Biometrical Genetics Pak Sham Shaun Purcell Twin Workshop

Biometrical Genetics Pak Sham & Shaun Purcell Twin Workshop, March 2002

Aims • To gain an appreciation of the scope and rationale of biometric genetics • To understand how the covariance structure of twin data is determined by genetic and environmental factors

Mendel’s Law of Segregation Maternal A 3 A 4 ½ A 1 Paternal A 2 ½ ½ ½ A 1 A 3 ¼ A 1 A 4 ¼ A 2 A 3 ¼ A 2 A 4 ¼

Classical Mendelian Traits • Dominant trait D, absence R – AA, Aa D; aa R • Recessive trait R, absence D – AA D; Aa, aa R • Co-dominant trait, X, Y, Z – AA X; Aa Y; aa Z
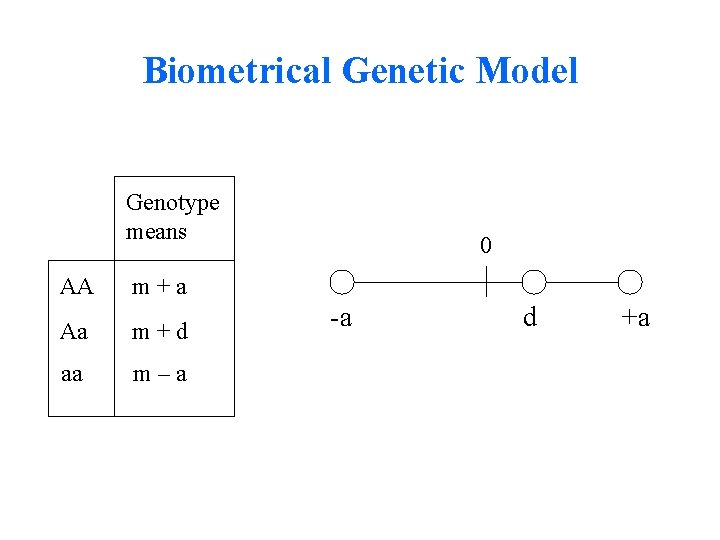
Biometrical Genetic Model Genotype means AA m+a Aa m+d aa m–a 0 -a d +a

Quantitative Traits • Mendel’s laws of inheritance apply to complex traits influenced by many genes • Assume 2 alleles per loci acting additively Genotype AA Aa aa Phenotype 1 0 -1 • Multiple loci – Normal distribution of continuous variation
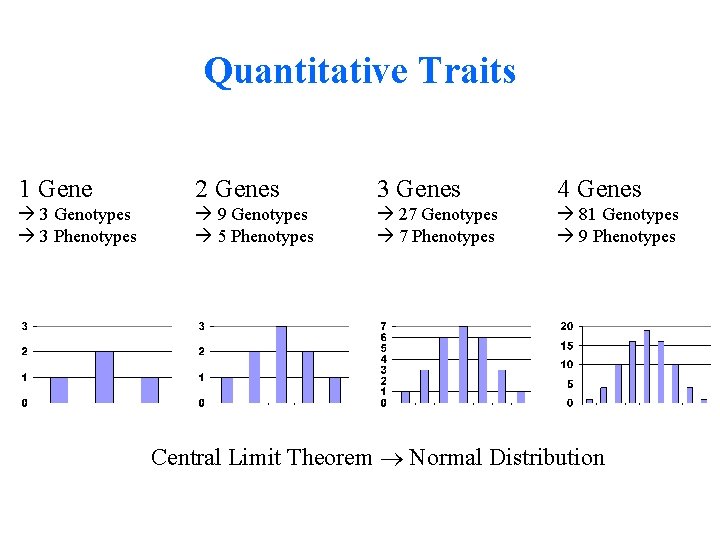
Quantitative Traits 1 Gene 2 Genes 3 Genes 4 Genes 3 Genotypes 3 Phenotypes 9 Genotypes 5 Phenotypes 27 Genotypes 7 Phenotypes 81 Genotypes 9 Phenotypes Central Limit Theorem Normal Distribution
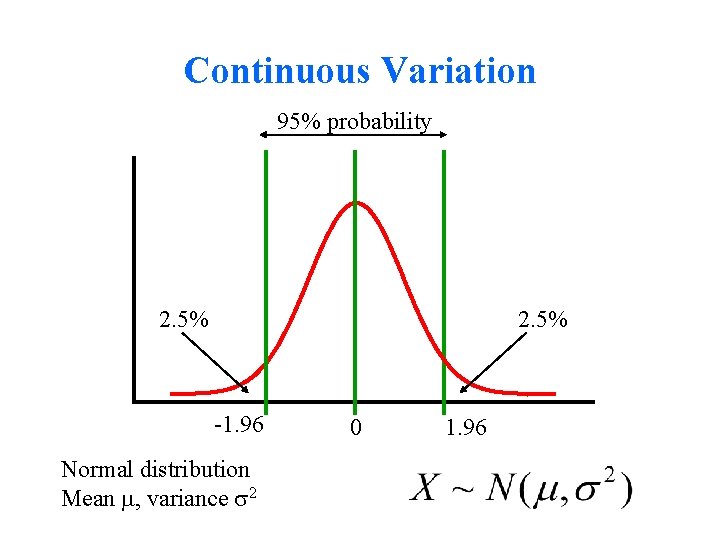
Continuous Variation 95% probability 2. 5% -1. 96 Normal distribution Mean , variance 2 0 1. 96

Familial Covariation Bivariate normal disttribution Relative 2 Relative 1
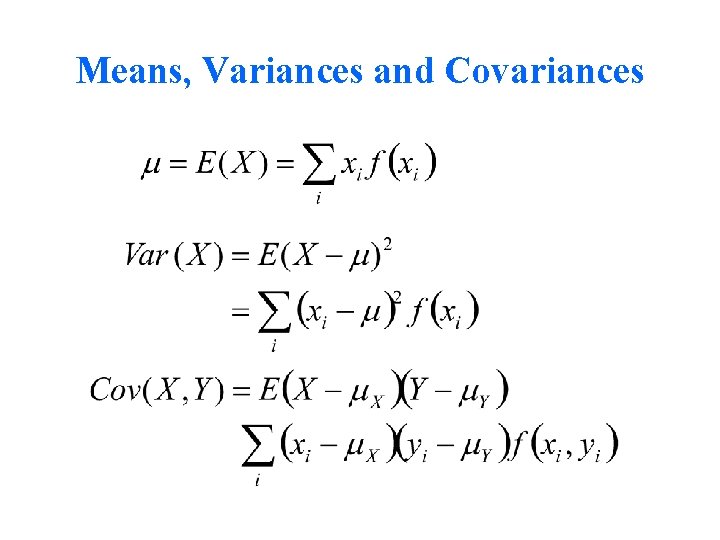
Means, Variances and Covariances

Covariance Algebra Forms Basis for Path Tracing Rules
![Covariance and Correlation is covariance scaled to range [-1, 1]. Covariance and Correlation is covariance scaled to range [-1, 1].](http://slidetodoc.com/presentation_image/38410829f14147e3bedef3482abbc0a4/image-12.jpg)
Covariance and Correlation is covariance scaled to range [-1, 1].

Genotype Frequencies (random mating) A a A p 2 pq p a qp q 2 q p q Hardy-Weinberg frequencies p(AA) = p 2 p(Aa) = 2 pq p(aa) = q 2
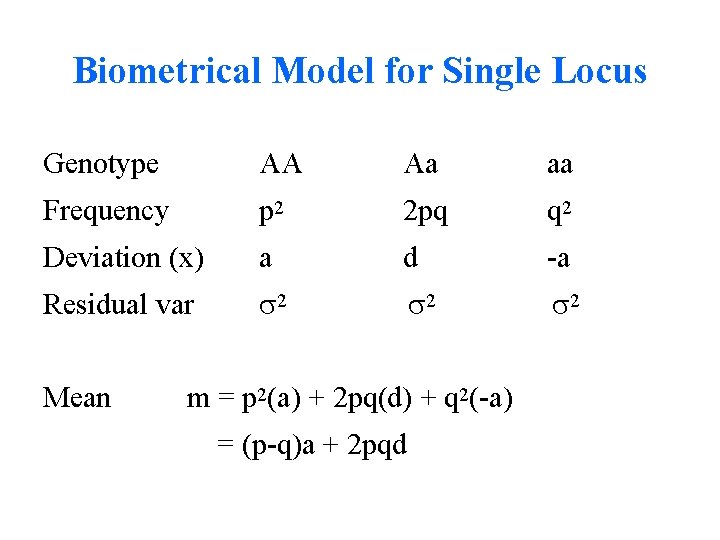
Biometrical Model for Single Locus Genotype AA Aa aa Frequency p 2 2 pq q 2 Deviation (x) a d -a Residual var 2 2 2 Mean m = p 2(a) + 2 pq(d) + q 2(-a) = (p-q)a + 2 pqd

Genetic Variance under Random Mating Genotype AA Aa aa Frequency p 2 2 pq q 2 (x-m)2 (a-m)2 (d-m)2 (-a-m)2 Variance = (a-m)2 p 2 + (d-m)22 pq + (-a-m)2 q 2 = 2 pq[d+(q-p)d]2 + (2 pqd)2 = VA + VD
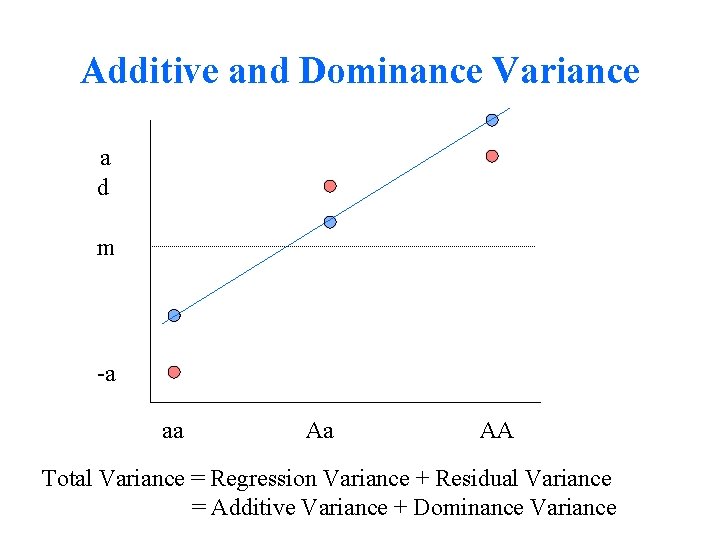
Additive and Dominance Variance a d m -a aa Aa AA Total Variance = Regression Variance + Residual Variance = Additive Variance + Dominance Variance

Cross-Products of Deviations for Pairs of Relatives AA Aa aa AA (a-m)2 Aa (a-m)(d-m)2 aa (a-m)(-a-m)(d-m) (-a-m)2 The covariance between relatives of a certain class is the weighted average of these cross-products, where each cross-product is weighted by its frequency in that class.

Covariance for MZ Twins AA Aa AA p 2 Aa 0 2 pq aa 0 0 aa q 2 Covariance = (a-m)2 p 2 + (d-m)22 pq + (-a-m)2 q 2 = 2 pq[d+(q-p)d]2 + (2 pqd)2 = VA + VD
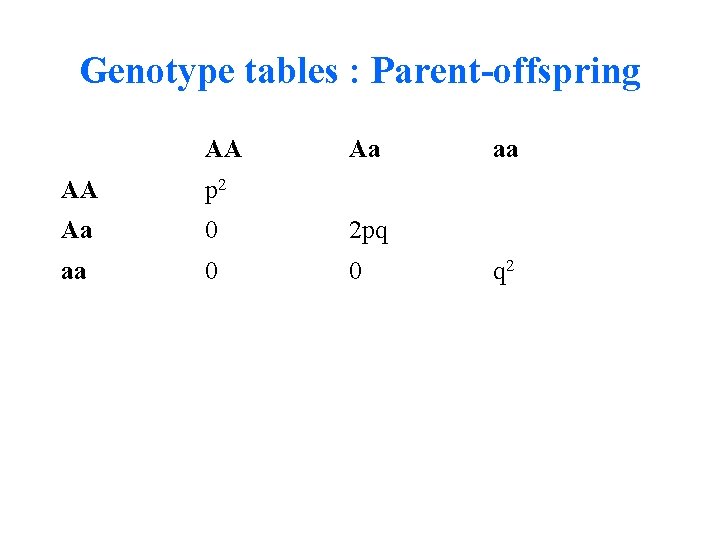
Genotype tables : Parent-offspring AA Aa AA p 2 Aa 0 2 pq aa 0 0 aa q 2
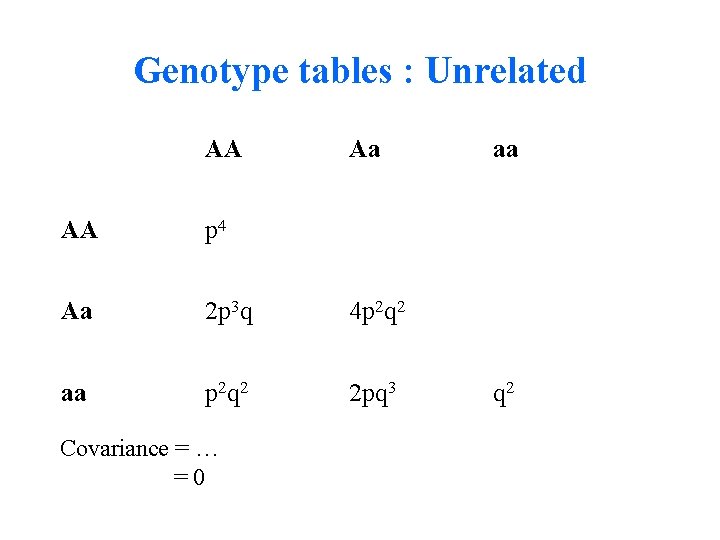
Genotype tables : Unrelated AA Aa AA p 4 Aa 2 p 3 q 4 p 2 q 2 aa p 2 q 2 2 pq 3 Covariance = … =0 aa q 2

Genotype tables : DZ twins AA Aa aa AA p 4 Aa 2 p 3 q 4 p 2 q 2 aa p 2 q 2 2 pq 3 • Weighted average – ¼ MZ twins – ½ Parent-offspring – ¼ Unrelated q 2 Covariance = … = ½VA+¼VD

Environmental components • Shared (C) – Correlation = 1 • Nonshared (E) – Correlation = 0
![ACE Model for twin data 1 [0. 5/1] E C e c PT 1 ACE Model for twin data 1 [0. 5/1] E C e c PT 1](http://slidetodoc.com/presentation_image/38410829f14147e3bedef3482abbc0a4/image-23.jpg)
ACE Model for twin data 1 [0. 5/1] E C e c PT 1 A a A C a c PT 2 E e

Implied covariance matrices
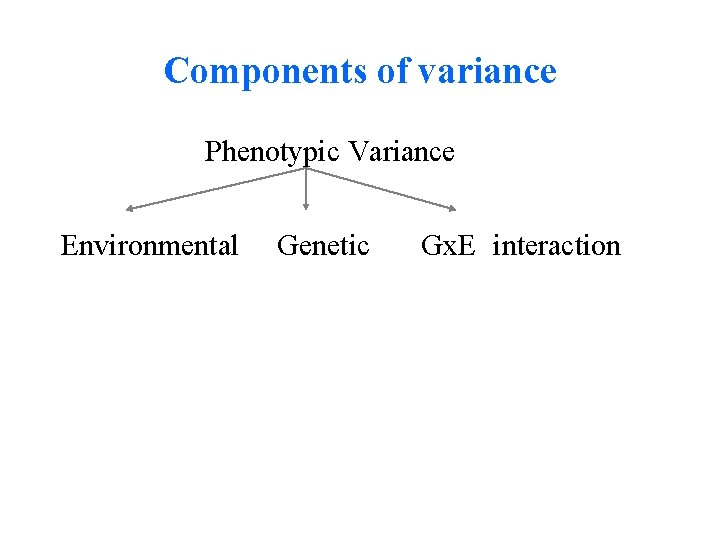
Components of variance Phenotypic Variance Environmental Genetic Gx. E interaction
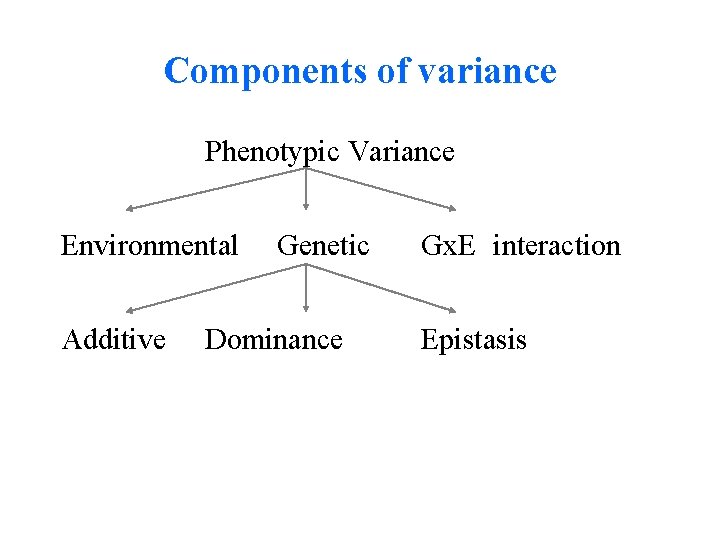
Components of variance Phenotypic Variance Environmental Additive Genetic Dominance Gx. E interaction Epistasis
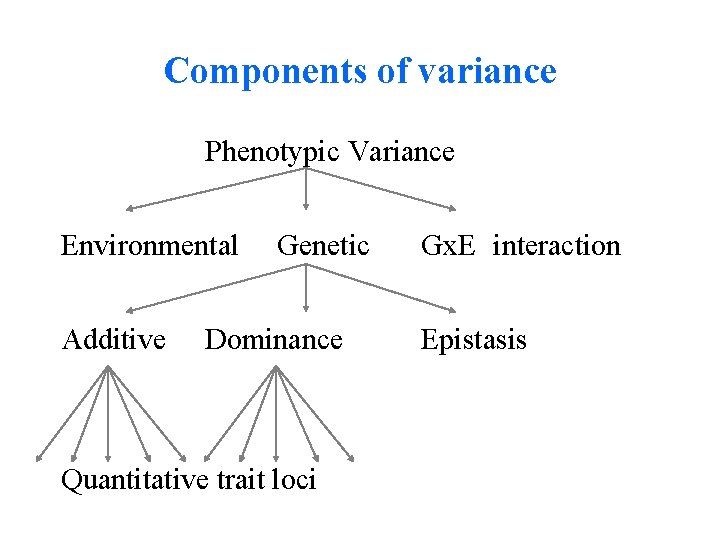
Components of variance Phenotypic Variance Environmental Additive Genetic Dominance Quantitative trait loci Gx. E interaction Epistasis


Parent-offspring
- Slides: 29