Combinatorial methods in Bioinformatics the haplotyping problem Paola

Combinatorial methods in Bioinformatics: the haplotyping problem Paola Bonizzoni DISCo Università di Milano-Bicocca 12/27/2021 1

Content n n n Motivation: biological terms Combinatorial methods in haplotyping Haplotyping via perfect phylogeny : the PPH problem Inference of incomplete perfect phylogeny: algorithms Incomplete pph and missing data Other models: open problems 12/27/2021 2
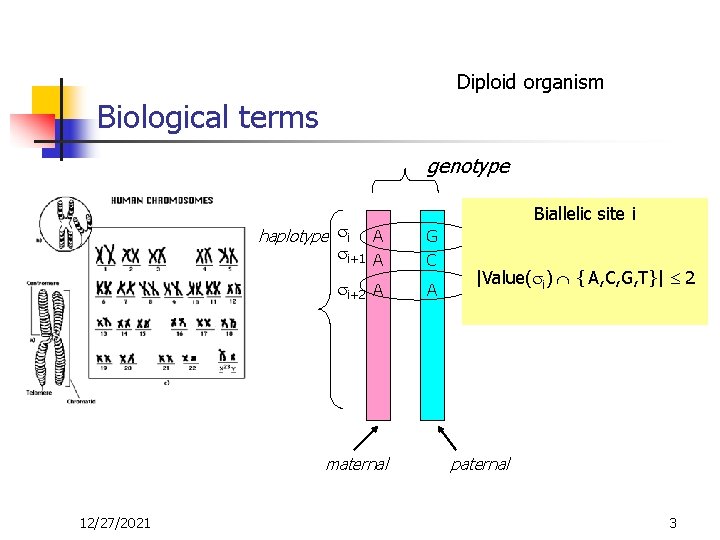
Diploid organism Biological terms genotype haplotype i Biallelic site i A A G C i+2 A A i+1 maternal 12/27/2021 heterozygous |Value( i) { A, C, G, T}| 2 homozygous paternal 3

Motivations n Human genetic variations are related to diseases (cancers, diabetes, osteoporoses) most common variation is the Single Nucleotide Polymorphism (SNP) on haplotypes in chromosomes The human genome project produces genotype sequences of humans n Computational methods to derive haplotypes from genotype data are demanded n Ongoing international Hap. Map project: find haplotype differences on large scale population data graphs n Set-cover problems Combinatorial methods: 12/27/2021 Optimization problems 4

Haplotyping: the formal model n Haplotype: m-vector n Genotype: m-sequence h=<0, 1, …, 0> over {0, 1}m g=<{0, 1}, …, {0, 0}, …{1, 1}> over {0, 1, *} g = <*, *…, 00, …, 1 1> Def. Haplotypes <h, k> solve genotype g iff : g(i)=* implies h(i)= k(i)= g(i) 12/27/2021 h(i) k(i) otherwise 5

Examples g =<0, *, 1, *, 0, 1, 1> k=<0, 0, 1, 1> g solved by <k, h> g h k h=<0, 1, 1, 0, 0, 1, 1> Clark inference rule h 1 g 1 h 2 h 1=<0, 0, 1, 1, 0, 1, 1> g 1 =<0, *, 1, *, 0, 1, 1> h 2=<0, 1, 1, 0, 0, 1, 1> g 2 =<0, 1, *, 0, 0, 1, 1> g 3 =<0, 0, *, *, 1, 1, 1> 12/27/2021 g 3 =<0, 1, 0, *, 0, 1, 1> h 1=<0, 0, 1, 1> h 2=<0, 1, 1, 0, 0, 1, 1> h 3=<0, 1, 0, 0, 0, 1, 1> g 3 =<0, 1, 0, *, 0, 1, 1> 6

Haplotype inference: the general problem Problem HI: Instance: a set G={g 1, …, g m} of genotypes and a set H={h 1, …, h n } of haplotypes, Solution: a set H’ of haplotypes that solves each genotype g in G s. t. H H’. n H’ derives from an inference RULE 12/27/2021 7

Type of inference rules n n Clark’s rule: haplotypes solve g by an iterative rule Gusfield coalescent model: haplotypes are related to genotypes by a tree model n Pedigree data: haplotypes are related to genotypes by a directed graph 12/27/2021 8
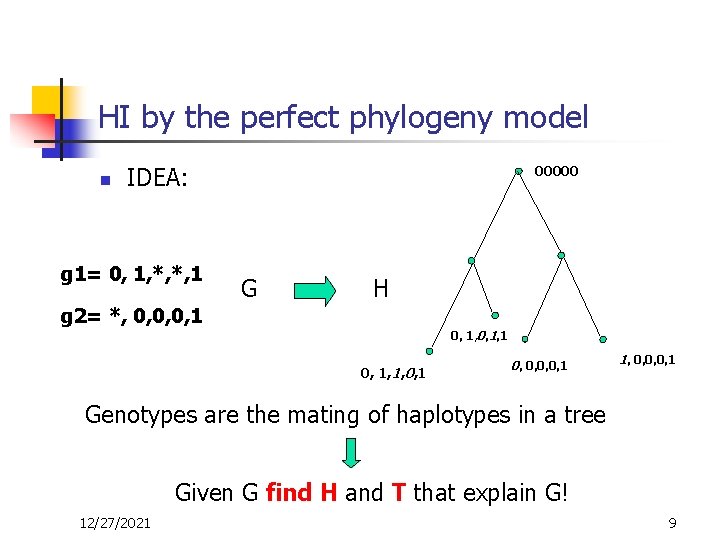
HI by the perfect phylogeny model n 00000 IDEA: g 1= 0, 1, *, *, 1 G H g 2= *, 0, 0, 0, 1, 1, 0, 1 0, 0, 1 1, 0, 0, 0, 1 Genotypes are the mating of haplotypes in a tree Given G find H and T that explain G! 12/27/2021 9
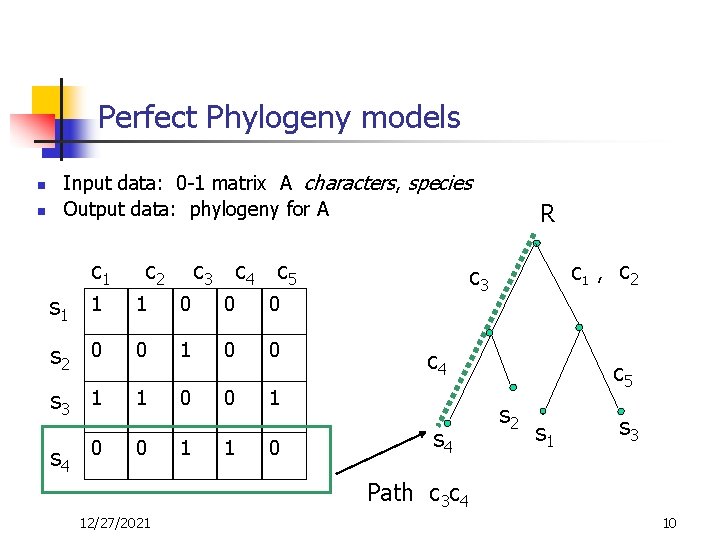
Perfect Phylogeny models n n Input data: 0 -1 matrix A characters, species Output data: phylogeny for A c 1 c 2 c 3 c 4 c 5 s 1 1 1 0 0 0 s 2 0 0 1 0 0 s 3 1 1 0 0 1 s 4 0 0 1 1 0 R c 3 C 1 , c 4 s 4 c 2 c 5 s 2 s 1 s 3 Path c 3 c 4 12/27/2021 10
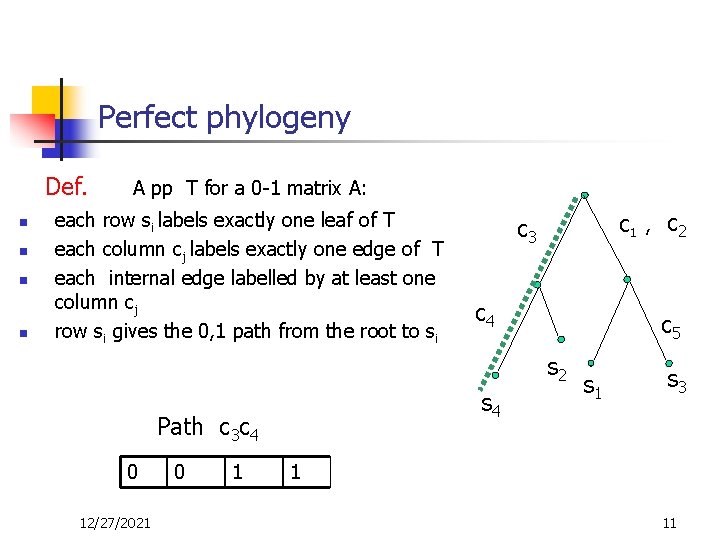
Perfect phylogeny Def. n n A pp T for a 0 -1 matrix A: each row si labels exactly one leaf of T each column cj labels exactly one edge of T each internal edge labelled by at least one column cj row si gives the 0, 1 path from the root to si c 3 C 1 , c 4 c 5 s 2 s 4 Path c 3 c 4 0 12/27/2021 0 1 c 2 s 1 s 3 1 11
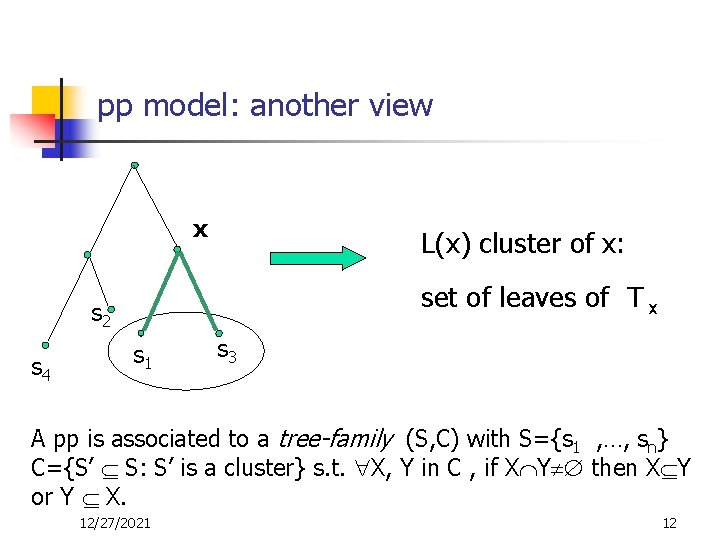
pp model: another view x L(x) cluster of x: set of leaves of T x s 2 s 4 s 1 s 3 A pp is associated to a tree-family (S, C) with S={s 1 , …, sn} C={S’ S: S’ is a cluster} s. t. X, Y in C , if X Y then X Y or Y X. 12/27/2021 12
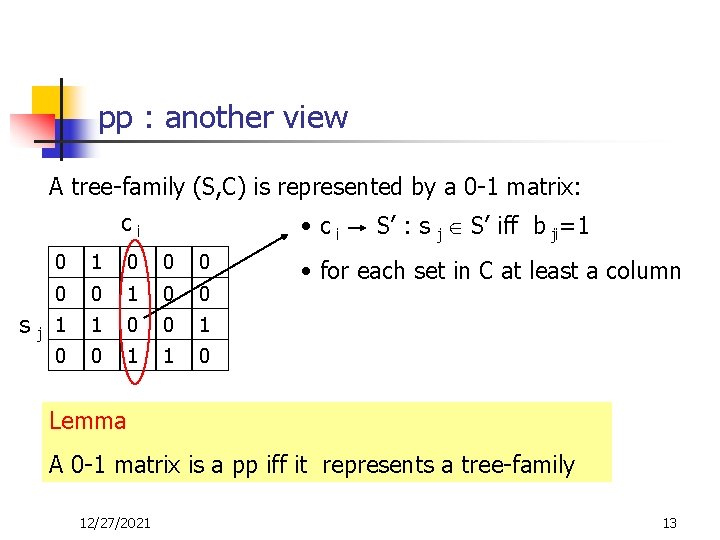
pp : another view A tree-family (S, C) is represented by a 0 -1 matrix: ci • ci S’ : s j S’ iff b ji=1 s j 0 1 0 0 1 1 0 • for each set in C at least a column Lemma A 0 -1 matrix is a pp iff it represents a tree-family 12/27/2021 13
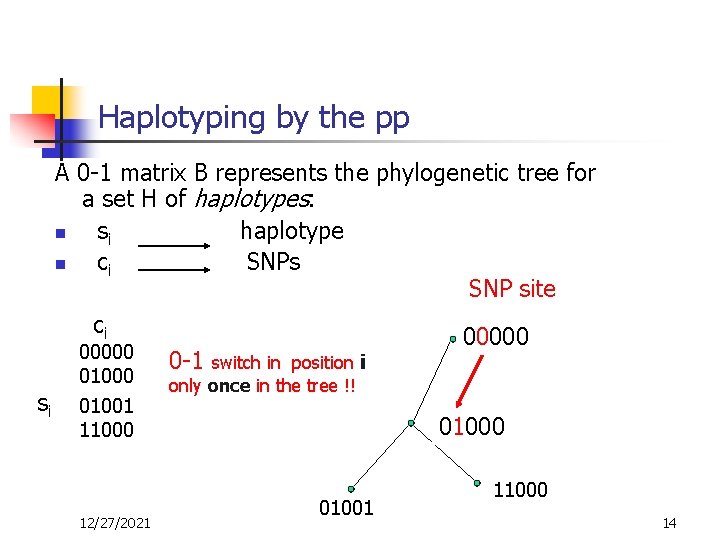
Haplotyping by the pp A 0 -1 matrix B represents the phylogenetic tree for a set H of haplotypes: n si haplotype n ci SNPs SNP site ci 00000 0 -1 switch in position i si 01000 01001 11000 12/27/2021 only once in the tree !! 01000 01001 11000 14

Haplotyping and the pp: observations n n The root of T may not be the haplotype 000000 0 -1 switch or 1 -0 switch (directed case) 00011 00000 00011 0 -1 switch 1 -0 01000 11000 01100 12/27/2021 01001 01010 01001 11010 15
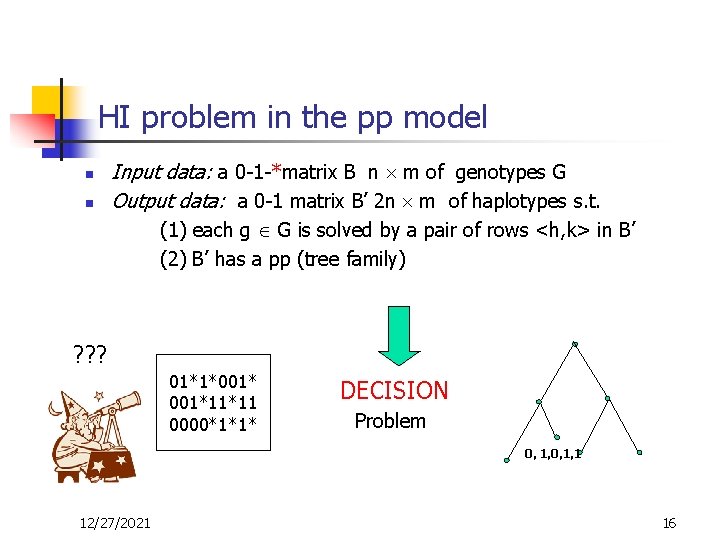
HI problem in the pp model n n Input data: a 0 -1 -*matrix B n m of genotypes G Output data: a 0 -1 matrix B’ 2 n m of haplotypes s. t. (1) each g G is solved by a pair of rows <h, k> in B’ (2) B’ has a pp (tree family) ? ? ? 01*1*001*11*11 0000*1*1* DECISION Problem 0, 1, 1 12/27/2021 16
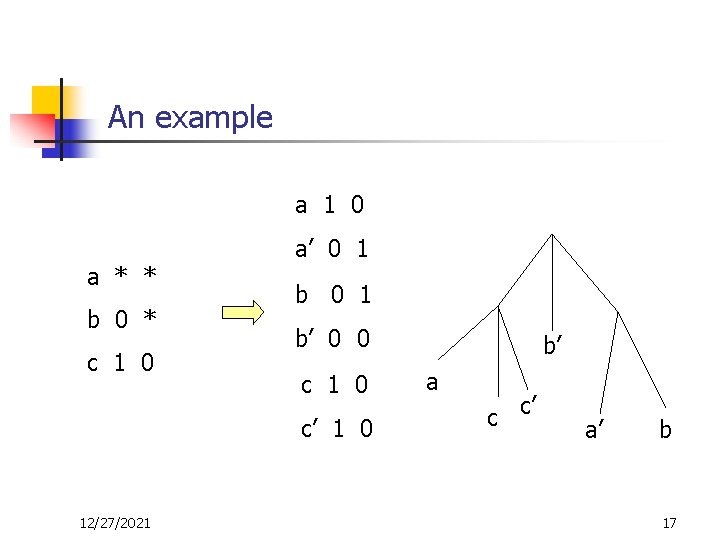
An example a 1 0 a * * b 0 * c 1 0 a’ 0 1 b’ 0 0 c 1 0 c’ 1 0 12/27/2021 b’ a c c’ a’ b 17

The pph problem: solutions n n n An undirected algorithm Gusfield Recomb 2002 An O(nm 2)- algorithm Karp et al. Recomb 2003 A linear time O(nm) algorithm ? ? Optimal algorithm A related problem: the incomplete directed pp (IDP) Inferring a pp from a 0 -1 -* matrix O(nm + klog 2(n+ m)) algorithm Peer, T. Pupko, R. Shamir, R. Sharan SIAM 2004 12/27/2021 18
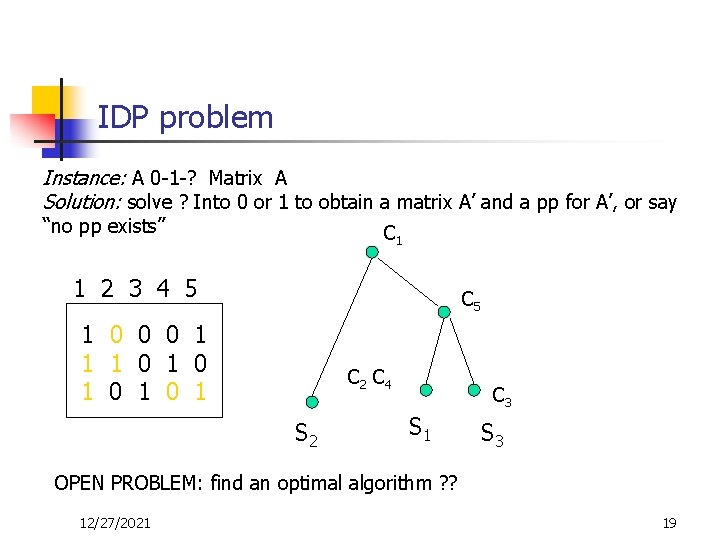
IDP problem Instance: A 0 -1 -? Matrix A Solution: solve ? Into 0 or 1 to obtain a matrix A’ and a pp for A’, or say “no pp exists” C 1 1 2 3 4 5 C 5 1 ? 0 0 0 1 1 ? ? 1 00 11 00 ? 0 1 ? 0 ? 1 1 C 2 C 4 S 2 C 3 S 1 S 3 OPEN PROBLEM: find an optimal algorithm ? ? 12/27/2021 19
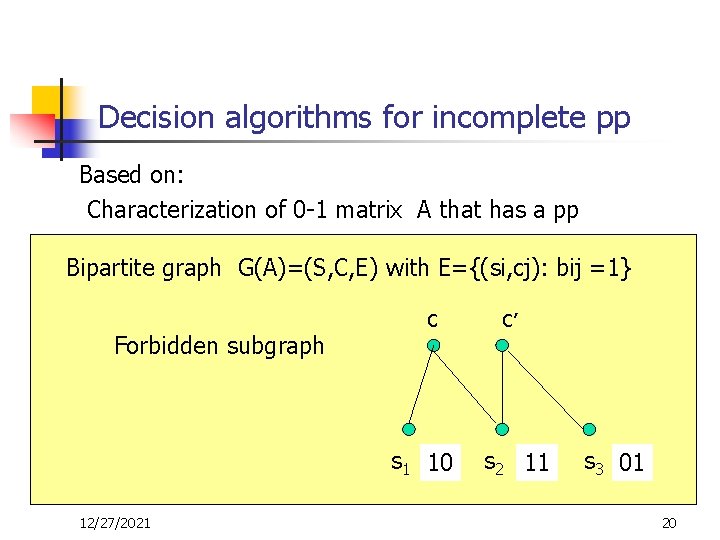
Decision algorithms for incomplete pp Based on: Characterization of 0 -1 matrix A that has a pp Bipartite graph G(A)=(S, C, E) with E={(si, cj): bij =1} -Tree family Forbidden subgraph 00 Y 01 11 12/27/2021 X 10 - forbidden submatrix – C’ c give a no certificate s 1 10 s 2 10 11 110 1 s 3 01 20

Test: a 0 -1 matrix A has a pp? O(nm) algorithm (Gusfield 1991) Steps: 1. Given A order {c 1, …, cm} as (decreasing) binary numbers A’ 2. Let L(i, j)=k , k = max{l <j: A’[i, l]=1} 3. Let index(j) = max{L(i, j): i} 4. Then apply th. n TH. A’ has a pp iff L(i, j) = index(j) for each (i, j) s. t. A’[i, j]=1 12/27/2021 21

Idea: 12/27/2021 22
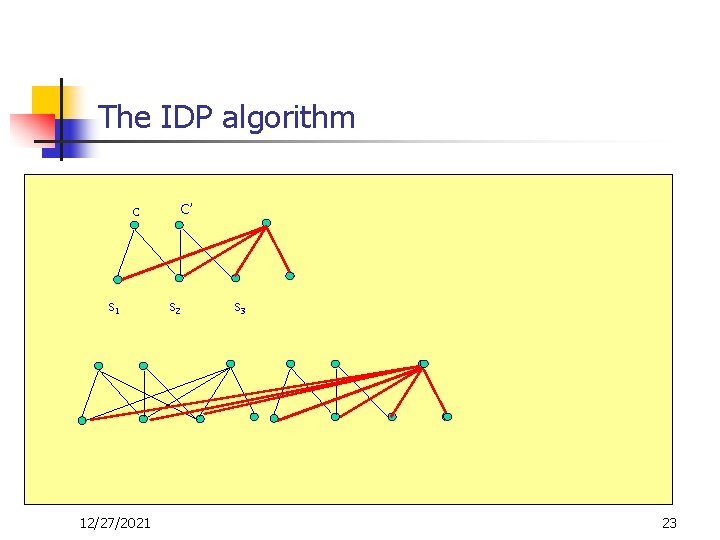
The IDP algorithm c s 1 12/27/2021 C’ s 2 s 3 23

Other HI problems via the pp model Incomplete 0 -1 -*-? matrix because of missing data: haplotypes pp (Ihpp) haplotype rows genotype pp (Igpp) genotype rows n Algorithms: n Ihpp = IDP given a row as a root (polynomial time) NP-complete otherwise n Igpp has polynomial solution under rich data hypothesis (Karp et al. Recomb 2004 – Icalp 2004 ) NP-complete otherwise 12/27/2021 24
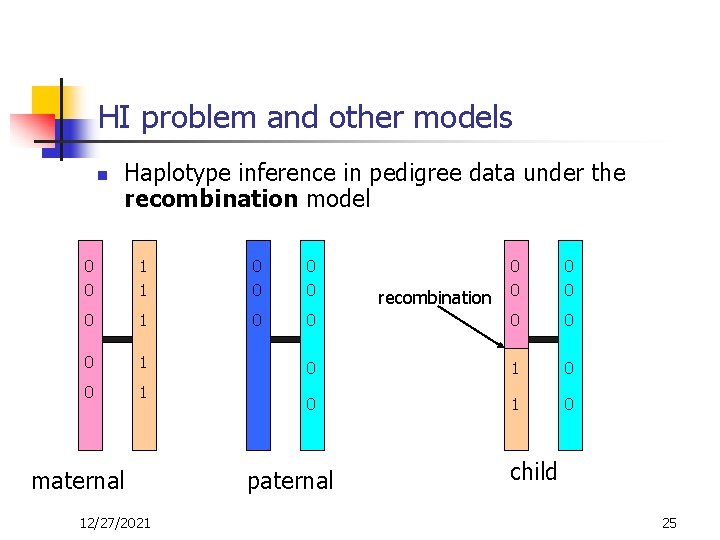
HI problem and other models n Haplotype inference in pedigree data under the recombination model 0 0 1 1 0 0 0 1 0 1 maternal 12/27/2021 0 0 0 0 1 0 paternal recombination child 25
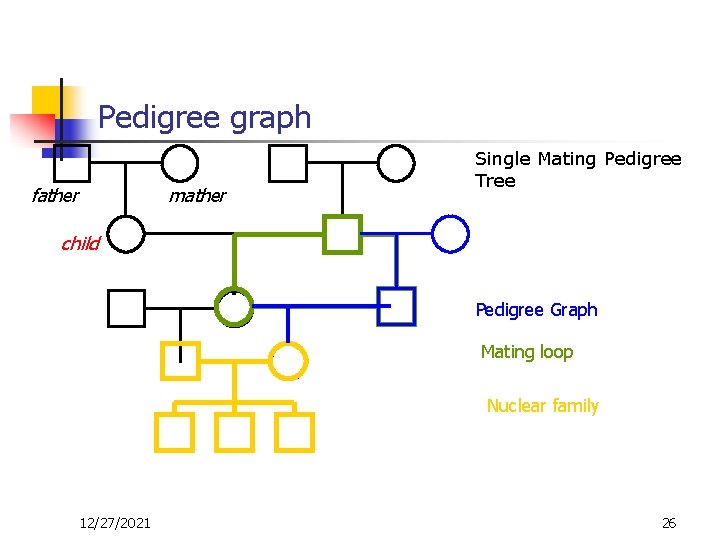
Pedigree graph father mather Single Mating Pedigree Tree child Pedigree Graph Mating loop Nuclear family 12/27/2021 26
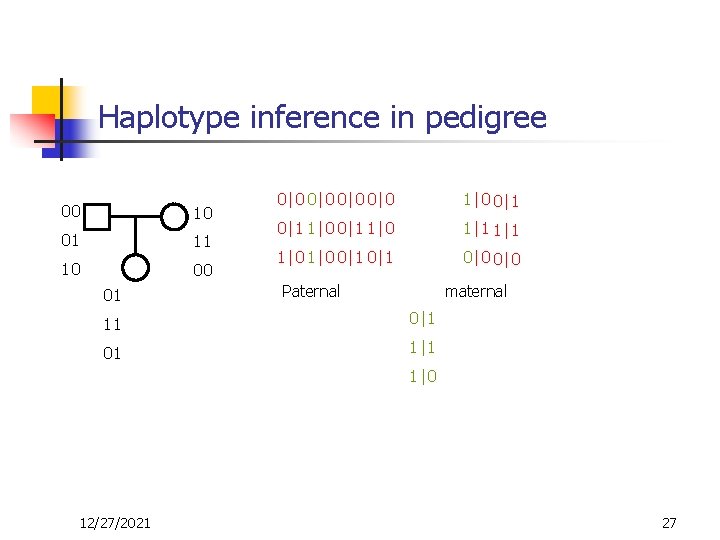
Haplotype inference in pedigree 00 10 01 11 10 00 01 0|0 0|0 0|0 1 1|0 0|1 1|0 1 1|1 1|0 0|1 0 0|0 Paternal maternal 11 0|1 01 01 1|1 11 1|0 10 12/27/2021 27
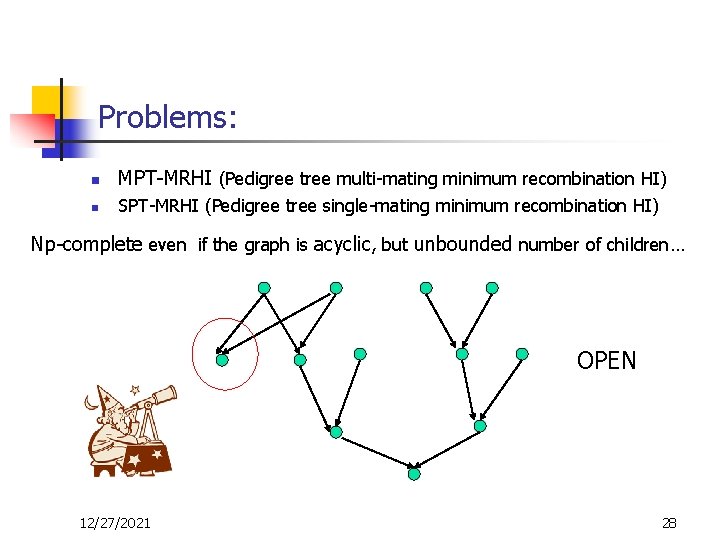
Problems: n MPT-MRHI (Pedigree tree multi-mating minimum recombination HI) n SPT-MRHI (Pedigree tree single-mating minimum recombination HI) Np-complete even if the graph is acyclic, but unbounded number of children… OPEN 12/27/2021 28

Conclusions 12/27/2021 29

References 12/27/2021 30
- Slides: 30