Regressionbased linkage analysis Shaun Purcell Pak Sham Penrose

Regression-based linkage analysis Shaun Purcell, Pak Sham.

• Penrose (1938) • quantitative trait locus linkage for sib pair data • Simple regression-based method • squared pair trait difference • proportion of alleles shared identical by descent (X – Y)2 = 2(1 – r) – 2 Q( ^ – 0. 5) + (HE-SD)
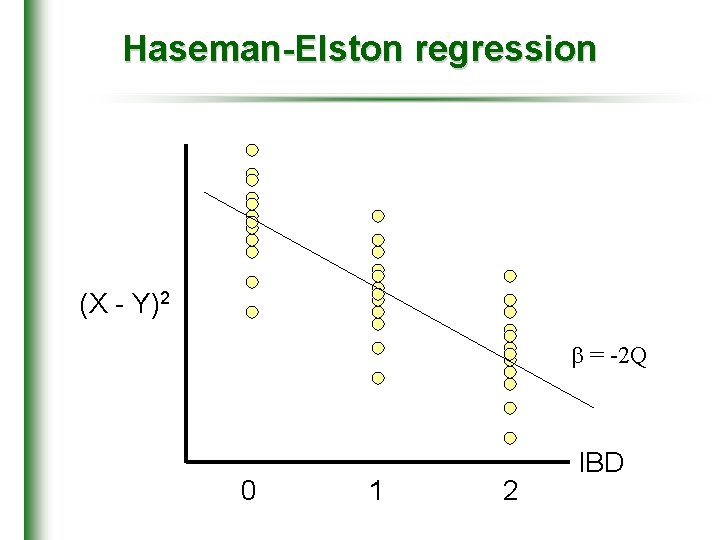
Haseman-Elston regression (X - Y)2 = -2 Q 0 1 2 IBD
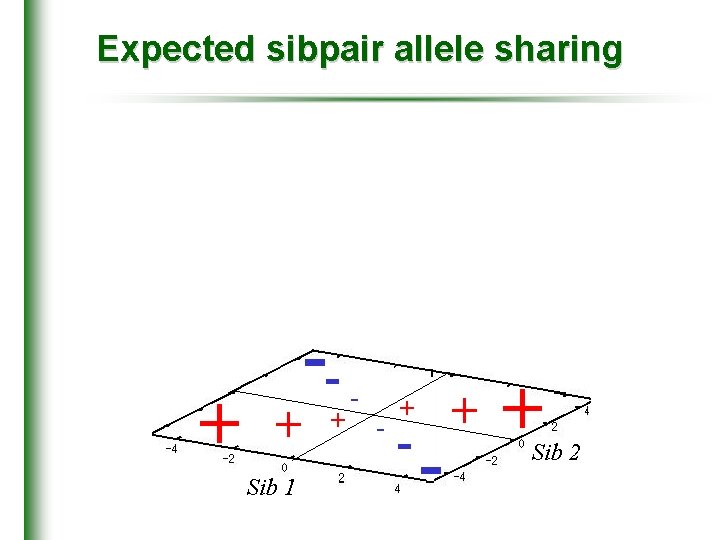
Expected sibpair allele sharing + + Sib 1 -+ - - + + Sib 2
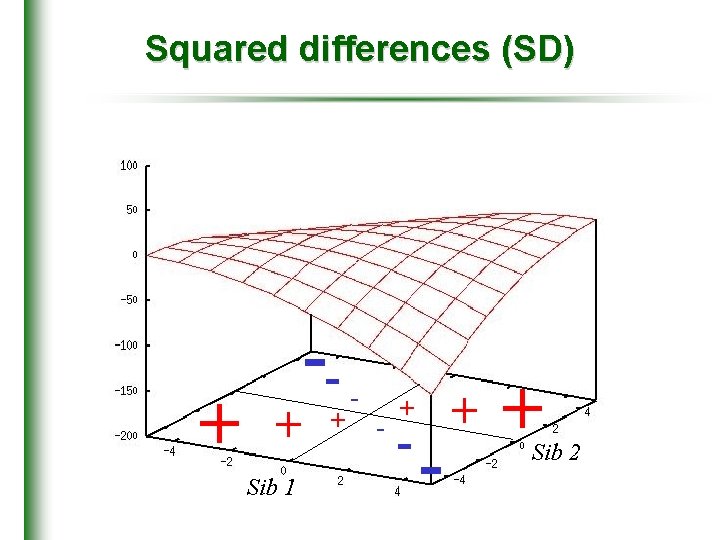
Squared differences (SD) + + Sib 1 -+ - - + + Sib 2

Sums versus differences • Wright (1997), Drigalenko (1998) • phenotypic difference discards sib-pair QTL linkage information • squared pair trait sum provides extra information for linkage • independent of information from HE-SD ^ – 0. 5) + (X + Y)2 = 2(1 + r) + 2 Q( (HE-SS)
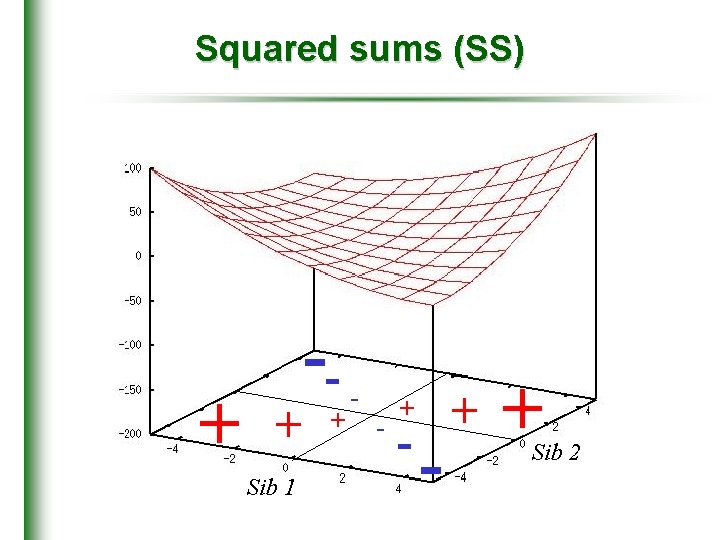
Squared sums (SS) + + Sib 1 -+ - - + + Sib 2
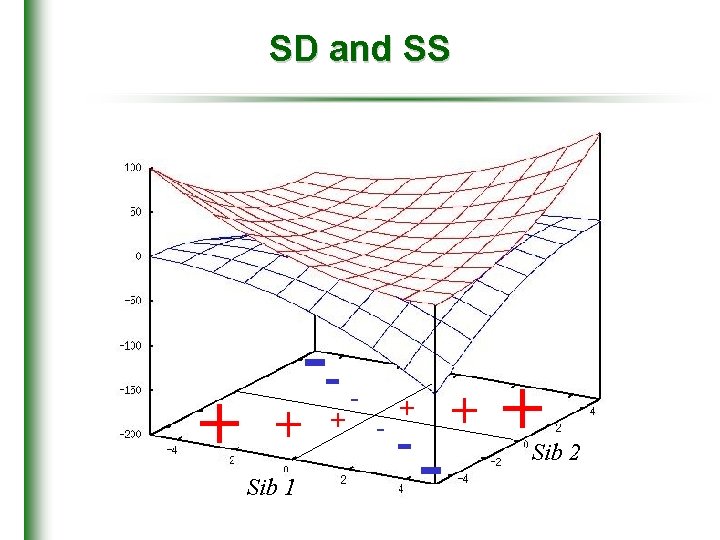
SD and SS + + Sib 1 -+ - - + + Sib 2

• New dependent variable to increase power • mean corrected cross-product (HE-CP) • other extensions • > 2 sibs in a sibship • multivariate • binary traits multiple trait loci and epistasis multiple markers other relative classes
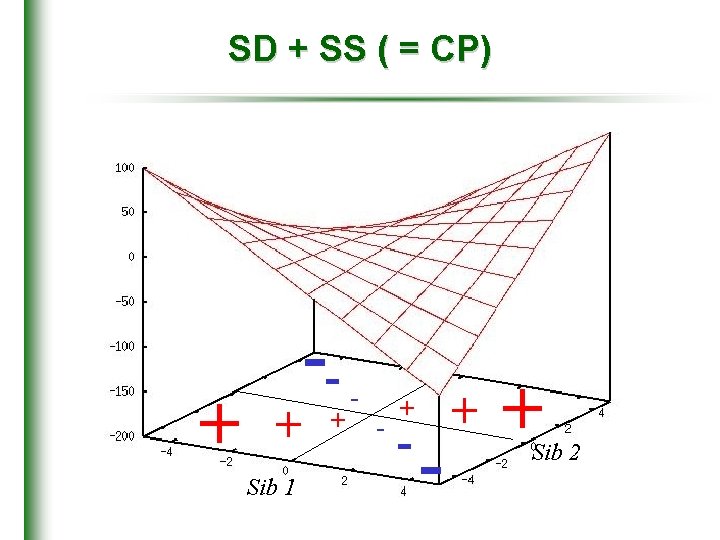
SD + SS ( = CP) + + Sib 1 -+ - - + + Sib 2

Xu et al • With residual sibling correlation • HE-CP in power, HE-SD in power • HE-CP • Propose a weighting scheme
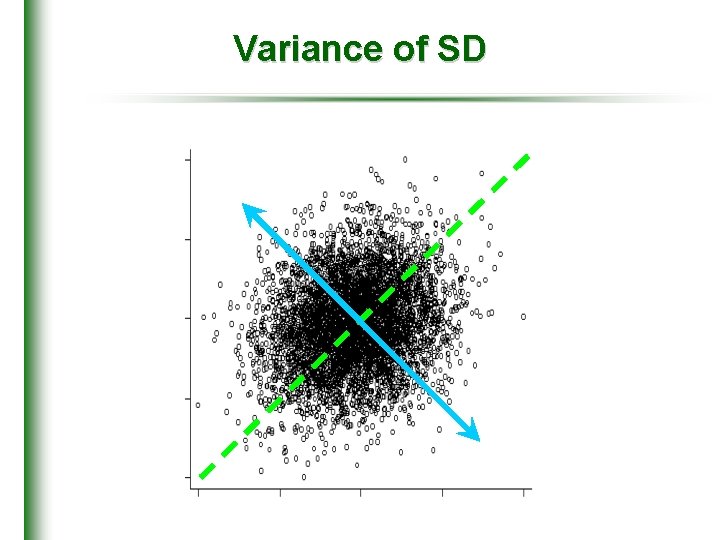
Variance of SD
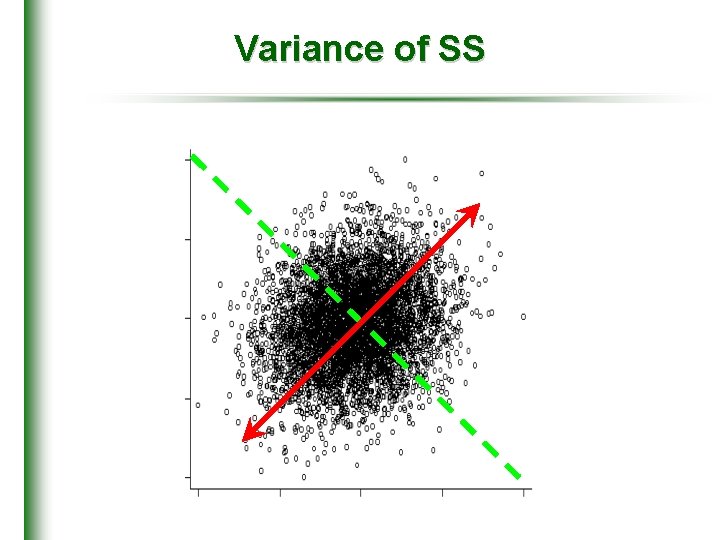
Variance of SS
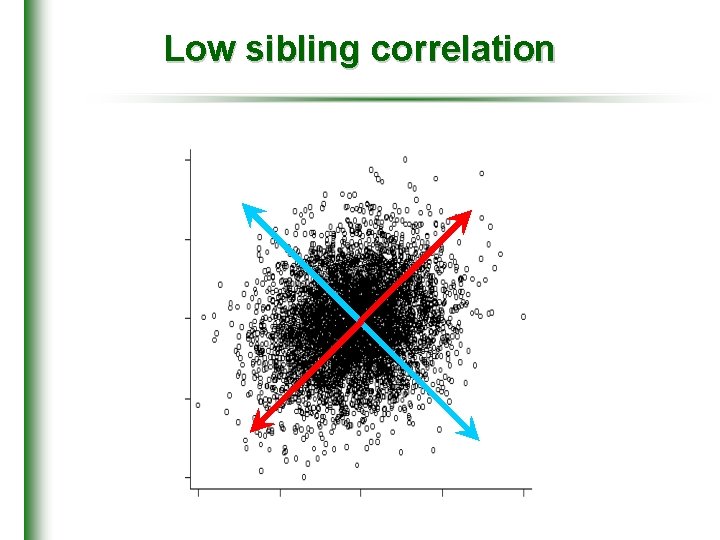
Low sibling correlation

Increased sibling correlation

• Clarify the relative efficiencies of existing HE methods • Demonstrate equivalence between a new HE method and variance components methods • Show application to the selection and analysis of extreme, selected samples

Haseman-Elston regressions (X – Y)2 = 2(1 – r) – 2 Q( – 0. 5) + HE-SD (X + Y)2 = 2(1 + r) + 2 Q( – 0. 5) + HE-SS XY = r + Q( – 0. 5) + HE-CP

NCPs for H-E regressions Dependent Variance of Dependent (X – Y)2 8(1 – r )2 (X + Y)2 8(1 + r )2 XY 1 + r 2 NCP per sibpair

Weighted H-E • Squared-sums and squared-differences • orthogonal components in the population • Optimal weighting • inverse of their variances

Weighted H-E • A function of • square of QTL variance • marker informativeness • complete information = 0. 0125 • sibling correlation • Equivalent to variance components • to second-order approximation • Rijsdijk et al (2000)

Combining into one regression • New dependent variable : • a linear combination of • squared-sum • and squared-difference • weighted by the population sibling correlation:
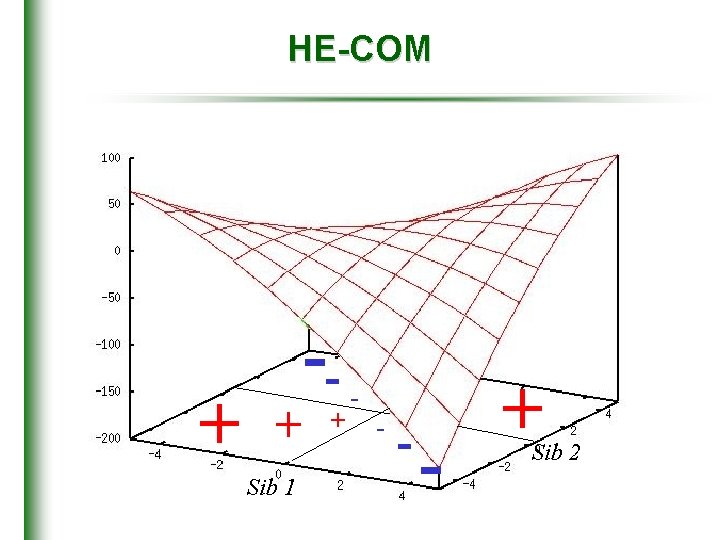
HE-COM + + Sib 1 -+ - - - + Sib 2

Simulation • Single QTL simulated • accounts for 10% of trait variance • 2 equifrequent alleles; additive gene action • assume complete IBD information at QTL • Residual variance • shared and nonshared components • residual sibling correlation : 0 to 0. 5 • 10, 000 sibling pairs • 100 replicates • 1000 under the null
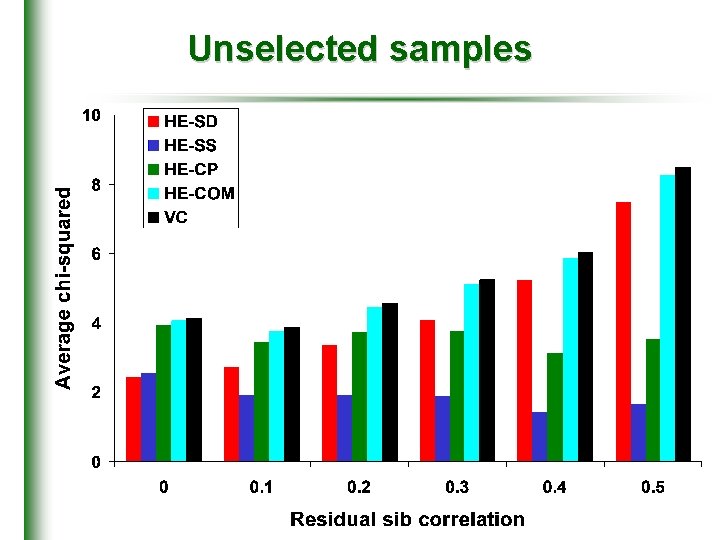
Unselected samples

Sample selection • A sib-pairs’ squared mean-corrected DV is proportional to its expected NCP • Equivalent to variance-components based selection scheme • Purcell et al, (2000)
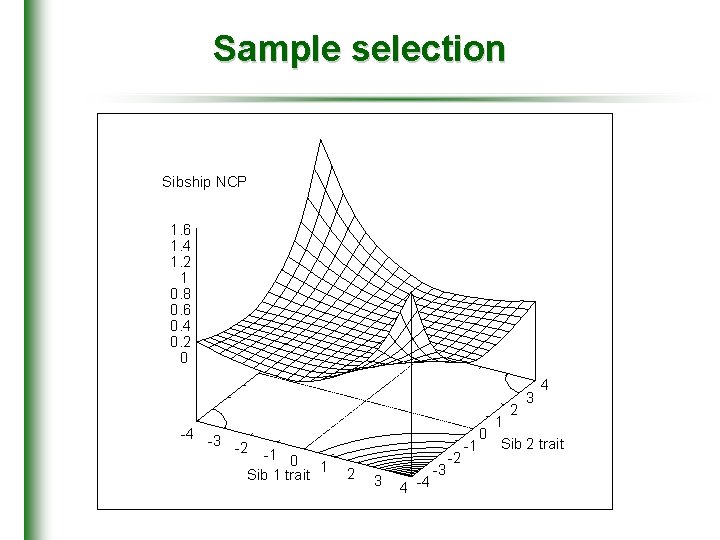
Sample selection Sibship NCP 1. 6 1. 4 1. 2 1 0. 8 0. 6 0. 4 0. 2 0 3 -4 -3 -2 -1 0 1 Sib 1 trait 2 3 4 -4 -3 -2 4 2 1 0 Sib 2 trait -1
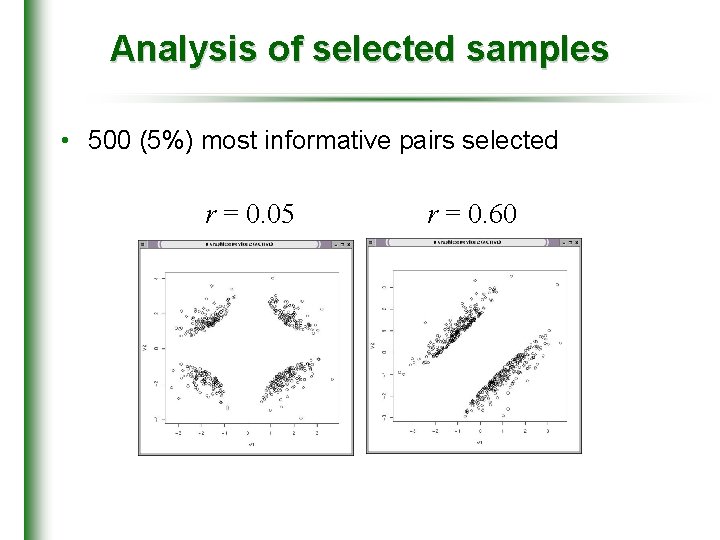
Analysis of selected samples • 500 (5%) most informative pairs selected r = 0. 05 r = 0. 60
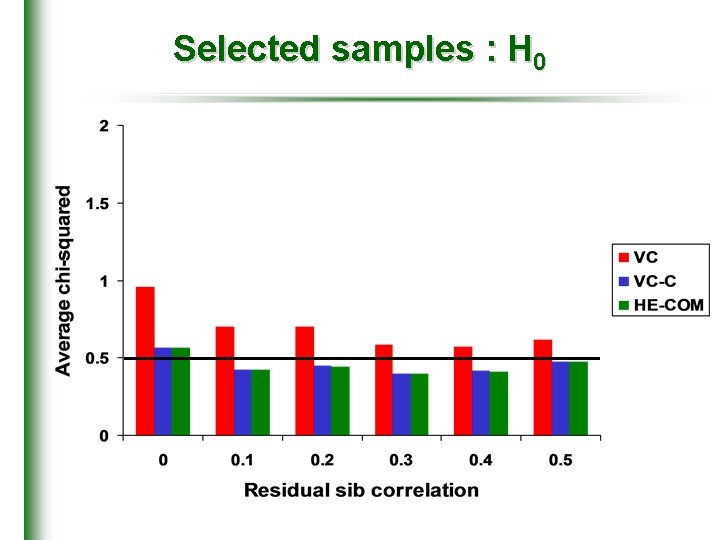
Selected samples : H 0
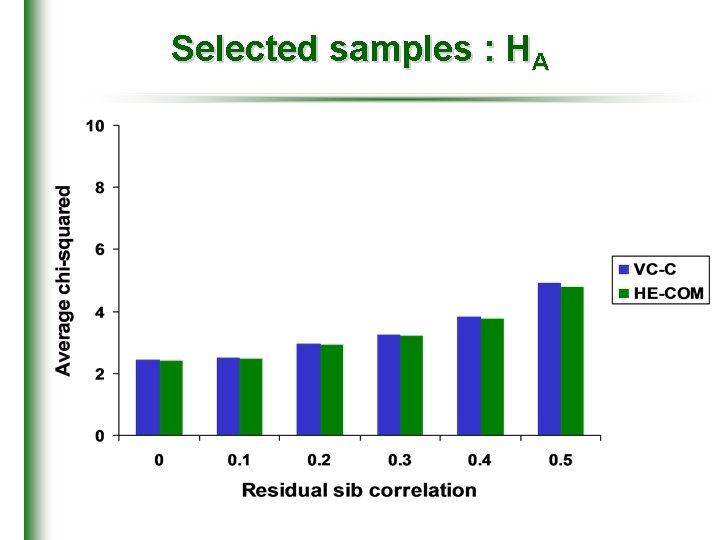
Selected samples : HA

• Variance-based weighting scheme • SD and SS weighted in proportion to the inverse of their variances • Implemented as an iterative estimation procedure • loses simple regression-based framework

• Product of pair values corrected for the family mean • for sibs 1 and 2 from the j th family, • Adjustment for high shared residual variance • For pairs, reduces to HE-SD

Conclusions • Advantages • Efficient • Robust • Easy to implement • Future directions • Weight by marker informativeness • Extension to general pedigrees

The End
- Slides: 33