Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2

Module 5 Small variant calling & annotation Guillaume Bourque Informatics on High-throughput Sequencing Data June 10 -11, 2015

Learning Objectives of Module • • Have an overview of the variant calling analysis workflow Understand the basic principles of variant calling Know what can improve the variant calls Learn how to filter and annotate variants Be able to call and annotate small variants Learn about the vcf format Visualize SNPs and indels in IGV Module 5: Small variant calling & annotation bioinformatics. ca
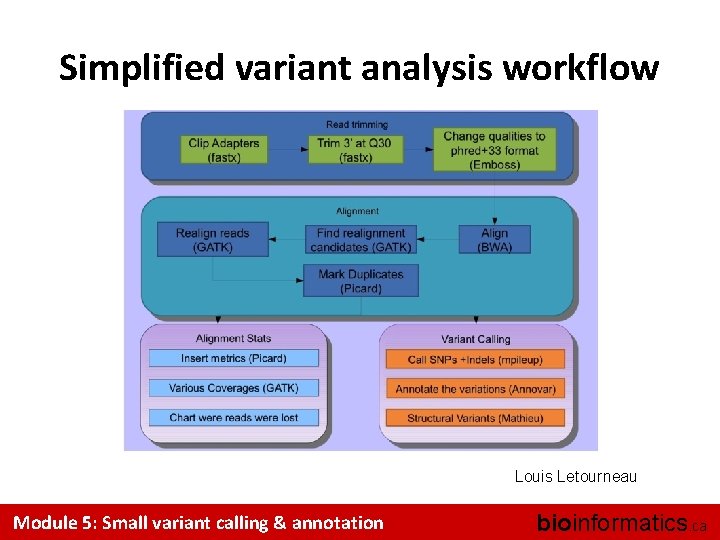
Simplified variant analysis workflow Louis Letourneau Module 5: Small variant calling & annotation bioinformatics. ca

Main analysis steps – – Quality control Pre-processing (trimming, remove adapters, …) Mapping (Module 2) Small variant calling (Module 5 – this module!) • SNP and indels calling • Variant filtering and annotation – Structural variant calling (Module 6) Module 5: Small variant calling & annotation bioinformatics. ca

Main analysis steps – – Quality control Pre-processing (trimming, remove adapters, …) Mapping (Module 2) Small variant calling (Module 5 – this module!) • SNP and indels calling • Variant filtering and annotation – Structural variant calling (Module 6) Module 5: Small variant calling & annotation bioinformatics. ca

Importance of quality control • Before you start an analysis, it’s very important to look at your raw data! • Are all of your samples sequenced using the same protocol and instruments? • Are there any technical issues affecting some of the samples? • This is especially important if you plan to compare different samples or different conditions Module 5: Small variant calling & annotation bioinformatics. ca
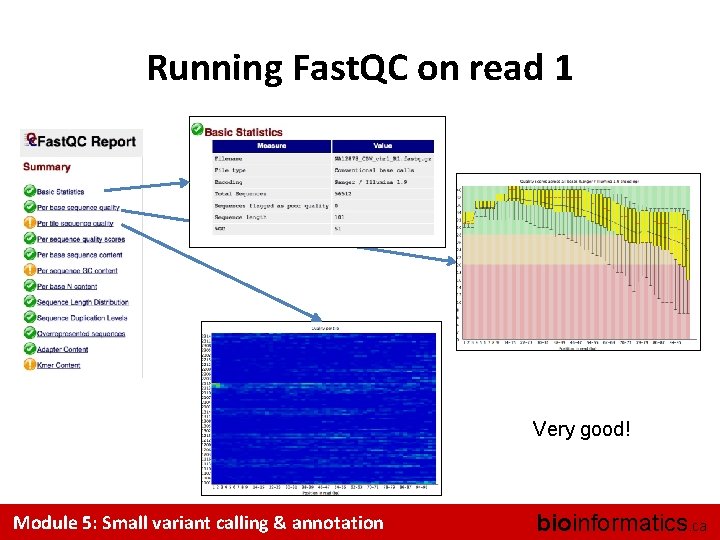
Running Fast. QC on read 1 Very good! Module 5: Small variant calling & annotation bioinformatics. ca

Running Fast. QC on read 2 Pretty good! Module 5: Small variant calling & annotation bioinformatics. ca

Adapters sequences in reads http: //www. illumina. com http: //srna-workbench. cmp. uea. ac. uk Module 5: Small variant calling & annotation 11 bioinformatics. ca
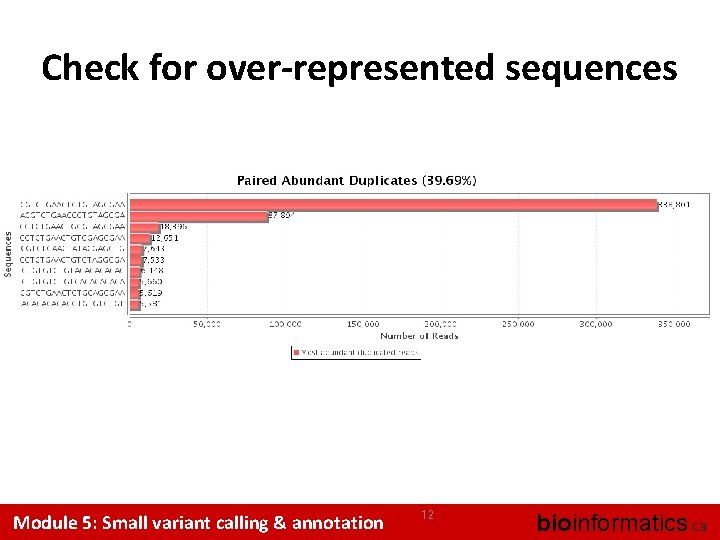
Check for over-represented sequences Module 5: Small variant calling & annotation 12 bioinformatics. ca

Read trimming tools For example, Trimmomatic performs a variety of useful trimming tasks for illumina paired-end and single ended data: – ILLUMINACLIP: Cut adapter and other illumina-specific sequences from the read. – SLIDINGWINDOW: Perform a sliding window trimming, cutting once the average quality within the window falls below a threshold. – LEADING: Cut bases off the start of a read, if below a threshold quality – TRAILING: Cut bases off the end of a read, if below a threshold quality – CROP: Cut the read to a specified length – HEADCROP: Cut the specified number of bases from the start of the read – MINLEN: Drop the read if it is below a specified length – TOPHRED 33: Convert quality scores to Phred-33 – TOPHRED 64: Convert quality scores to Phred-64 Module 5: Small variant calling & annotation 13 bioinformatics. ca

Main analysis steps – – Quality control Pre-processing (trimming, remove adapters, …) Mapping (Module 2) Small variant calling (Module 5 – this module!) • SNP and indels calling • Variant filtering and annotation – Structural variant calling (Module 6) Module 5: Small variant calling & annotation bioinformatics. ca

Goal Michael Strömberg Module 5: Small variant calling & annotation bioinformatics. ca

SNP Discovery: Goal Michael Strömberg sequencing errors Module 5: Small variant calling & annotation SNP bioinformatics. ca

Base quality QPhred = – 10 log 10 P (error) = 10 – Q / 10 Module 5: Small variant calling & annotation QPhred P (error) 10 10% 20 1% 30 0. 1% 40 0. 01% bioinformatics. ca
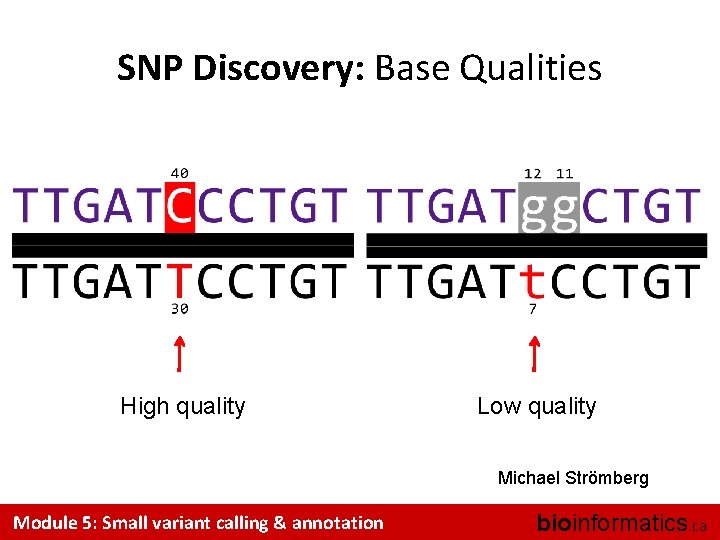
SNP Discovery: Base Qualities High quality Low quality Michael Strömberg Module 5: Small variant calling & annotation bioinformatics. ca

SNPs & Bayesian Statistics # of individuals base quality allele call in read Michael Strömberg Module 5: Small variant calling & annotation bioinformatics. ca

Strategies that improve variant calling • • Local realignment Duplicate marking Base quality recalibration Population structure and imputation Module 5: Small variant calling & annotation bioinformatics. ca

Strategies that improve variant calling • • Local realignment Duplicate marking Base quality recalibration Population structure and imputation Module 5: Small variant calling & annotation bioinformatics. ca
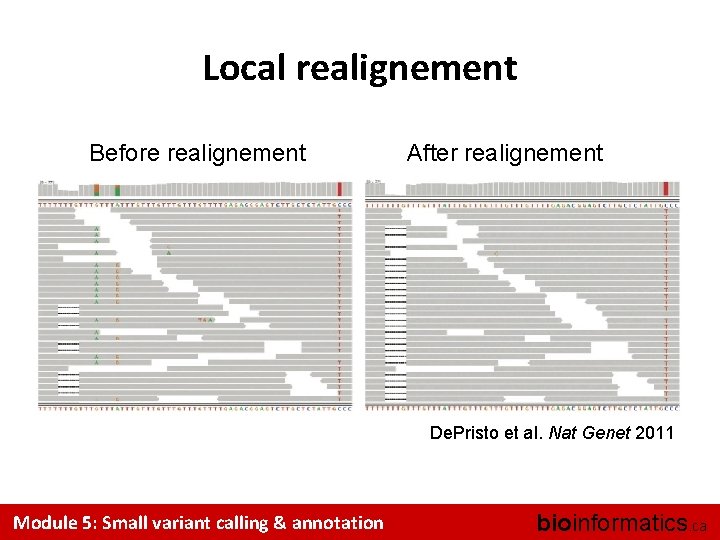
Local realignement Before realignement After realignement De. Pristo et al. Nat Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca

Strategies that improve variant calling • • Local realignment Duplicate marking Base quality recalibration Population structure and imputation Module 5: Small variant calling & annotation bioinformatics. ca

Duplicate marking www. broadinstitute. org Module 5: Small variant calling & annotation bioinformatics. ca

Strategies that improve variant calling • • Local realignment Duplicate marking Base quality recalibration Population structure and imputation Module 5: Small variant calling & annotation bioinformatics. ca
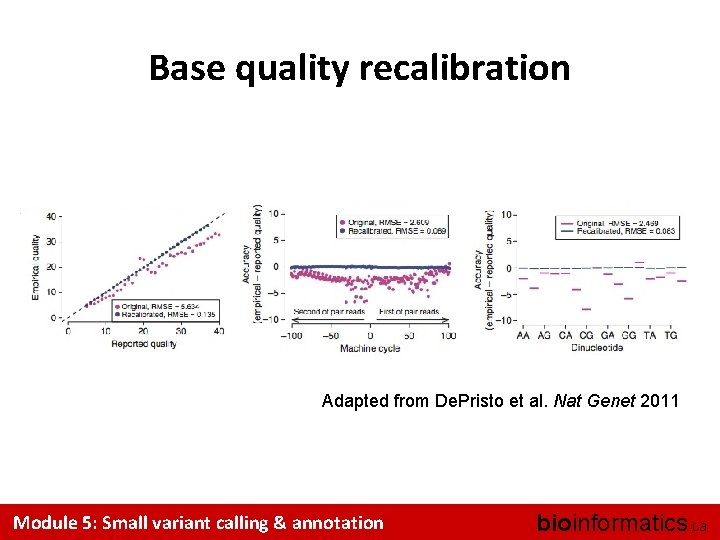
Base quality recalibration Adapted from De. Pristo et al. Nat Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca

Strategies that improve variant calling • • Local realignment Duplicate marking Base quality recalibration Population structure and imputation Module 5: Small variant calling & annotation bioinformatics. ca

Using haplotypes for base calling • Suppose that only 2 haplotypes have been observed in a population: Chr 1: . . A. . T. . . . G. . Chr 1: . . C. . G. . . . A. . • And that you observe the following reads: . . . A. . . . N. . . G. . . A. . N. . . . G. . . • Can you guess the value of N ? Module 5: Small variant calling & annotation bioinformatics. ca
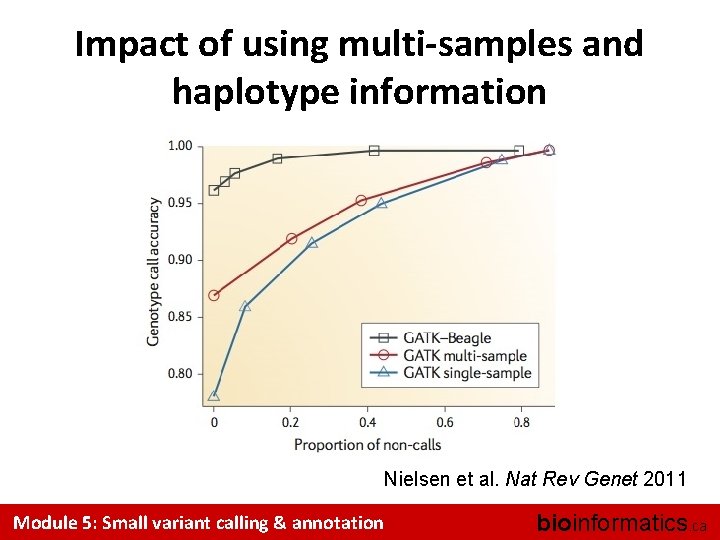
Impact of using multi-samples and haplotype information Nielsen et al. Nat Rev Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca
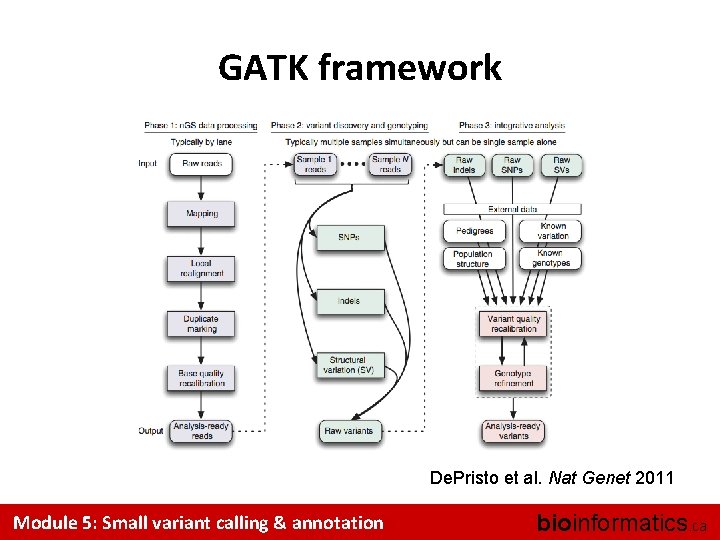
GATK framework De. Pristo et al. Nat Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca
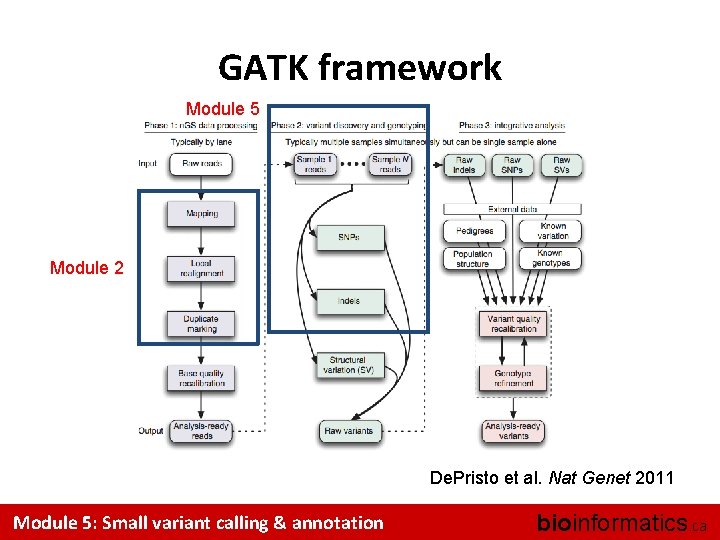
GATK framework Module 5 Module 2 De. Pristo et al. Nat Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca
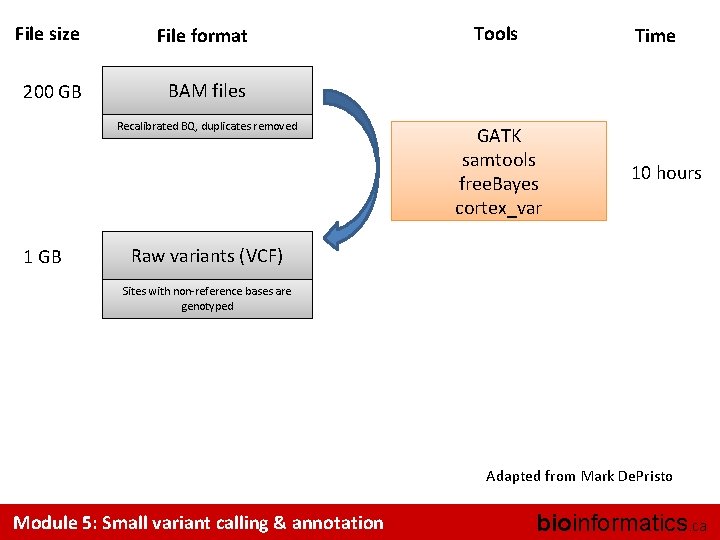
File size File format 200 GB BAM files Recalibrated BQ, duplicates removed 1 GB Tools Time GATK samtools free. Bayes cortex_var 10 hours Raw variants (VCF) Sites with non-reference bases are genotyped Adapted from Mark De. Pristo Module 5: Small variant calling & annotation bioinformatics. ca

Main analysis steps – – Quality control Pre-processing (trimming, remove adapters, …) Mapping (Module 2) Small variant calling (Module 5 – this module!) • SNP and indels calling • Variant filtering and annotation – Structural variant calling (Module 6) Module 5: Small variant calling & annotation bioinformatics. ca

VCF format Mandatory header line Reference base Quality score Alternative base Allele frequency, read depth, etc. https: //samtools. github. io/hts-specs/VCFv 4. 2. pdf Module 5: Small variant calling & annotation bioinformatics. ca

Variant filtering Raw variant calls have a lot of false positives. How to filter? 1. Manual filtering based on different parameters (e. g. using GATK Variant. Filtration or snp. Sift): – Based on quality score, depth of coverage, etc. – Difficult and requires time and expertise 2. Learn the filters from the data itself (e. g. GATK Variant. Recalibrator): – Better rank-order variants based on their likelihood of being real Module 5: Small variant calling & annotation bioinformatics. ca

QC: Hap. Map & db. SNP • International Hap. Map Project (phase III) – 1301 individuals in 11 populations genotyped – ~1 SNP per 2 kb – Proxy for false negatives • db. SNP (build 130) – 14 million SNPs in human genome – Varying quality – Proxy for false positives Michael Strömberg Module 5: Small variant calling & annotation bioinformatics. ca
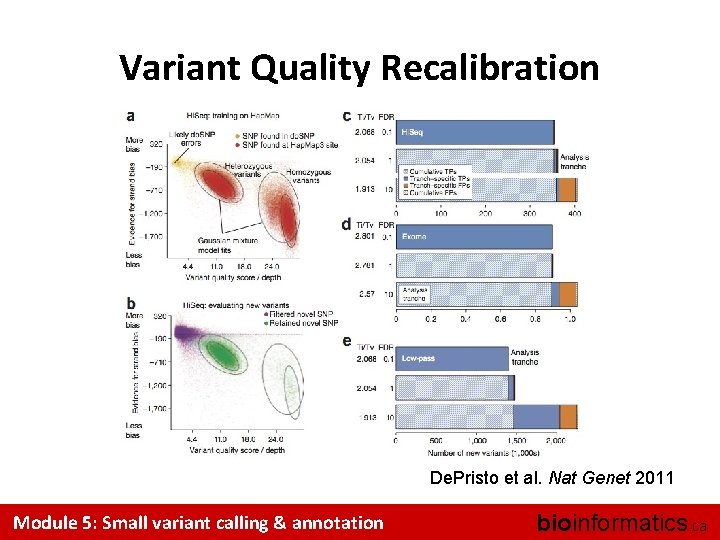
Variant Quality Recalibration De. Pristo et al. Nat Genet 2011 Module 5: Small variant calling & annotation bioinformatics. ca
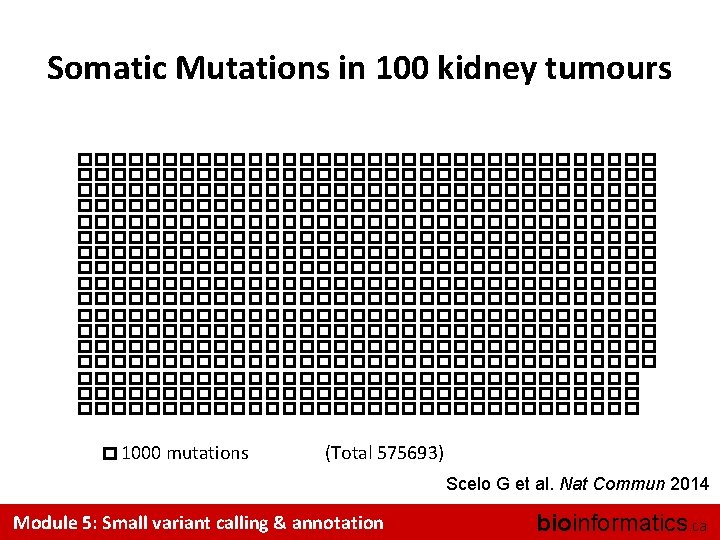
Somatic Mutations in 100 kidney tumours 1000 mutations (Total 575693) Scelo G et al. Nat Commun 2014 Module 5: Small variant calling & annotation bioinformatics. ca
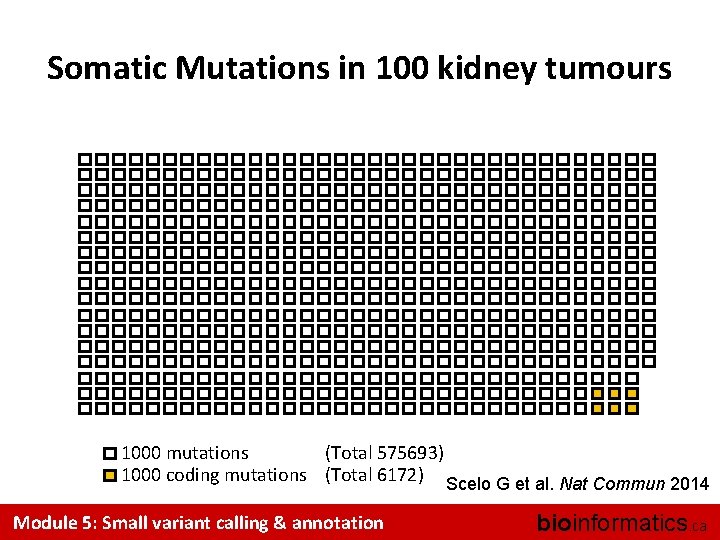
Somatic Mutations in 100 kidney tumours 1000 mutations (Total 575693) 1000 coding mutations (Total 6172) Scelo G et al. Nat Commun 2014 Module 5: Small variant calling & annotation bioinformatics. ca

Annotating variants with Snp. Eff • Annotations using reference genomes • Calculate effects: – Coding (e. g. Syn, Non-Syn, Stop gained, Splice) – Non-coding (e. g. TFBS) • Basic prioritizations (putative impact): {HIGH, MODERATE, LOW, MODIFIER} • And many other things… Pablo Cingolani Module 5: Small variant calling & annotation bioinformatics. ca
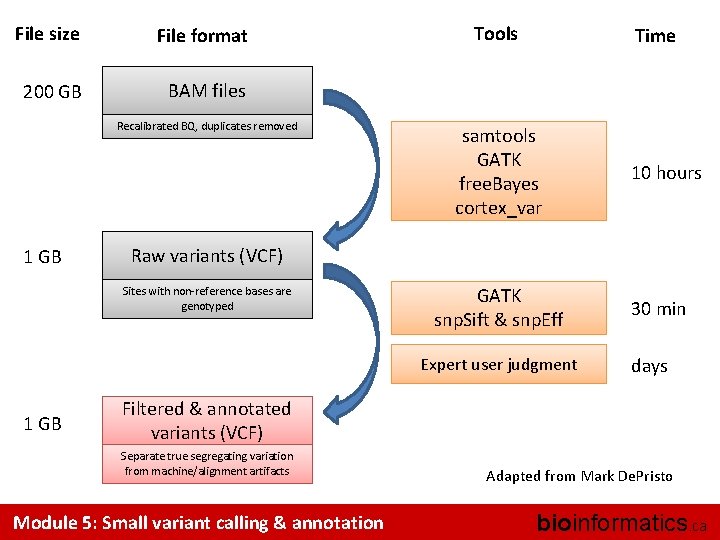
File size File format 200 GB BAM files Recalibrated BQ, duplicates removed 1 GB Tools Time samtools GATK free. Bayes cortex_var Raw variants (VCF) Sites with non-reference bases are genotyped GATK snp. Sift & snp. Eff Expert user judgment 1 GB 10 hours 30 min days Filtered & annotated variants (VCF) Separate true segregating variation from machine/alignment artifacts Module 5: Small variant calling & annotation Adapted from Mark De. Pristo bioinformatics. ca

Lab time! Module 5: Small variant calling & annotation bioinformatics. ca

We are on a Coffee Break & Networking Session Module 5: Small variant calling & annotation bioinformatics. ca
- Slides: 43