Molecular Biology Restriction enzymes Restriction enzymes q Endonuclease

Molecular Biology Restriction enzymes

Restriction enzymes q Endonuclease Cleaves internal phosphodiester linkages q Recognize specific double stranded DNA sequences q Different endonucleases recognize different sequences q Recognize palindrome sequences q

Palindromes q The same sequence is read in the 5’ » 3’ direction on both strands 5’-G G A T C C-3’ 3’-C C T A G G-5’
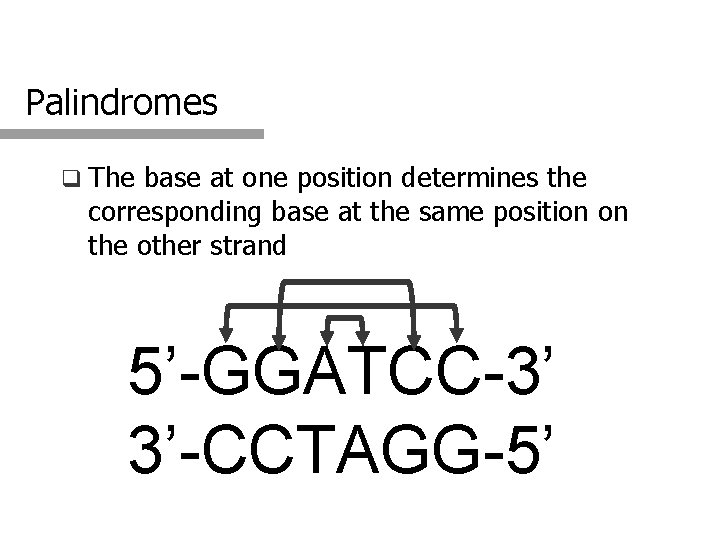
Palindromes q The base at one position determines the corresponding base at the same position on the other strand 5’-GGATCC-3’ 3’-CCTAGG-5’

Palindromes q Complete the following palindromes q. CAC_ __ q. GT _ TA _ q. AT _ _ CACGTC GTATAC ATAT

Palindromes q Problem q How many palindromes of 4 bases are possible? q N 1 N 2 N 3 N 4 q. Position 1: 2: 3: 4: q Therefore: 4 4 1 1 choices (A, G, C, or T) choice (determined by position 2) choice (determined by position 1) 4 X 1 X 1 =16

Problem q How many palindromes does the enzyme Acc. I recognize? q GTMKAC (M= A or C; K = G or T) 1 X 1 X 2 X 1 X 1=2

Frequency of Occurrence q Probability q of finding a given palindrome Depends on the base distribution q Ex. 50% G or C or 1/4 for each base q What is the probability of occurrence of GGATCC? q 1/46 ou 1 divided by (0. 25)-6 = 1/4096 q How many times would this enzyme be expected to cut in a 10 kb genome? q 1/4096 X 10000 = 2. 44 q. Round off to nearest whole number. Therefore 2 X

Frequency of Occurrence q Problem q What is the probability of finding any 4 base palindrome in a genome which is 50% G or C q Number of different sequences of 4 bases q 44 = 256 q Number of different palindromes of 4 bases q 4 X 1 X 1 = 16 q Probability of finding a 4 base palindrome q 16/256 = 1/16

Frequency of Occurrence q Unequal distribution of bases Determine the sequence of each possible palindrome q Independently determine probability of each palindrome according to base distribution q Determine the sum of probabilities q Ex. Acc. I cuts GTMKAC (M= A or C; K = G or T) q Base distribution: 70% G/C, 30% A/T q 2 palindromes possible q q. GTATAC: (0. 35)(0. 15)(0. 35) = 1/16125 q. GTCGAC: (0. 35)(0. 15)(0. 35) = 1/2962 q Probability: 1/2502
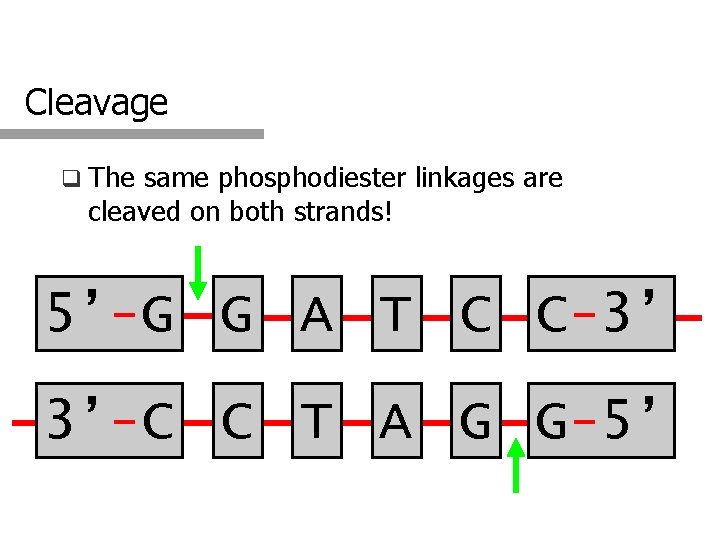
Cleavage q The same phosphodiester linkages are cleaved on both strands! 5’-G G A T C C-3’ 3’-C C T A G G-5’
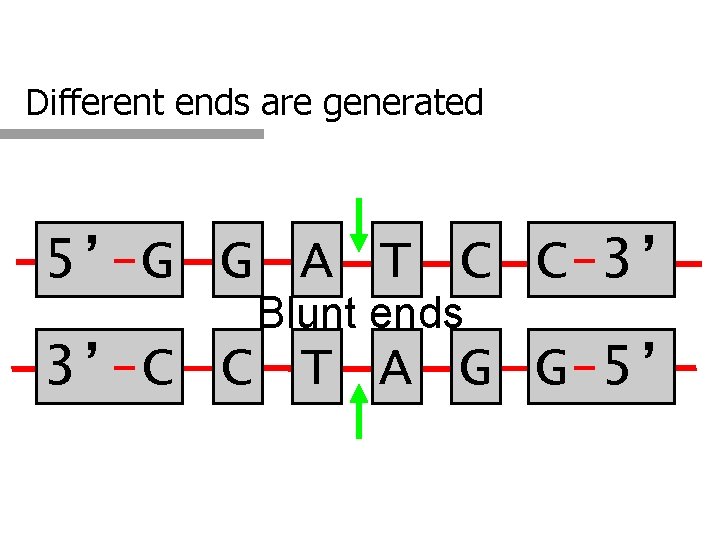
Different ends are generated 5’-G G A T C C-3’ Blunt ends 3’-C C T A G G-5’
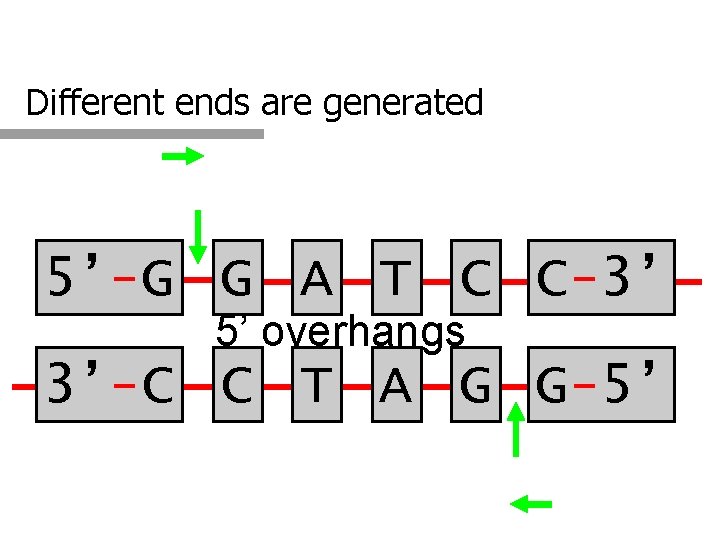
Different ends are generated 5’-G G A T C C-3’ 5’ overhangs 3’-C C T A G G-5’
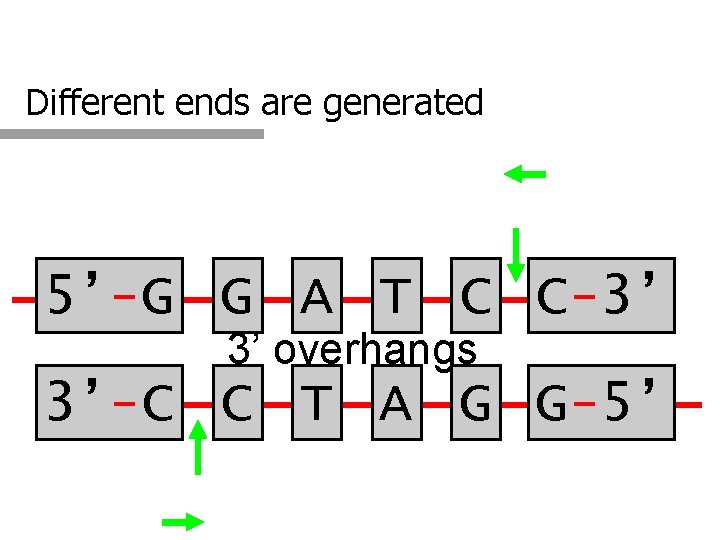
Different ends are generated 5’-G G A T C C-3’ 3’ overhangs 3’-C C T A G G-5’
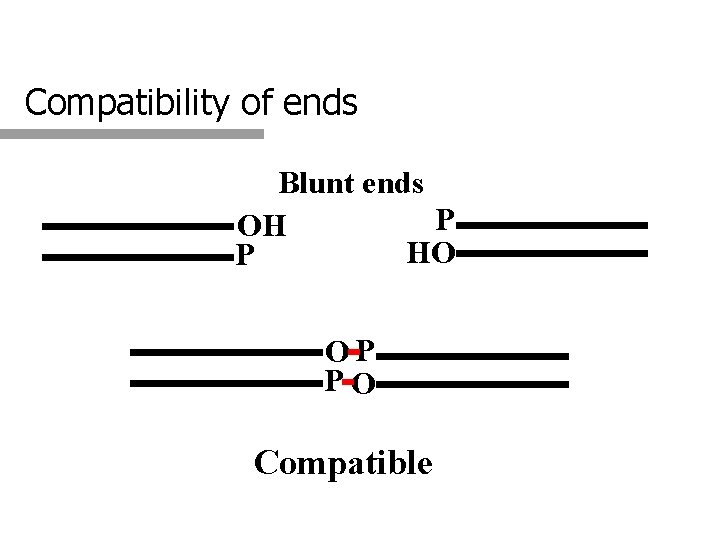
Compatibility of ends Blunt ends P OH HO P OP PO Compatible
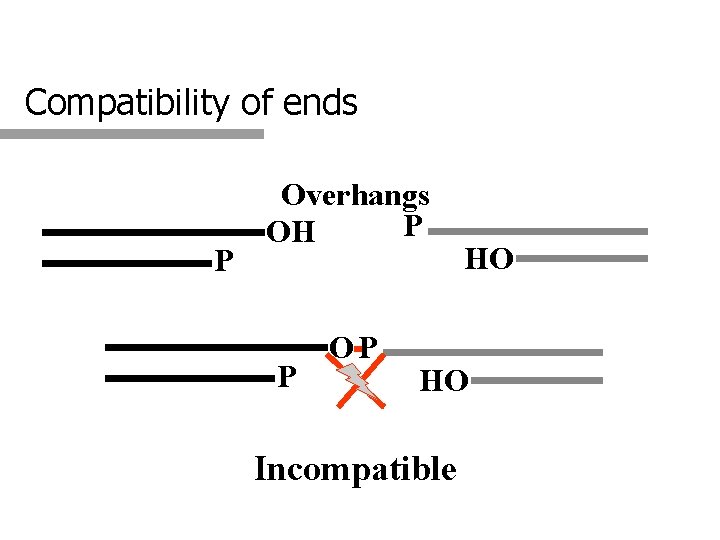
Compatibility of ends P Overhangs P OH P OP HO HO Incompatible
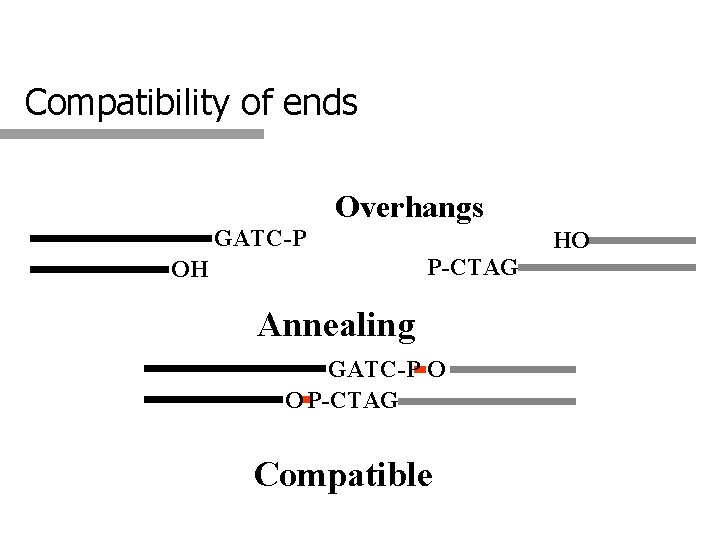
Compatibility of ends Overhangs GATC-P HO P-CTAG OH Annealing GATC-P O O P-CTAG Compatible
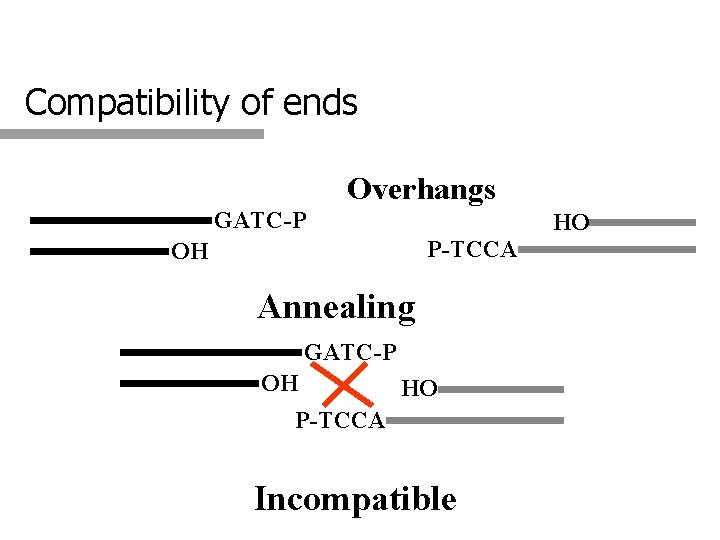
Compatibility of ends Overhangs GATC-P HO P-TCCA OH Annealing GATC-P OH HO P-TCCA Incompatible

Restriction Maps

Compaible ends q Which restriction sites in List 1 would generate compatible ends with those generated from List 2? List 1 List 2 1. 2. 3. 4. A. B. C. D. GA▼TATC AAAT ▼ TT CGC ▼ CGC A ▼ CGCGT 2+A & 2+D CCAT ▼ GGC CAC ▼ GTG CCGC ▼ GG CCATAT ▼ ATGG

Restriction maps q Determining the positions of restriction sites Linear DNA maps q Circular DNA maps (plasmids) q Maps of inserts within vectors q Mapping genomes q Southern analysis q

Approach Determine whether the DNA has digested 2. Is the digestion complete or partial? 3. How many cuts? 4. Determine the relative positions 1.
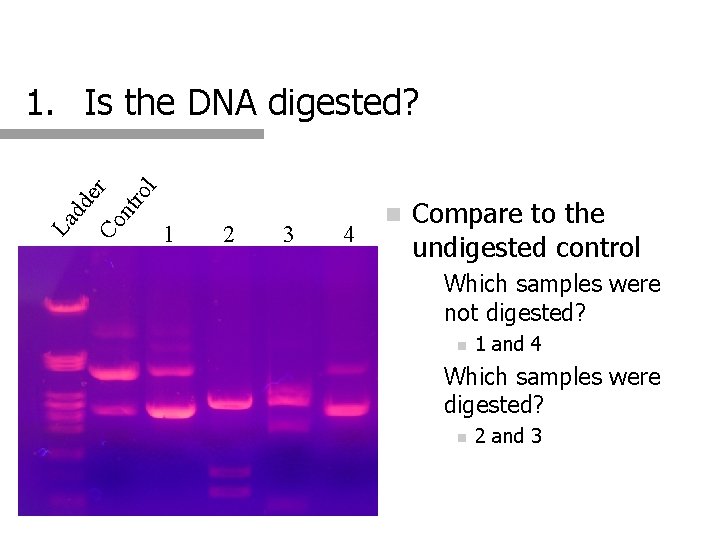
l ro nt Co La dd er 1. Is the DNA digested? 1 2 3 4 n Compare to the undigested control n Which samples were not digested? n n 1 and 4 Which samples were digested? n 2 and 3

2. Is the digestion complete? q Complete q digestion All the DNA molecules are cleaved at all the possible sites q Partial digestion A fraction of the molecules are not digested q Partial undigested q A fraction of the molecules were digested, but not at all the possible sites q Partial digestion q
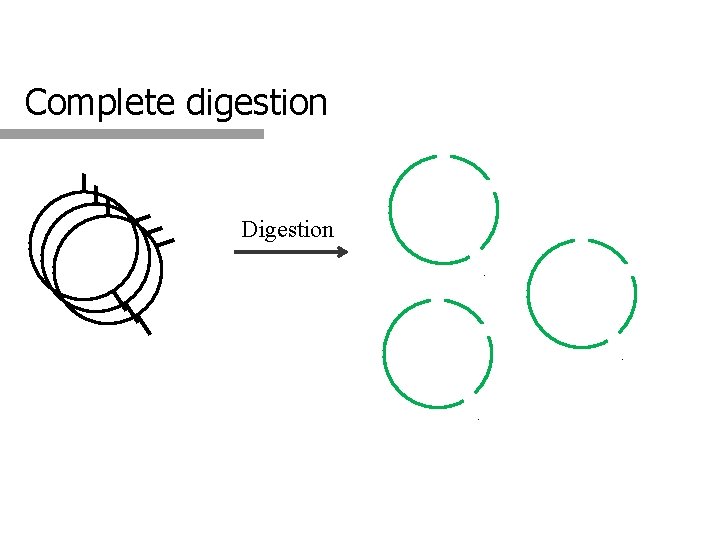
Complete digestion Digestion
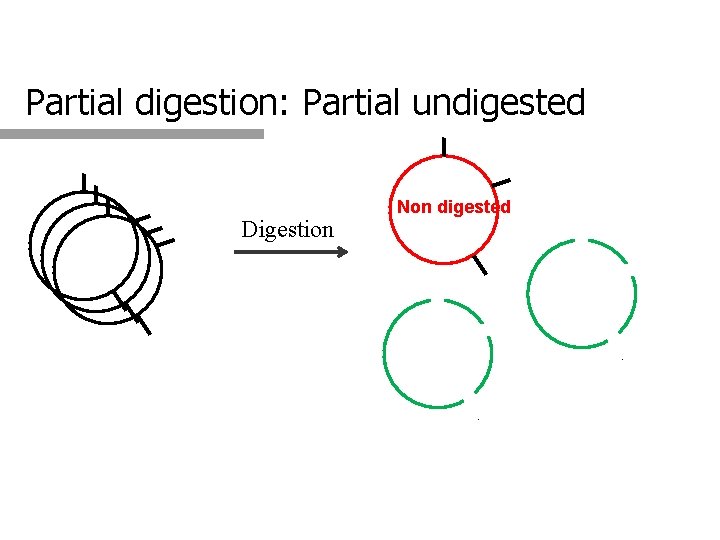
Partial digestion: Partial undigested Digestion Non digested
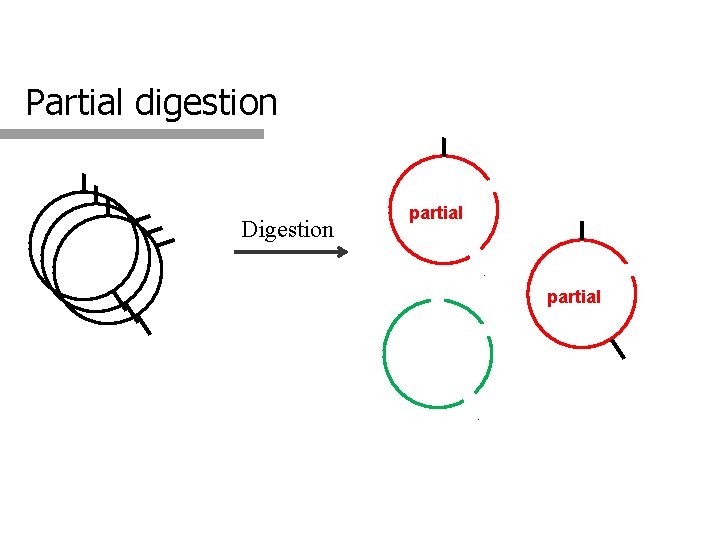
Partial digestion Digestion partial
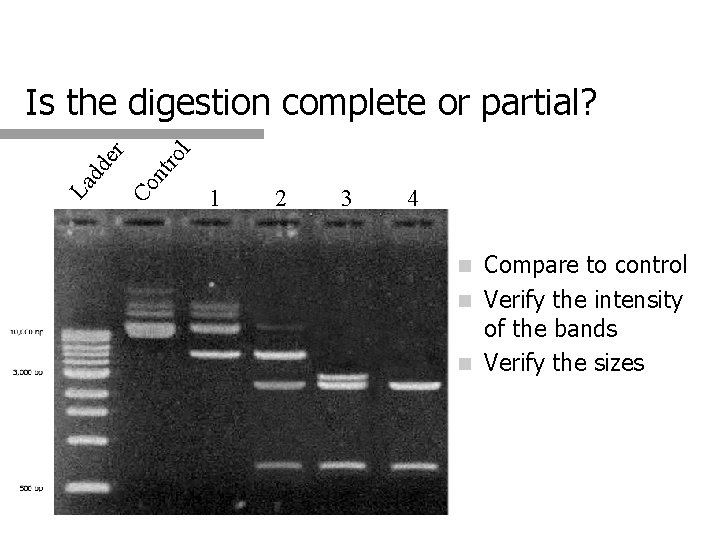
l ro nt Co La dd er Is the digestion complete or partial? 1 2 3 4 Compare to control n Verify the intensity of the bands n Verify the sizes n

3. How many cuts? q Number of sites Circular DNA = number of bands q Linear DNA = Number of bands – 1 q 4. Determine the relative positions q The fragment sizes represent the distances between the sites
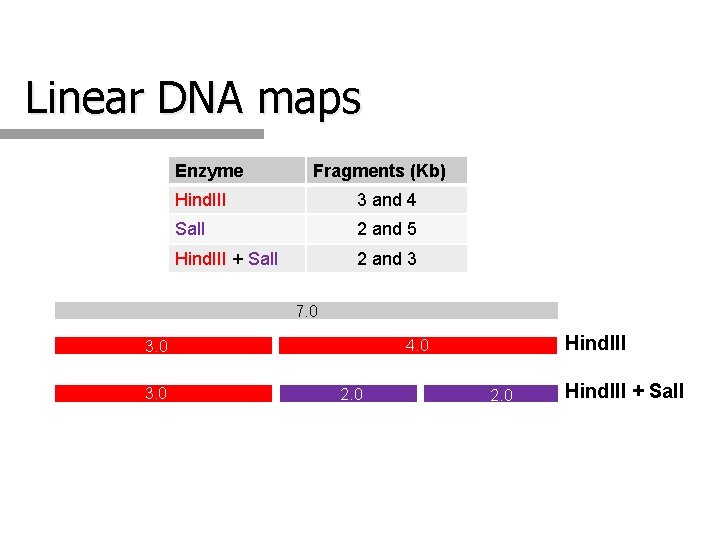
Linear DNA maps Enzyme Fragments (Kb) Hind. III 3 and 4 Sal. I 2 and 5 Hind. III + Sal. I 2 and 3 7. 0 3. 0 Hind. III 4. 0 3. 0 2. 0 Hind. III + Sal. I
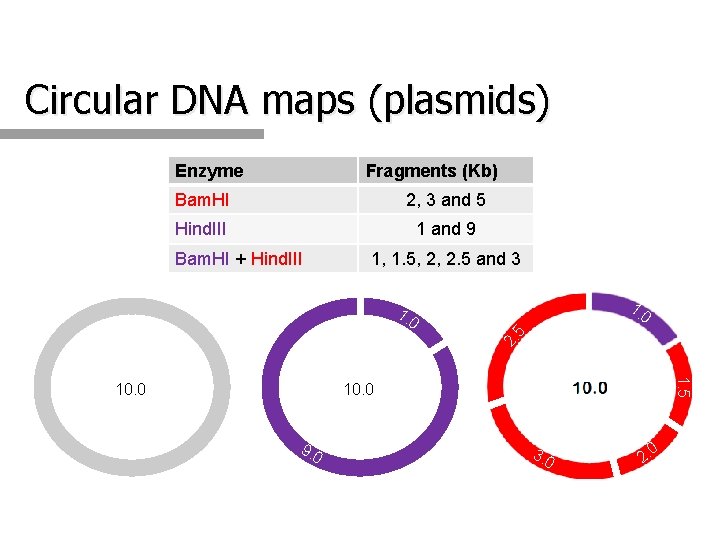
Circular DNA maps (plasmids) Enzyme Fragments (Kb) Bam. HI 2, 3 and 5 Hind. III 1 and 9 Bam. HI + Hind. III 1, 1. 5, 2, 2. 5 and 3 7. 0 1. 0 2. 5 1. 0 1. 5 10. 0 9. 0 3. 0 0 2.
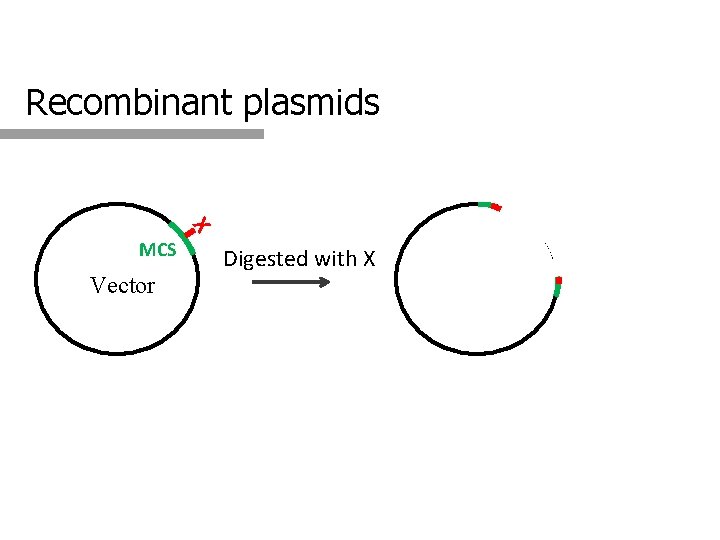
Recombinant plasmids X MCS Vector Digested with X
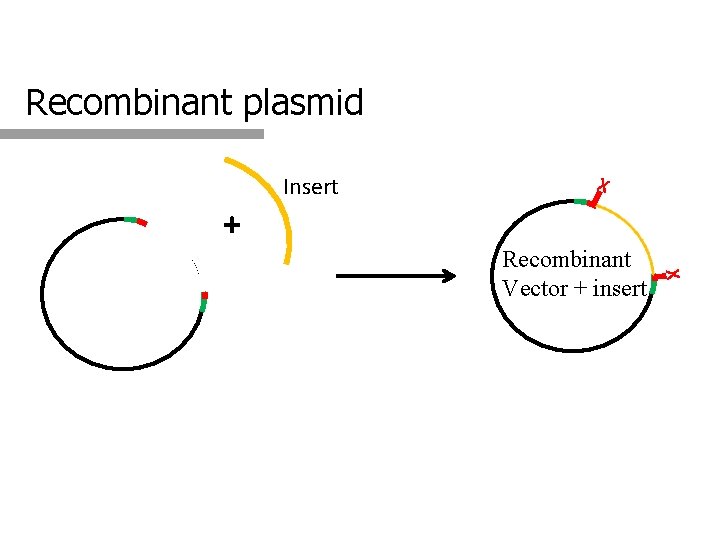
Recombinant plasmid Insert X + X Recombinant Vector + insert
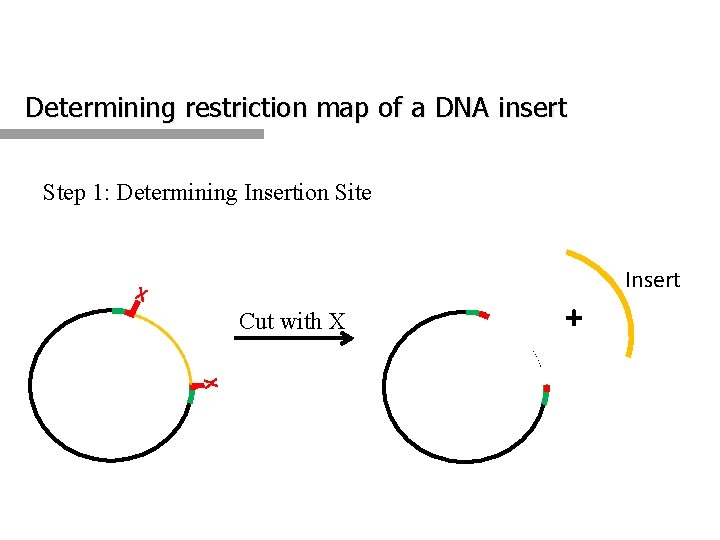
Determining restriction map of a DNA insert Step 1: Determining Insertion Site Insert X Cut with X + X

Insertion maps 1. Total size • 2. Insert size • Enzyme Fragments Bam. HI 7. 7 Kb Eco. RI 1. 0, 3. 7 Kb Pst. I 2. 0 and 5. 7 Xba. I 2. 7 and 5. 0 3. 7. 7 Kb 7. 7 – 2. 7 = 5. 0 Kb Insertion site • Generates 2 fragments one of which is the size of the vector • Xba. I

Determining Relative Orientation Insertion site delimits Right & Left Ex. If Pst is the insertion site: Bam is to the left. If Xba is the insertion site, then Bam is to the right
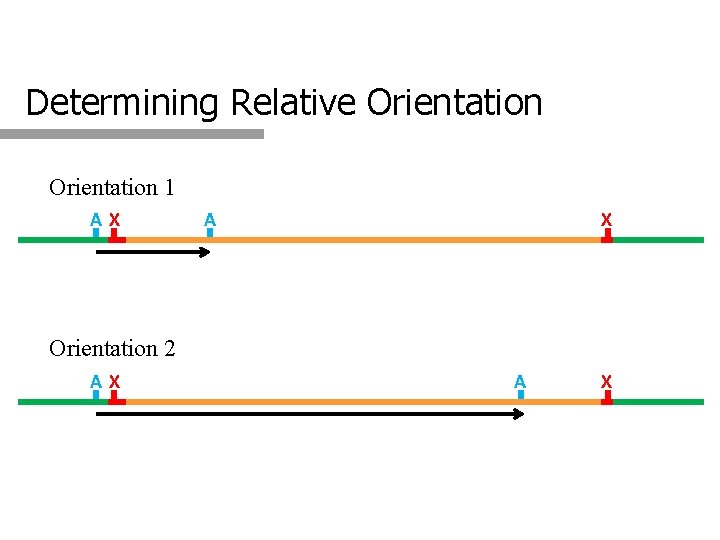
Determining Relative Orientation 1 AX A X Orientation 2 AX A X
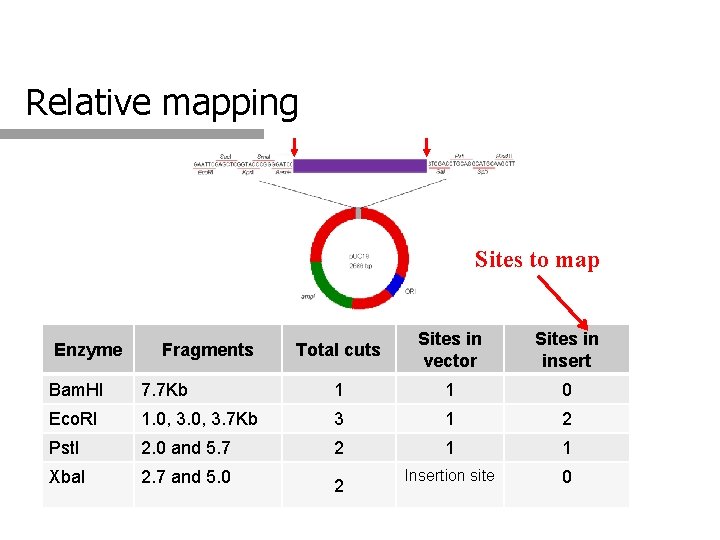
Relative mapping Sites to map Enzyme Fragments Total cuts Sites in vector Sites in insert Bam. HI 7. 7 Kb 1 1 0 Eco. RI 1. 0, 3. 7 Kb 3 1 2 Pst. I 2. 0 and 5. 7 2 1 1 Xba. I 2. 7 and 5. 0 2 Insertion site 0
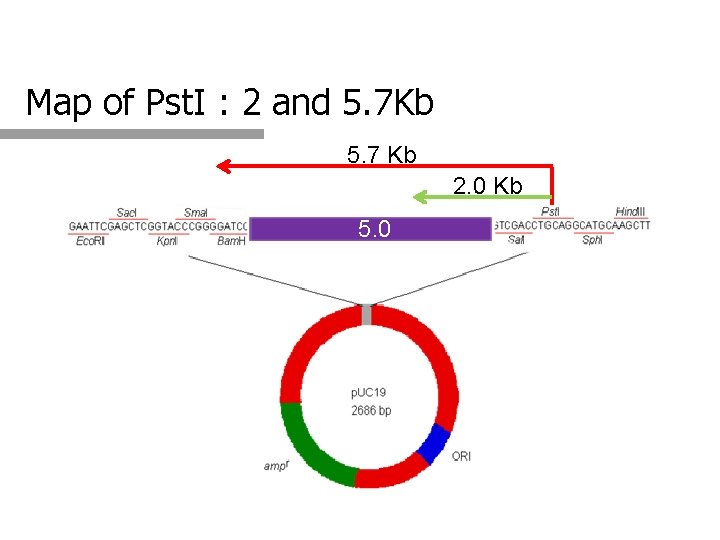
Map of Pst. I : 2 and 5. 7 Kb 5. 7 Kb 2. 0 Kb 5. 0
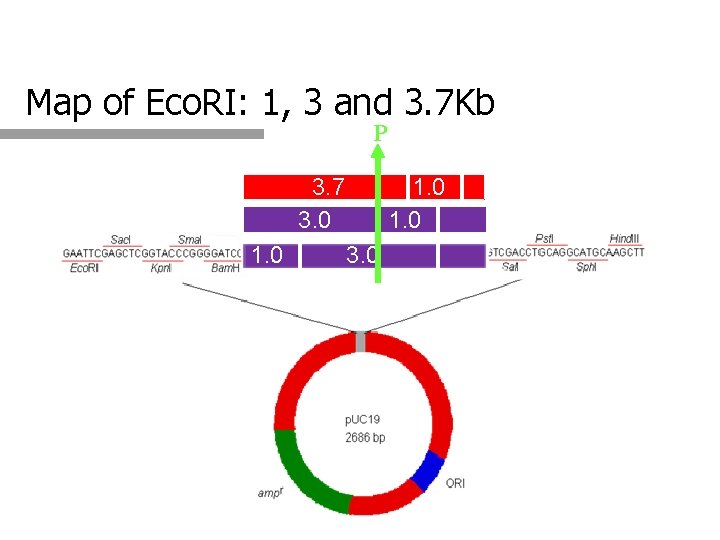
Map of Eco. RI: 1, 3 and 3. 7 Kb P 3. 7 3. 0 1. 0 3. 0

Restriction analysis of large DNA q Purpose Find the position of a sequence in a large DNA fragment q Find similar sequences in other organisms q Analyze non-sequenced DNA q

Digestion of Large DNA q Number of fragments makes analysis impossible

Southern Analysis q Separate DNA fragments according to size q Denature q Hybridize with single stranded probe representing region of interest q Probe allows visualization of only the region of interest

Southern Probe Hybridize to probe Agarose gel
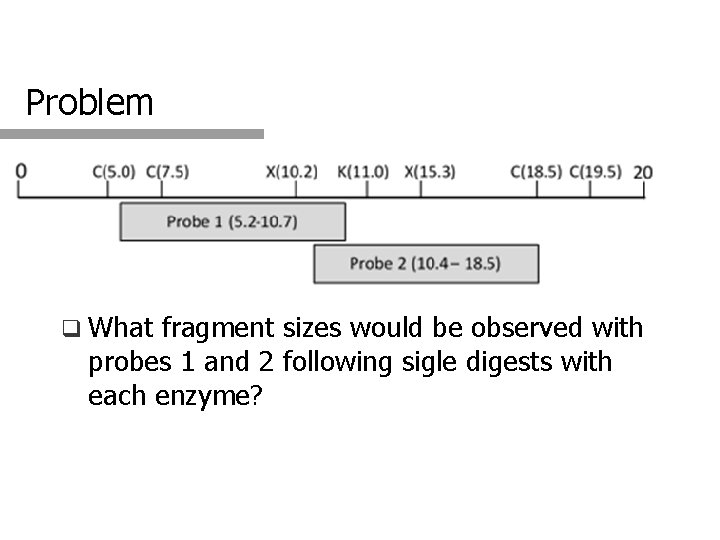
Problem q What fragment sizes would be observed with probes 1 and 2 following sigle digests with each enzyme?

Solution Enzyme C K X Probe 1 2. 5 & 11 kb >9. 8 & 10. 2 kb Probe 2 11 kb >9 & >11 kb >9. 8 kb
- Slides: 46