Solving Haplotyping Inference Parsimony problem using a polynomial

Solving Haplotyping Inference Parsimony problem using a polynomial class representative formulation and a set covering formulation Martine Labbé Alessandra Godi Université Libre de Bruxelles IASI (CNR) Roma Airo Winter 2007 - Cortina d’Ampezzo, February 5 th -9 th, 2007
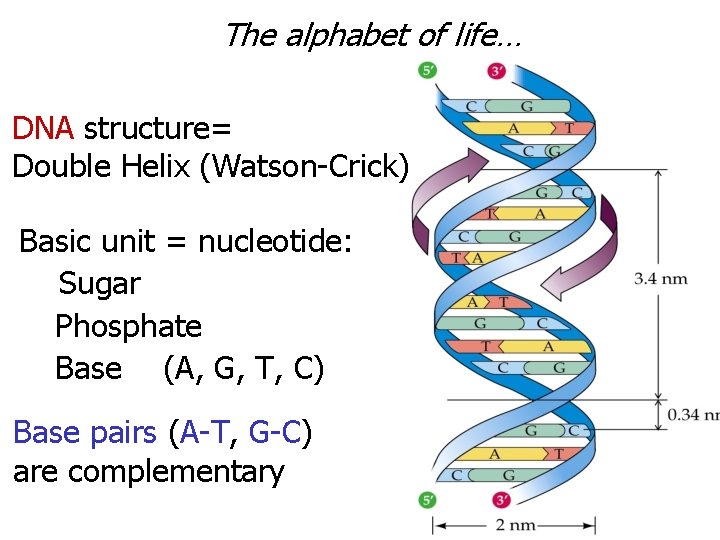
The alphabet of life… DNA structure= Double Helix (Watson-Crick) Basic unit = nucleotide: Sugar Phosphate Base (A, G, T, C) Base pairs (A-T, G-C) are complementary
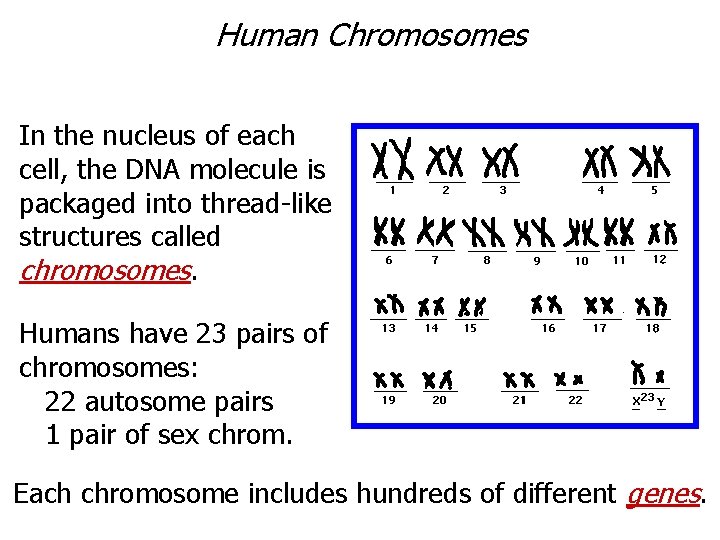
Human Chromosomes In the nucleus of each cell, the DNA molecule is packaged into thread-like structures called chromosomes. Humans have 23 pairs of chromosomes: 22 autosome pairs 1 pair of sex chrom. Each chromosome includes hundreds of different genes.
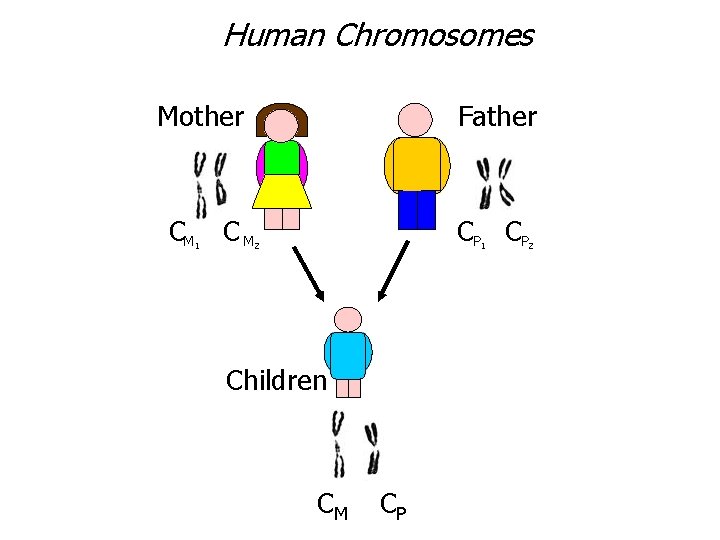
Human Chromosomes Mother CM C M 1 Father CP CP 2 1 Children CM CP 2
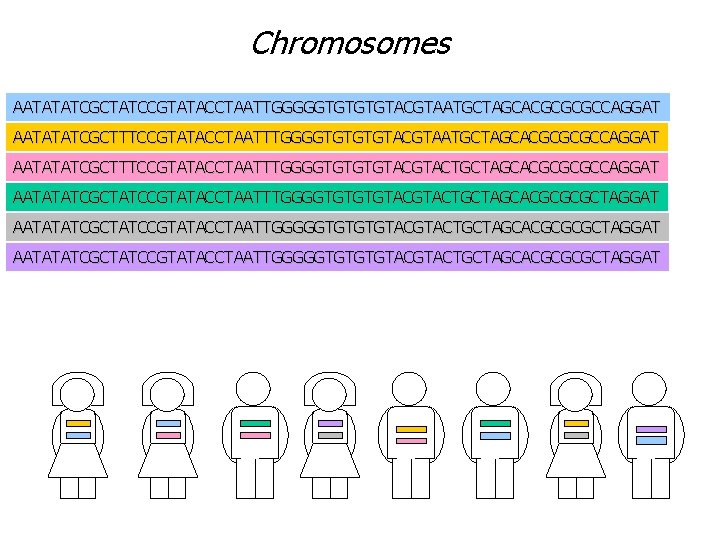
Chromosomes AATATATCGCTATCCGTATACCTAATTGGGGGTGTGTGTACGTAATGCTAGCACGCGCGCCAGGAT AATATATCGCTTTCCGTATACCTAATTTGGGGTGTACGTACTGCTAGCACGCGCGCCAGGAT AATATATCGCTATCCGTATACCTAATTTGGGGTGTGTGTACGTACTGCTAGCACGCGCGCTAGGAT AATATATCGCTATCCGTATACCTAATTGGGGGTGTACGTACTGCTAGCACGCGCGCTAGGAT

Chromosomes A single ‘copy’ of a chromosome is called haplotype, while a description of the mixed data on the two ‘copies’ is called genotype. For disease association studies, haplotype data is more valuable than genotype data, but haplotype data is hard to collect. Genotype data is easy to collect.

SNPs All humans are 99, 99 % identical. Diversity? polymorphism. A SNP is a Single Nucleotide Polymorphism - a site in the genome where two different nucleotides appear with sufficient frequency in the population (say each with 5% frequency or more).
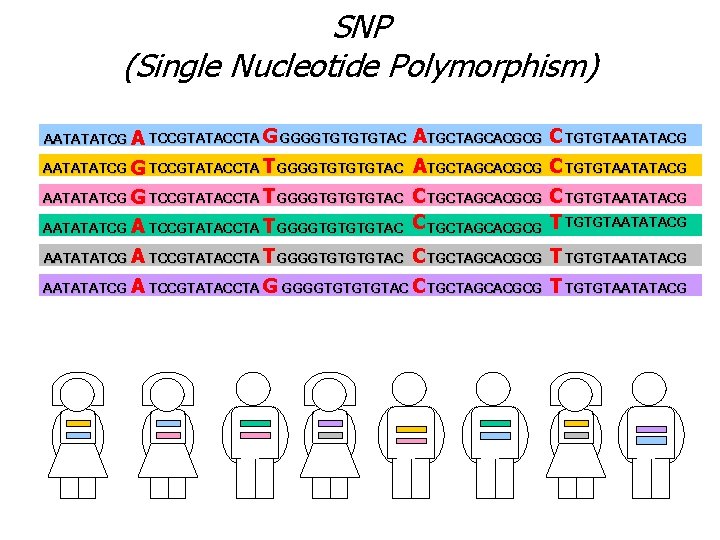
SNP (Single Nucleotide Polymorphism) AATATATCG A TCCGTATACCTA G GGGGTGTAC A TGCTAGCACGCG C TGTGTAATATACG AATATATCG G TCCGTATACCTA T GGGGTGTAC A TGCTAGCACGCG C TGTGTAATATACG C TGCTAGCACGCG T TGTGTAATATACG AATATATCG G TCCGTATACCTA T GGGGTGTGTGTAC AATATATCG A TCCGTATACCTA T GGGGTGTAC C TGCTAGCACGCG T TGTGTAATATACG AATATATCG A TCCGTATACCTA G GGGGTGTAC C TGCTAGCACGCG T TGTGTAATATACG
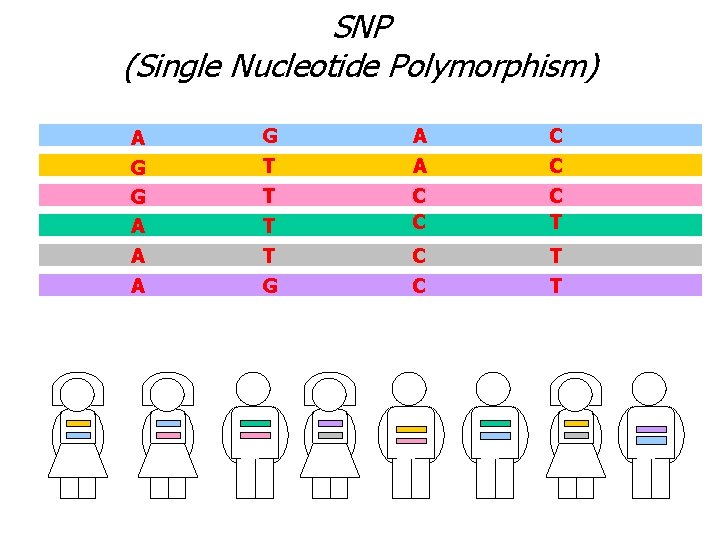
SNP (Single Nucleotide Polymorphism) A G A C G G A A A T T G A C C T T
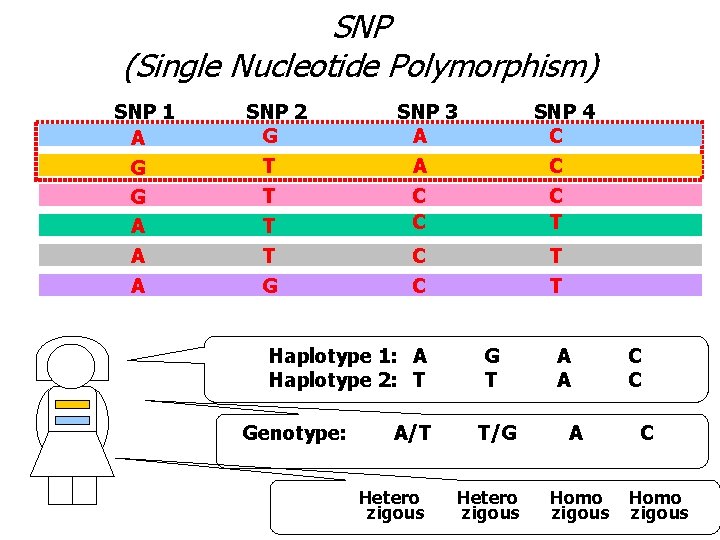
SNP (Single Nucleotide Polymorphism) SNP 1 A G G A A A SNP 2 G T T G SNP 3 A A C C T T Haplotype 1: A Haplotype 2: T Genotype: SNP 4 C A/T Hetero zigous G T T/G Hetero zigous A A A Homo zigous C C C Homo zigous
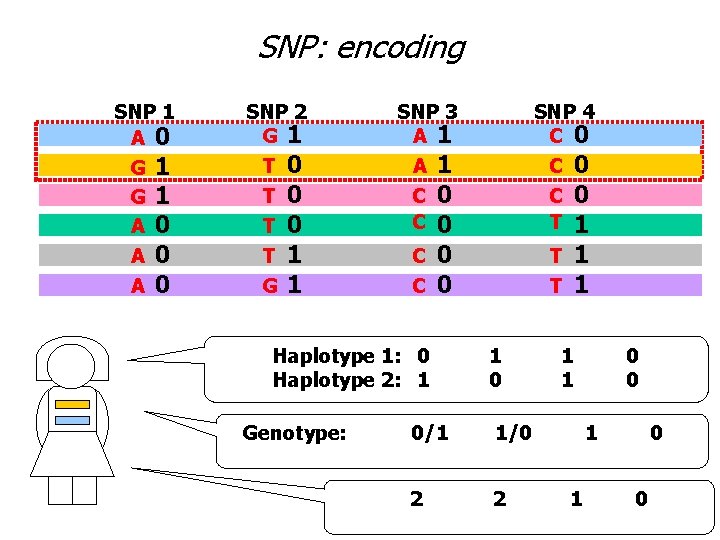
SNP: encoding SNP 1 A 0 G G A A A 1 1 0 0 0 SNP 2 G 1 T T G 0 0 0 1 1 SNP 3 A 1 1 0 0 C 0 0 0 1 T 1 A C C Haplotype 1: 0 Haplotype 2: 1 Genotype: SNP 4 C 0 C C T 1 0 0/1 1/0 2 2 1 1 0 0

Haplotyping of a population Given a set of genotypes G (strings on {0, 1, 2}n alphabet), find a set of “generating” haplotypes H (strings on {0, 1}n alphabet). genotype individual
The GENOME is the set of genetic information which lies in the DNA sequence of each living organism. The DNA sequence is a linear disposition of 4 different molecule, nucleotide, or bases: A, T, C, G. The bases are paired each other by hydrogen bonds.

The DNA implies differences between the individuals of the same species. What makes us different from each other is called polymorphism.
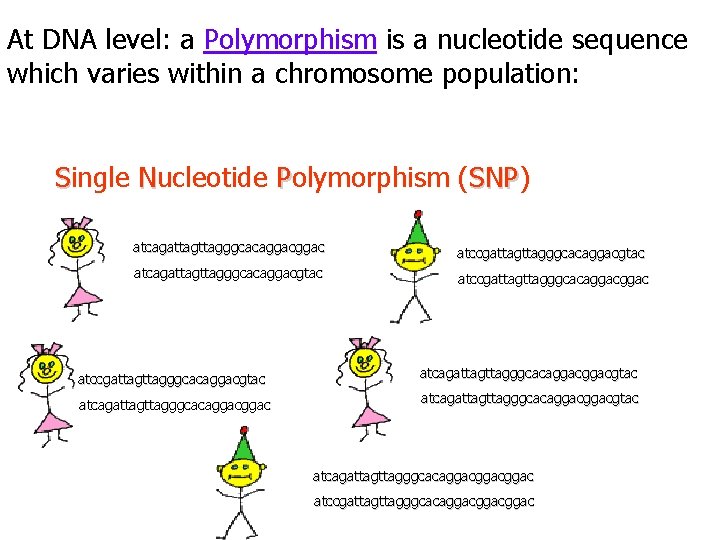
At DNA level: a Polymorphism is a nucleotide sequence which varies within a chromosome population: Single Nucleotide Polymorphism (SNP) SNP atcagattagggcacaggac atccgattagggcacaggacgtac atcagattagggcacaggacgtac atccgattagggcacaggacgtac atcagattagttagggcacaggacggac atcagattagggcacaggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
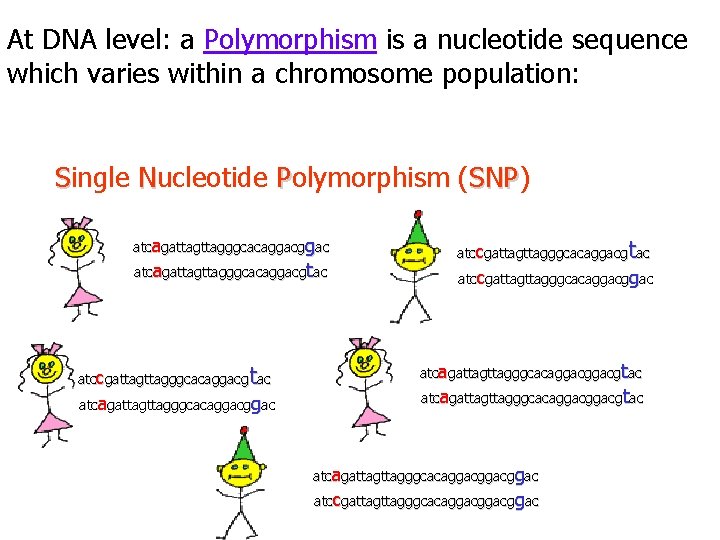
At DNA level: a Polymorphism is a nucleotide sequence which varies within a chromosome population: Single Nucleotide Polymorphism (SNP) SNP atcagattagggcacaggacgtac t atccgattagggcacaggacg ac atcagattagggcacaggac t atccgattagggcacaggacg ac atccgattagggcacaggac atcagattagttagggcacaggacggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
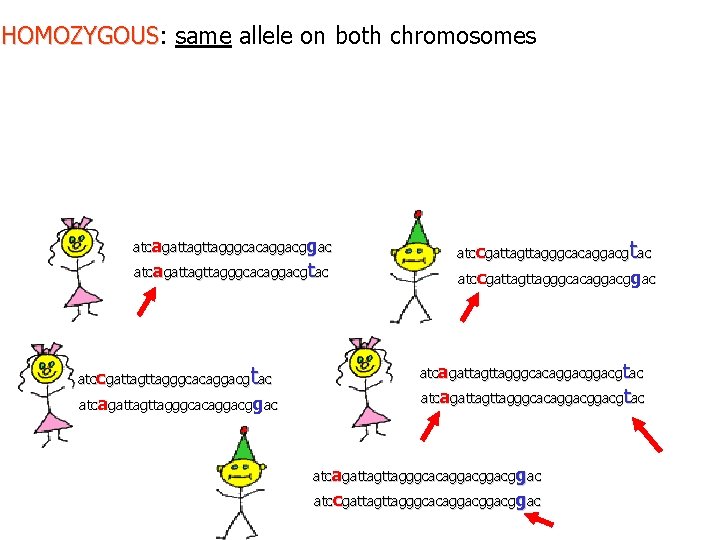
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS atcagattagggcacaggacgtac t atccgattagggcacaggacg ac atcagattagggcacaggac t atccgattagggcacaggacg ac atccgattagggcacaggac atcagattagttagggcacaggacggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS ETEROZYGOUS: different alleles ETEROZYGOUS atcagattagggcacaggacgtac t atccgattagggcacaggacg ac atcagattagggcacaggac t atccgattagggcacaggacg ac atccgattagggcacaggac atcagattagttagggcacaggacggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS ETEROZYGOUS: different alleles ETEROZYGOUS atcagattagggcacaggacgtac t atccgattagggcacaggacg ac atcagattagggcacaggac t atccgattagggcacaggacg ac atccgattagggcacaggac atcagattagttagggcacaggacggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS ETEROZYGOUS: different alleles ETEROZYGOUS HAPLOTYPES: chromosome at SNP level HAPLOTYPES atcagattagggcacaggacgtac t atccgattagggcacaggacg ac atcagattagggcacaggac t atccgattagggcacaggacg ac atccgattagggcacaggac atcagattagttagggcacaggacggacgtac atcagattagggcacaggacggac atccgattagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS ETEROZYGOUS: different alleles ETEROZYGOUS HAPLOTYPES: chromosome at SNP level HAPLOTYPES a a c a g t c c t g t a a t g a c t g g
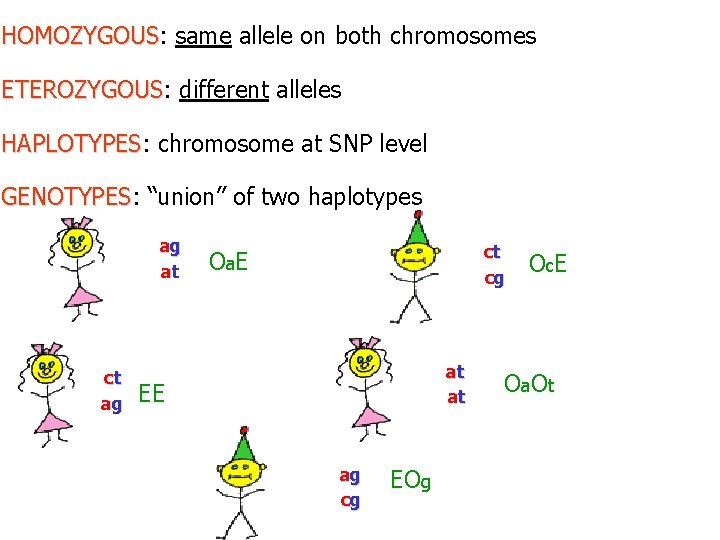
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS ETEROZYGOUS: different alleles ETEROZYGOUS HAPLOTYPES: chromosome at SNP level HAPLOTYPES GENOTYPES: “union” of two haplotypes GENOTYPES ag at ct O c. E cg O a E at at ct ag EE ag cg EOg O a O t
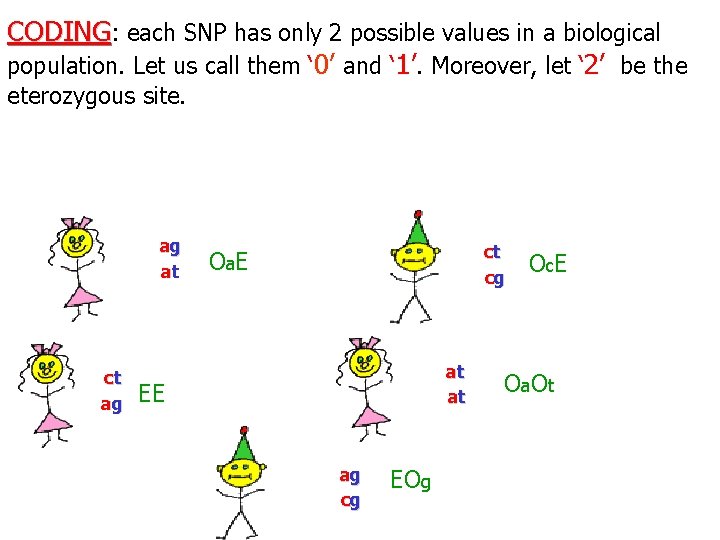
CODING: each SNP has only 2 possible values in a biological population. Let us call them ‘ 0’ and ‘ 1’. Moreover, let ‘ 2’ be the eterozygous site. ag at ct O c. E cg O a E at at ct ag EE ag cg EOg O a O t
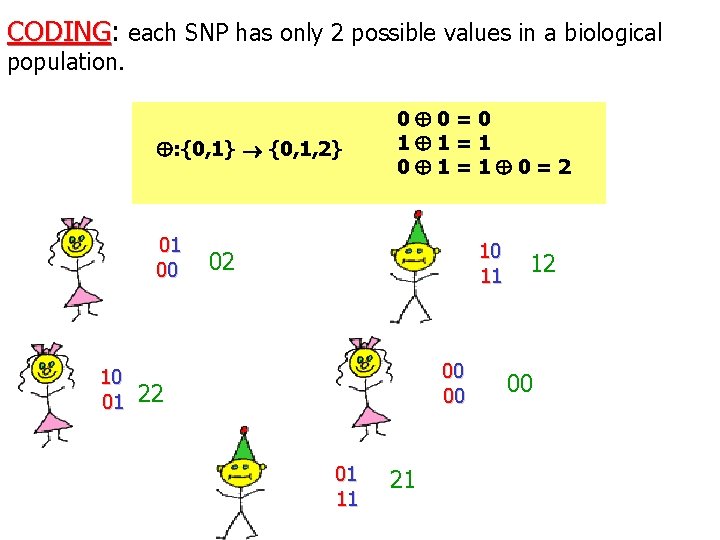
CODING: CODING each SNP has only 2 possible values in a biological population. : {0, 1} {0, 1, 2} 01 00 0 0=0 1 1=1 0=2 10 02 11 00 10 01 22 00 01 11 21 12 00
HAPLOTYPING of a population Given a set G (strings in {0, 1, 2}n), find a set of generator haplotypes H (strings in {0, 1}n) genotype individual

HAPLOTYPING of a population: State of the Art Perfect Phylogeny (Bafna, Gusfield, Yooseph 02) Estimation of haplotype frequencies (probabilistic studies: Fallin – Shork, 00) Parsimony Objective (Gusfield 02, Brown 05)

HAPLOTYPING of a population: Parsimony Objective (NP-hard) Combinatorial Methods (Gusfield 2002, Brown 2004, LANCIA –Rizzi, 2002): Exponential and Polynomial ILP formulations Rule-based methods (HAPINFER - Clark 1990): Starting from genotypes, haplotypes are inferred Statistical methods (PHASE- Stephens 2004, HAPLOTYPER – Niu 2001, GERBIL – Shamir 2005)
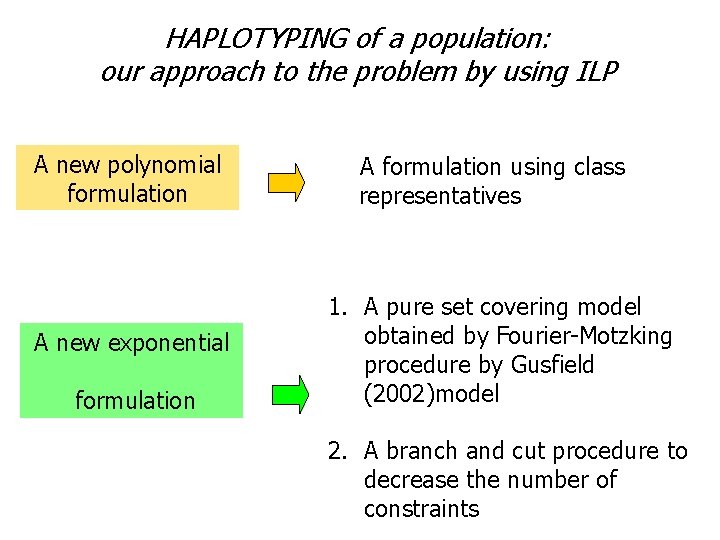
HAPLOTYPING of a population: our approach to the problem by using ILP A new polynomial formulation A formulation using class representatives A new exponential 1. A pure set covering model obtained by Fourier-Motzking procedure by Gusfield (2002)model formulation 2. A branch and cut procedure to decrease the number of constraints

A new polynomial formulation Main idea: class representatives G={g 1, g 2, …, gm} genotypes of length n I={h 1, …, hq} a solution of the problem Each haplotype induces a subset of ordinated genotypes, and each geno belongs to exactly two of these subsets: h 1 {gi, gj, gk, …} = Si h 2 {gi, gl, gr, gs…} = Si’ h 3 {gk, gl, gs, gt…} = Sk …. The smallest index geno identifies the subset; the prime appears if the correspondent index has been already used. …. K = {1, 2, …, m} K’ = {1’, 2’, …, m’}

A new polynomial formulation VARIABLES 1 yk{i, j}= k K i, j K K’ If geno gk belongs to two subset of geno’s, one having a geno with smallest index equal to i and the other one having the geno with smallest index j 0 Otherwise

A new polynomial formulation Ex: g 1= 021, g 2= 002, g 3 = 012 h 1 = 001 {g 1, g 2} = S 1 h 2 = 011 {g 1, g 3} = S 1’ y 1{1, 1’} = 1 Let us note that some y variables do not exist: y 2{1’, 2’} = 0 If y 2{1’, 2’} = 1 S 1={g 1, …. } S 1’={g 1, g 2…. } S 2={g 2, …} S 2’={g 2, …} Absurd!!!

A new polynomial formulation VARIABLES xi = i K K’ 1 If there exists a subset of geno’s of the solution having geno i as geno with smallest index 0 1 zi, p = i K K’ p SNP 0 Otherwise It is the value of the p-th coordinate of the haplo explaining the subset of geno’s used in the solution and having geno i as geno with smallest index OBJECTIVE FUNCTION: min xi i K K’

A new polynomial formulation CONSTRAINTS: 1. xi’ i K, i K’ 2. yk{i, j} 1 k K i, j K K’, i≤k, j≤k

A new polynomial formulation CONSTRAINTS: yk{i, j} + yk{i, j} ≤ xi 3. j K K’, j≥i 3 a. j K K’, j<i yk{k, k’} ≤ xk’ k K i K K’, k K i = k’

A new polynomial formulation CONSTRAINTS: 4 a. 4 b. 4 c. zi, p= 0 i K K’ p SNP s. t. gi(p)=0 zi, p= 1 i K K’ p SNP s. t. gi(p)=1 zi, p + zj, p = 1 {i, j} K K’ p SNP s. t. gi(p)=2

A new polynomial formulation CONSTRAINTS: 5. zi, p ≤ 1 - yk{i, j} xi j K K’, j≥i 5 a. yk{k, k’} + zk’, p ≤ 1 j K K’, j<i k K p SNP : gk(p)=0 i K K’ k K, i = k’ p SNP : gk(p)=0

A new polynomial formulation CONSTRAINTS: 6. zi, p ≥ yk{i, j} + yk{i, j} j K K’, j≥i 6 a. zk’, p ≥ yk{k, k’} j K K’, j<i k K p SNP : gk(p)=1 i K K’ k K, i = k’ p SNP : gk(p)=1

A new polynomial formulation CONSTRAINTS: 7. 7 a. zi, p + zj, p ≥ yk{i, j} zi, p + zj, p ≤ 2 - yk{i, j} k K p SNP : gk(p)=2 i, j K K’
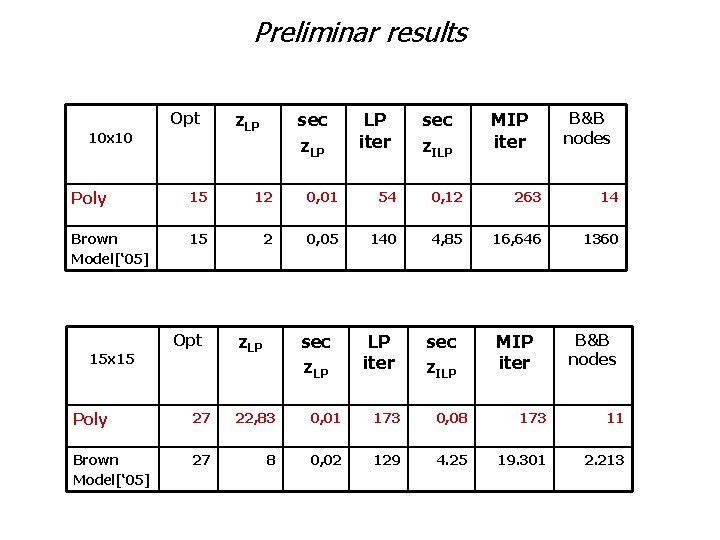
Preliminar results Opt 10 x 10 z. LP sec z. LP LP iter sec z. ILP MIP iter B&B nodes Poly 15 12 0, 01 54 0, 12 263 14 Brown Model[‘ 05] 15 2 0, 05 140 4, 85 16, 646 1360 Opt 15 x 15 z. LP sec z. LP LP iter sec z. ILP MIP iter B&B nodes Poly 27 22, 83 0, 01 173 0, 08 173 11 Brown Model[‘ 05] 27 8 0, 02 129 4. 25 19. 301 2. 213
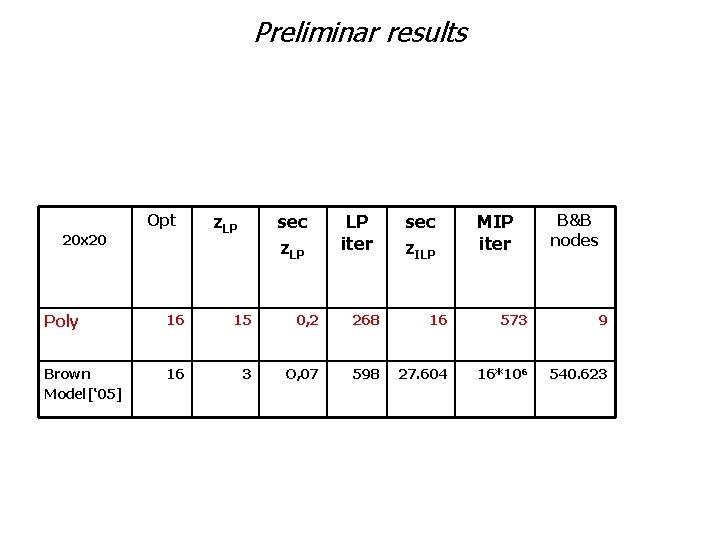
Preliminar results Opt 20 x 20 z. LP sec z. LP LP iter sec z. ILP MIP iter B&B nodes Poly 16 15 0, 2 268 16 573 9 Brown Model[‘ 05] 16 3 O, 07 598 27. 604 16*106 540. 623

From Gusfield’s formulation (2002)… ^ Let G be the genotype set and H the set of haplotypes which are compatible with some genotype in G. For each g G ^ Pg = {(h 1, h 2) con h 1, h 2 H | h 1 h 2 = g} INTEGER VARIABLES Xh 1 if h is chosen 0 otherwise y 1 if (h 1, h 2) is 2 selected h 1, h 2 0 otherwise

From Gusfield’s formulation (2002)… OBJECTIVE FUNCTION min Xh ^ h H CONSTRAINTS 1. y h , h 1 (h 1, h 2) Pg 1 g G 2 2. y X h 1, h 2 (h 1, h 2) Pg , g G 3. y X h 2 h 1, h 2 (h 1, h 2) Pg , g G
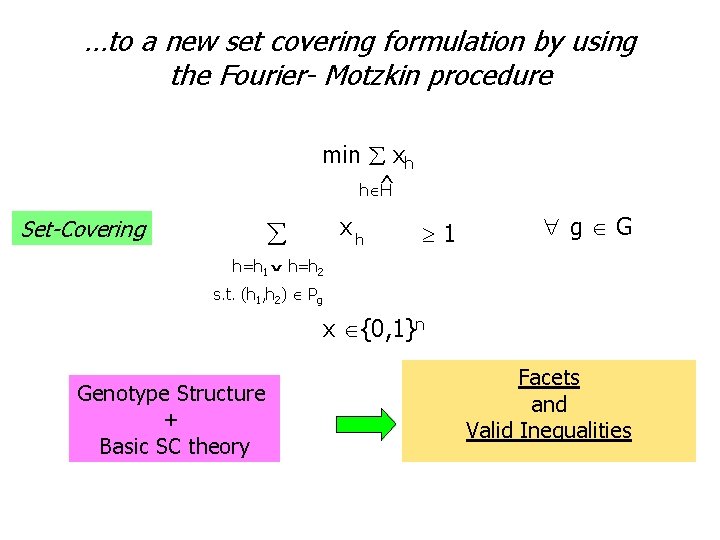
…to a new set covering formulation by using the Fourier- Motzkin procedure min xh ^ h H Set-Covering xh 1 g G h=h 1 h=h 2 ˇ s. t. (h 1, h 2) Pg x {0, 1}n Genotype Structure + Basic SC theory Facets and Valid Inequalities
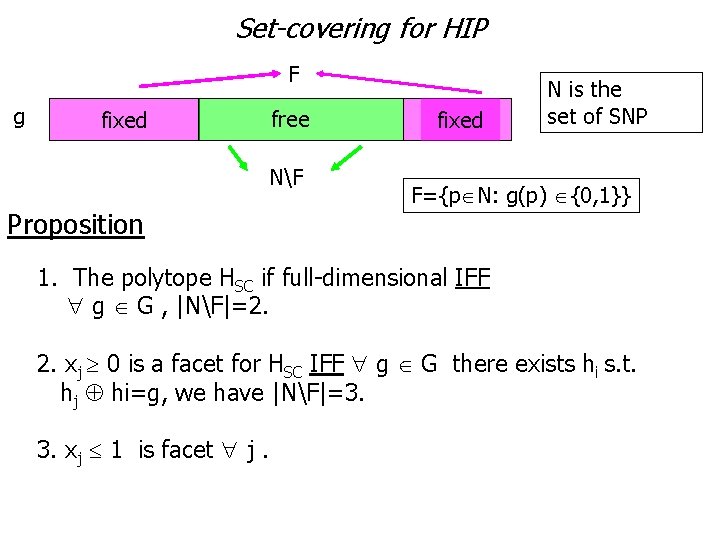
Set-covering for HIP F g free fixed NF Proposition fixed N is the set of SNP F={p N: g(p) {0, 1}} 1. The polytope HSC if full-dimensional IFF g G , |NF|=2. 2. xj 0 is a facet for HSC IFF g G there exists hi s. t. hj hi=g, we have |NF|=3. 3. xj 1 is facet j.
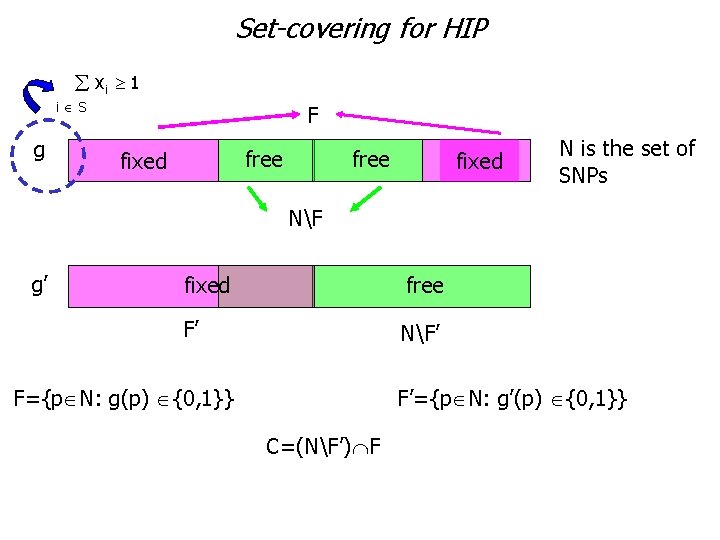
Set-covering for HIP xi 1 i S g F free fixed N is the set of SNPs NF g’ fixed free F’ NF’ F={p N: g(p) {0, 1}} F’={p N: g’(p) {0, 1}} C=(NF’) F

Set-covering for HIP Theorem Let us consider a genotype g and a subset S of haplotypes which are associated to a minimal set covering inequality: xh 1. h S This inequality is facet defining IFF for each genotype g’ g one of the following conditions holds: |C|=|(NF’) F| 3 |C|=|(NF’) F|= 2 e (NF) (NF’)

Set-covering for HIP NOTE: For the following cases: 1 st case: If |C|=|(NF’) F|= 2 (NF) (NF’) = 2 nd case : If |C|= |{p}|=1 3 rd case : If C= the set covering inequality is dominated by another one that can be defined by using a SEQUENTIAL LIFTING procedure.

Set-covering for HIP: main idea To overcome the exponential structure of the formulation: 1. Add only set-covering inequalities which are facet-defining 2. Add them in branch and cut procedure

Set-covering for HIP: a branch and cut procedure x* a fractional solution of a subproblem of the original one g: (h 1, h 2 ) (h 3, h 4) (h 5, h 6) (h 7, h 8) All set covering inequalities associated with g have the following structure: x{1 or 2}+ x{3 or 4} + x{5 or 6}+ x{7 or 8} ≥ 1

Set-covering for HIP: a branch and cut procedure We want to find a set covering inequality of g that violates x* min {x*1, x*2} + min {x*3, x*4} + min {x*5, x*6} + min {x*7, x*8} < 1 If it esists, we have found a set covering inequality which cut off x* !!! We choose to add it to the system only if it is facet-defining.
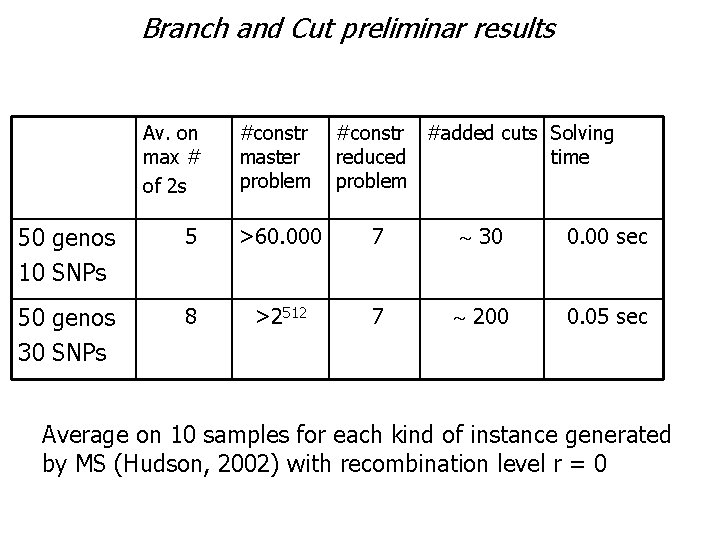
Branch and Cut preliminar results Av. on max # of 2 s #constr master problem #constr #added cuts Solving reduced time problem 50 genos 10 SNPs 5 >60. 000 7 30 0. 00 sec 50 genos 30 SNPs 8 >2512 7 200 0. 05 sec Average on 10 samples for each kind of instance generated by MS (Hudson, 2002) with recombination level r = 0

Future Works On Polynomial formulation: 1. Strengthening of the model by Clique inequalities on genotype conflict graph 2. Cplex Concert Technologies 3. More test vs other polynomial formuations On Exponential formulation: 1. Implementation of Lifting Procedure 2. More test in comparison with Gusfield formulation
- Slides: 52