stable base pairing is not required.
- Slides: 44