Epistasis Multilocus Modelling Shaun Purcell Pak Sham SGDP























- Slides: 37

Epistasis / Multi-locus Modelling Shaun Purcell, Pak Sham SGDP, Io. P, London, UK

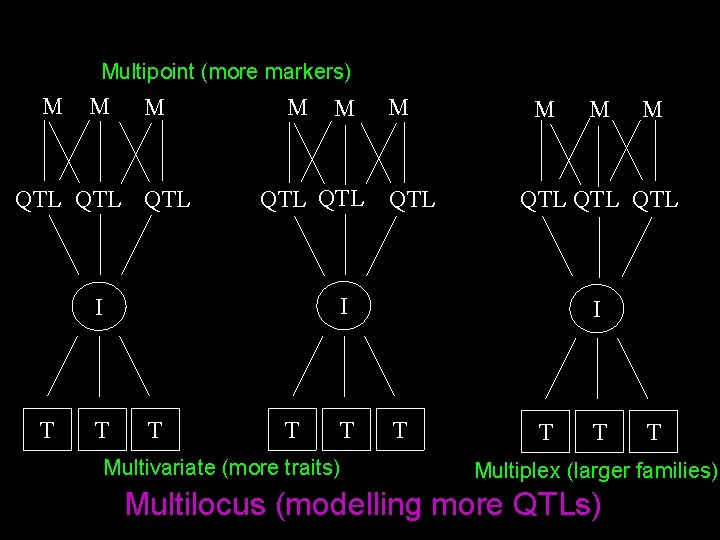
Multipoint (more markers) M M M QTL QTL M T T Multivariate (more traits) M M QTL QTL I I T M I T T Multiplex (larger families) Multilocus (modelling more QTLs)

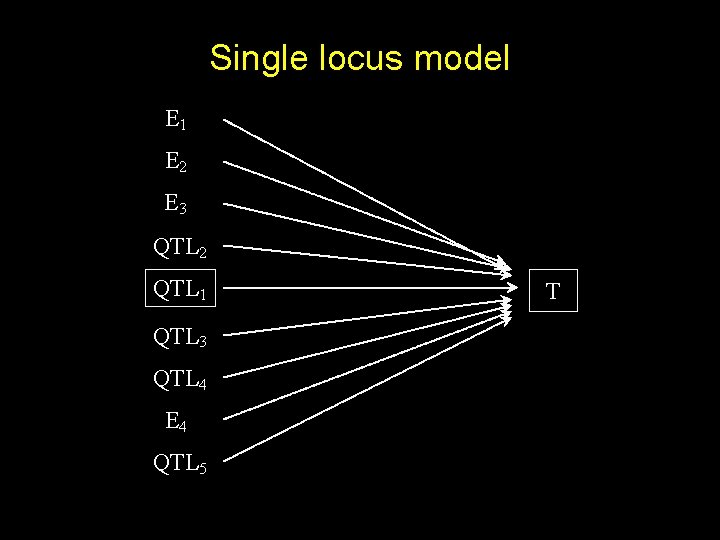
Single locus model E 1 E 2 E 3 QTL 2 QTL 1 QTL 3 QTL 4 E 4 QTL 5 T

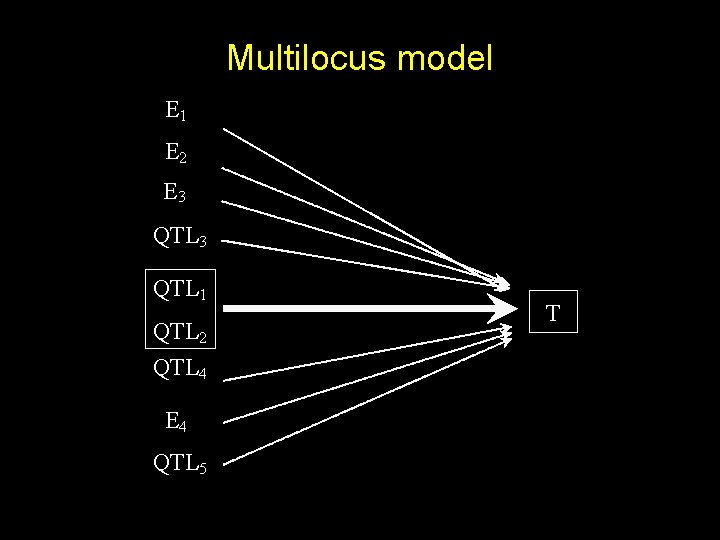
Multilocus model E 1 E 2 E 3 QTL 1 QTL 2 QTL 4 E 4 QTL 5 T

GENE x GENE Interaction GENE x GENE INTERACTION : Epistasis Additive genetic effects : alleles at a locus and across loci independently sum to result in a net phenotypic effect Nonadditive genetic effects : effects of an allele modified by the presence of other alleles (either at the same locus or at different loci)

Nonadditive genetic effects Dominance an allele interaction occurring within one locus Epistasis an interaction occurring between the alleles at two (or more) different loci Additionally, nonadditivity may arise if the effect of an allele is modified by the presence of certain environments

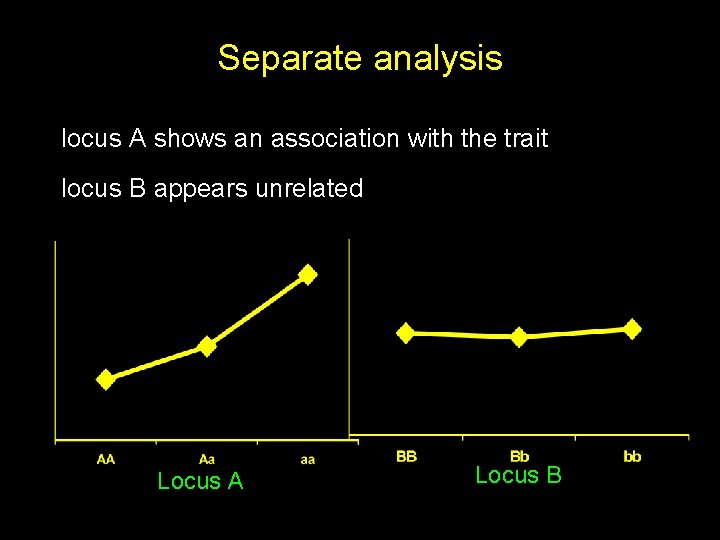
Separate analysis locus A shows an association with the trait locus B appears unrelated Locus A Locus B

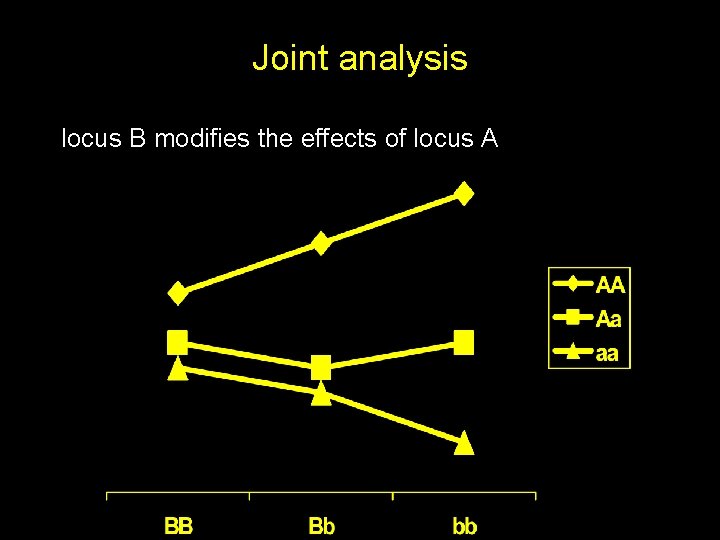
Joint analysis locus B modifies the effects of locus A

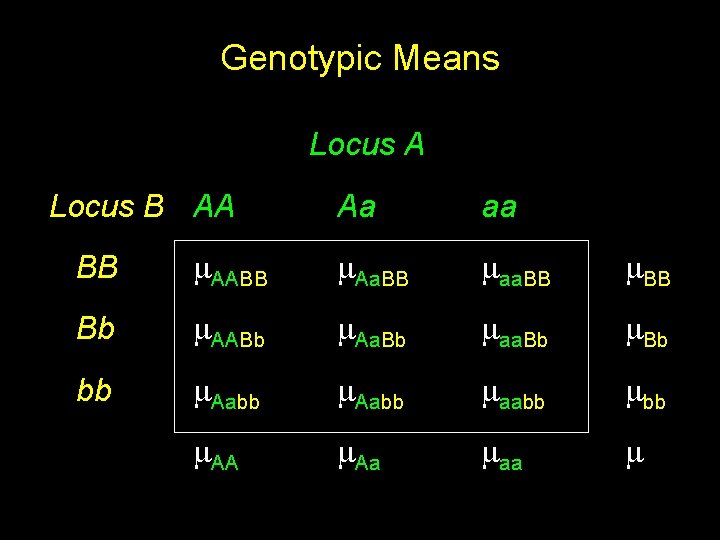
Genotypic Means Locus A Locus B AA Aa aa BB AABB Aa. BB aa. BB Bb AABb Aa. Bb aa. Bb bb Aabb aabb AA Aa aa

Partitioning of effects Locus A M P Locus B

4 main effects M P Additive effects

6 twoway interactions M P Dominance M P

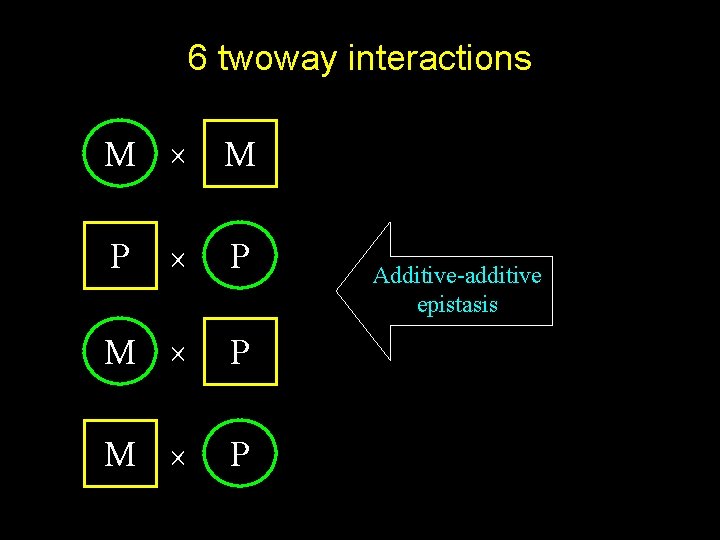
6 twoway interactions M M P M P P Additive-additive epistasis

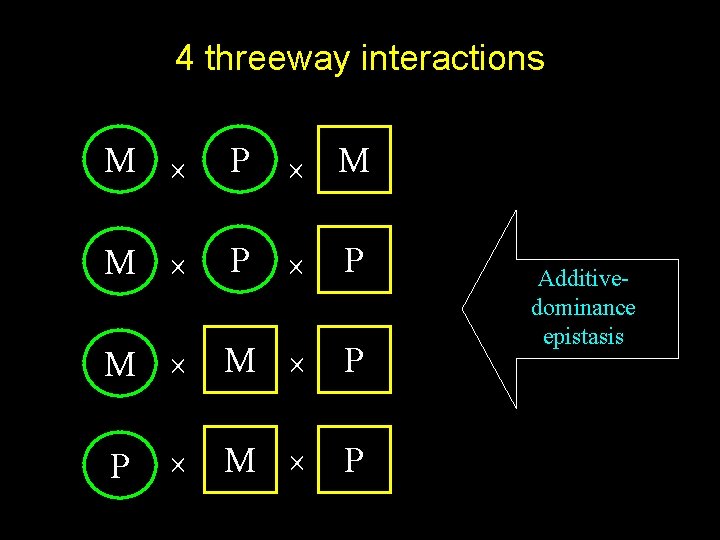
4 threeway interactions M P M M P P M M P P Additivedominance epistasis

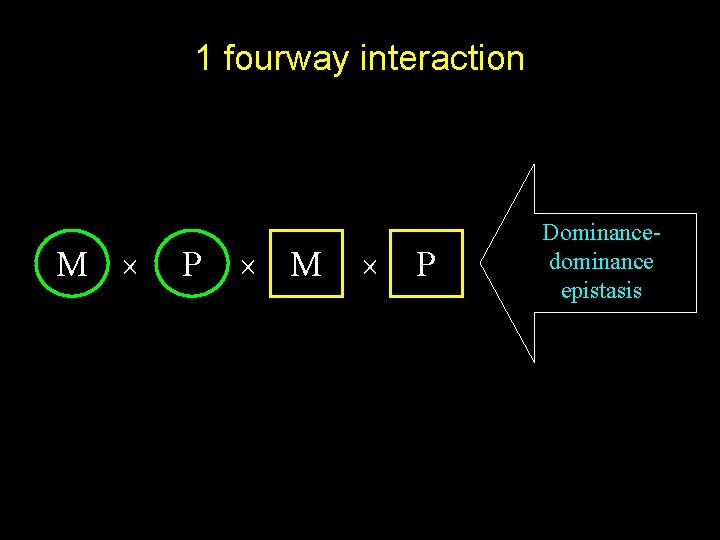
1 fourway interaction M P M P Dominancedominance epistasis

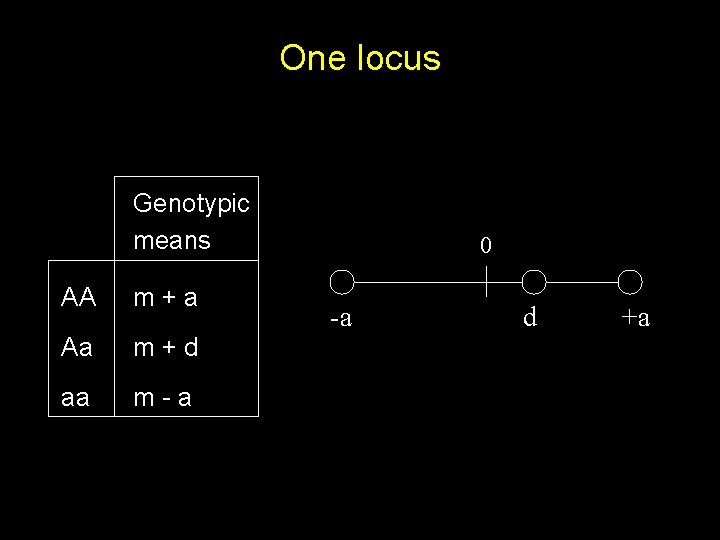
One locus Genotypic means AA m+a Aa m+d aa m-a 0 -a d +a

Two loci AA Aa aa BB m + a. A + a. B + aa m + d. A + a. B + da m – a. A + a. B – aa Bb m + a. A + d. B + ad m + d. A + d. B + dd m – a. A + d. B – ad bb m + a. A – a. B – aa m + d. A – a. B – da m – a. A – a. B + aa

Research questions How can epistasis be modelled under a variance components framework? How powerful is QTL linkage to detect epistasis? How does the presence of epistasis impact QTL detection when epistasis is not modelled?

Variance components QTL linkage : single locus model P=A+D+S+N Var (P) = 2 A + 2 D + 2 S + 2 N Under H 01 : 2 2 ++ 2 2 Cov(P 1, P 2) = ½ 22 AA++z ¼ DD SS where ½==proportion proportionof ofallelessharedidentical-bydescent (ibd) between siblings at that locus z ==probability ¼ prior probability of complete allele sharing ibd between siblings at that locus

Covariance matrix Sib 1 2 A + 2 D + 2 S + 2 N Sib 2 2 A + z 2 D + 2 S Sib 1 2 A + 2 D + 2 S + 2 N Sib 2 ½ 2 A + ¼ 2 D + 2 S Sib 2 2 A + z 2 D + 2 S 2 A + 2 D + 2 S + 2 N Sib 2 ½ 2 A + ¼ 2 D + 2 S 2 A + 2 D + 2 S + 2 N

QTL linkage : two locus model P = A 1 + D 1 + A 2 + D 2 + A 1 A 1 + A 1 D 2 + D 1 A 2 + D 1 D 2 +S+N Var (P) = 2 A + 2 D + 2 AA + 2 AD + 2 DA + 2 DD + 2 S + 2 N

Under linkage : Cov(P 1, P 2) = 2 A + z 2 D + 2 A + z 2 AD + z 2 DA + zz 2 DD + 2 S Under null : Cov(P 1, P 2) = ½ 2 A + ¼ 2 D + E( ) 2 A+E( z) 2 AD +E(z ) 2 DA+ E(zz) 2 DD + 2 S

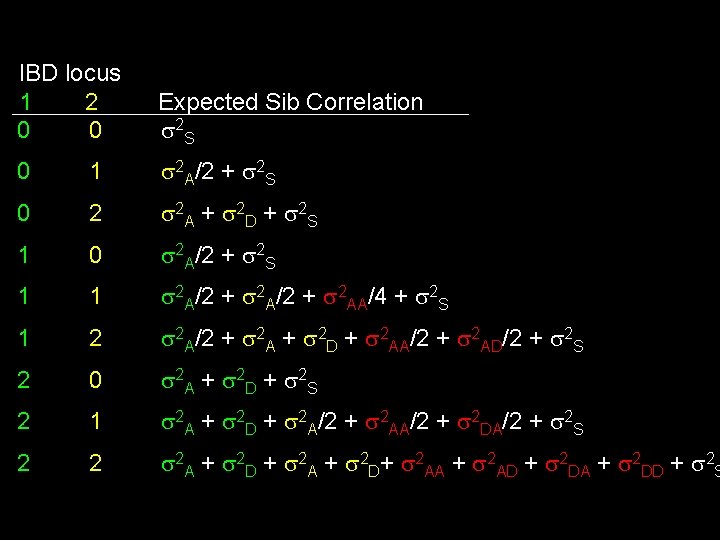
IBD locus 1 2 0 0 Expected Sib Correlation 2 S 0 1 2 A/2 + 2 S 0 2 2 A + 2 D + 2 S 1 0 2 A/2 + 2 S 1 1 2 A/2 + 2 AA/4 + 2 S 1 2 2 A/2 + 2 A + 2 D + 2 AA/2 + 2 AD/2 + 2 S 2 0 2 A + 2 D + 2 S 2 1 2 A + 2 D + 2 A/2 + 2 AA/2 + 2 DA/2 + 2 S 2 2 2 A + 2 D + 2 A + 2 D+ 2 AA + 2 AD + 2 DA + 2 DD + 2 S

Joint IBD sharing for two loci For unlinked loci, Locus A 0 Locus B 1 2 0 1/16 1/8 1/16 1/4 1 1/8 2 1/16 1/8 1/16 1/4 1/4 1/2

Joint IBD sharing for two linked loci at QTL 2 0 1/2 1 at QTL 1 0 1/2 1

Potential importance of epistasis “… a gene’s effect might only be detected within a framework that accommodates epistasis…” Locus A Locus B A 1 A 1 A 1 A 2 A 2 A 2 Freq. 0. 25 0. 50 0. 25 B 1 B 1 0. 25 0 0 1 0. 25 B 1 B 2 0. 50 0 0. 5 0 0. 25 B 2 B 2 0. 25 1 0 0 0. 25 Marginal

Power calculations for epistasis Specify genotypic means, allele frequencies residual variance Calculate under full model and submodels variance components expected non-centrality parameter (NCP)

Submodels Apparent variance components - biased estimate of variance component - i. e. if we assumed a certain model (i. e. no epistasis) which, in reality, is different from the true model (i. e. epistasis) Enables us to explore the effect of misspecifying the model

Detecting epistasis The test for epistasis is based on the difference in fit between - a model with single locus effects and epistatic effects and - a model with only single locus effects, Enables us to investigate the power of the variance components method to detect epistasis

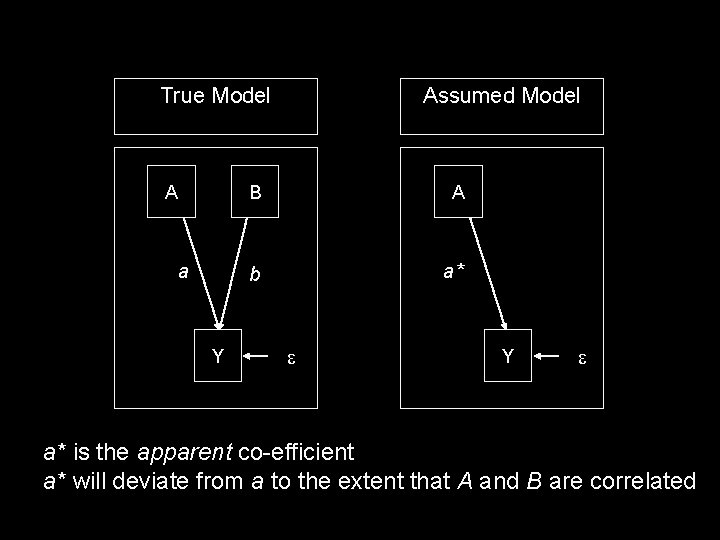
True Model A a Y Assumed Model B A b a* Y a* is the apparent co-efficient a* will deviate from a to the extent that A and B are correlated

Full VA 1 VD 1 VA 2 VD 2 VAA VAD VDA VDD - DD V*A 1 V*D 1 V*A 2 V*D 2 V*AA V*AD V*DA - - AD V*A 1 V*D 1 V*A 2 V*D 2 V*AA - - AA V*A 1 V*D 1 V*A 2 V*D 2 - - -D V*A 1 - V*A 2 - - -A V*A 1 - - - - H 0 - - - - VS and VN estimated in all models

Example 1 : epi 1. mx Genotypic Means B 1 B 1 B 1 B 2 B 2 B 2 A 1 A 1 0 0 1 A 1 A 2 0 0. 5 0 A 2 A 2 1 0 0 Allele frequencies A 1 = 50% ; B 1 = 50% QTL variance 20% Shared residual variance 40% Nonshared residual variance 40% Sample N 10, 000 unselected pairs Recombination fraction Unlinked (0. 5)

Example 2 : epi 2. mx Genotypic Means B 1 B 1 B 1 B 2 B 2 B 2 A 1 A 1 0 1 2 A 1 A 2 0 1 2 A 2 A 2 2 1 0 Allele frequencies A 1 = 90% ; B 1 = 50% QTL variance 10% Shared residual variance 20% Nonshared residual variance 70% Sample N 2, 000 unselected pairs Recombination fraction 0. 1

Exercise Using the module, are there any configurations of means, allele frequencies and recombination fraction that result in only epistatic components of variance? How does linkage between two epistatically interacting loci impact on multilocus analysis?

Poor power to detect epistasis Detection = reduction in model fit when a term is dropped Apparent variance components “soak up” variance attributable to the dropped term artificially reduces the size of the reduction

Epistasis as main effect Epistatic effects detected as additive effects “Main effect” versus “interaction effect” blurred for linkage, main effects and interaction effects are partially confounded

Probability Function Calculator http: //statgen. iop. kcl. ac. uk/bgim/ Genetic Power Calculator http: //statgen. iop. kcl. ac. uk/gpc/