Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2
Module 6 Structural variant calling Guillaume Bourque Informatics on High-throughput Sequencing Data June 10 -11, 2015

Learning Objectives of Module • To understand what are structural variants (SVs) • To appreciate how SVs are discovered from NGS data • To appreciate the strengths and weaknesses of each SV discovery strategy • To recognize the sequence alignment SV “signals” • To be able to visually explore read support for SVs Module 6: Structural variant calling bioinformatics. ca

Structural Variants (SVs): Genomic rearrangements that affect >50 bp (or 100 bp, or 1 Kb) of sequence, including: • deletions • novel insertions • inversions • mobile-element transpositions • duplications • translocations Adapted from Alkan et al. Nat Rev Genet 2011 Module 6: Structural variant calling bioinformatics. ca
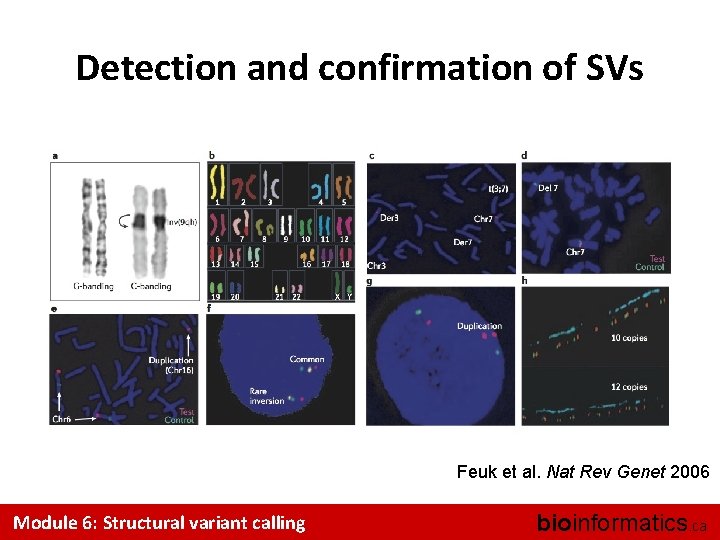
Detection and confirmation of SVs Feuk et al. Nat Rev Genet 2006 Module 6: Structural variant calling bioinformatics. ca

Structural variants in cancer Can higher resolution maps help identify recurrent aberrations and driver mutations in cancer? Module 6: Structural variant calling bioinformatics. ca

Classes of SVs • Copy number variants (CNVs): – Deletions – Duplications • Copy neutral rearrangements: – Inversions – Translocations • Other structural variants: – Novel insertions – Mobile-element transpositions Module 6: Structural variant calling bioinformatics. ca

Classes of SVs Alkan et al. Nat Rev Genet 2011 Module 6: Structural variant calling bioinformatics. ca

Our understanding is driven by technology Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca

Array-based detection of CNVs Alkan et al. Nat Rev Genet 2011 Module 6: Structural variant calling bioinformatics. ca

Detecting SVs from NGS data Meyerson et al. Nat Rev Genet 2010 Module 6: Structural variant calling bioinformatics. ca
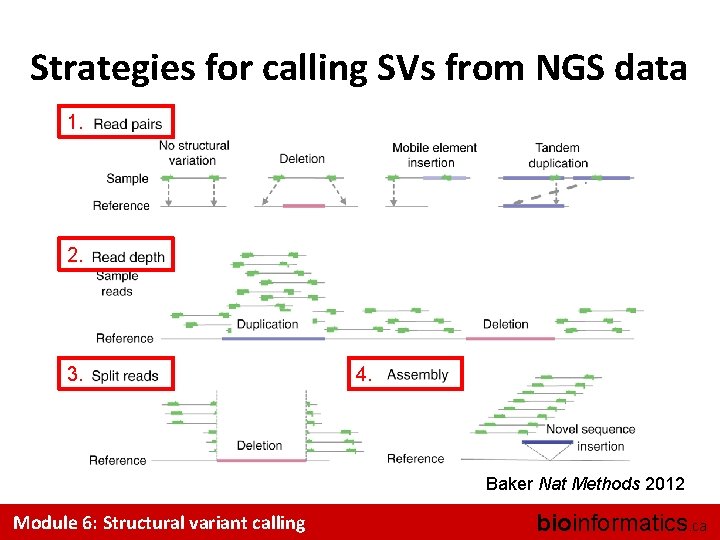
Strategies for calling SVs from NGS data 1. 2. 3. 4. Baker Nat Methods 2012 Module 6: Structural variant calling bioinformatics. ca

Strategies for calling SVs from NGS data 1. Baker Nat Methods 2012 Module 6: Structural variant calling bioinformatics. ca
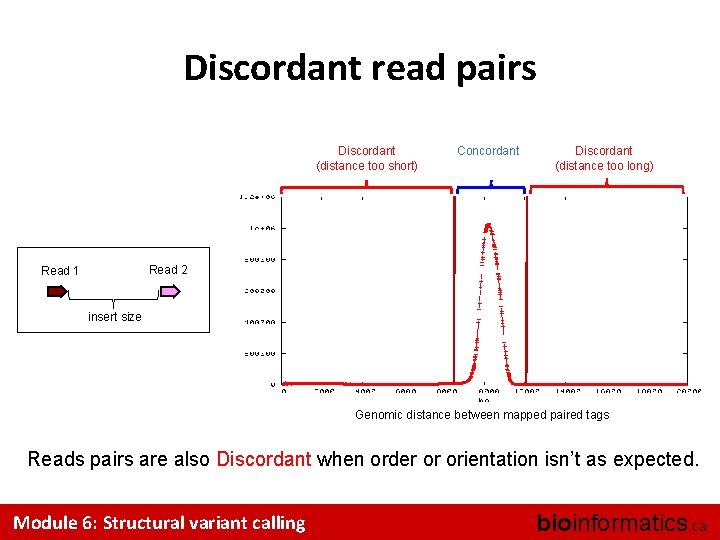
Discordant read pairs Discordant (distance too short) Concordant Discordant (distance too long) Read 2 Read 1 insert size Genomic distance between mapped paired tags Reads pairs are also Discordant when order or orientation isn’t as expected. Module 6: Structural variant calling bioinformatics. ca
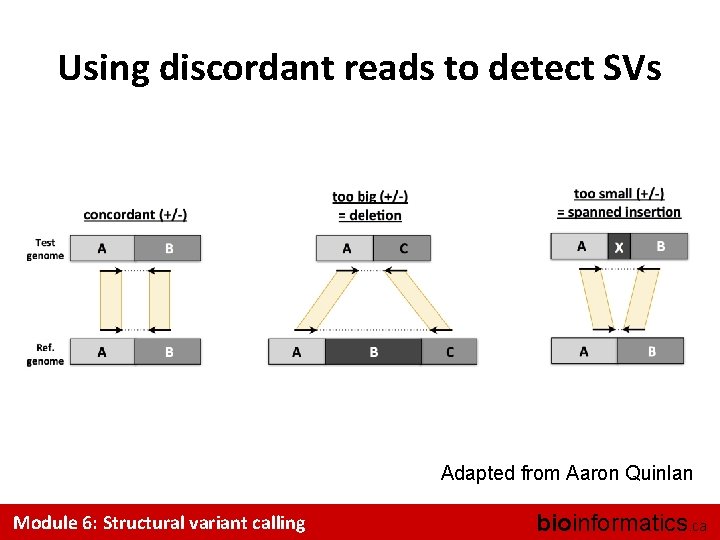
Using discordant reads to detect SVs Adapted from Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
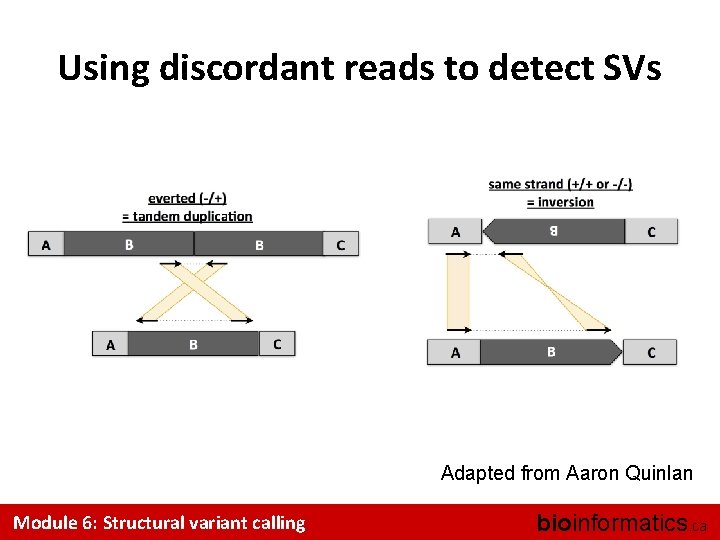
Using discordant reads to detect SVs Adapted from Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
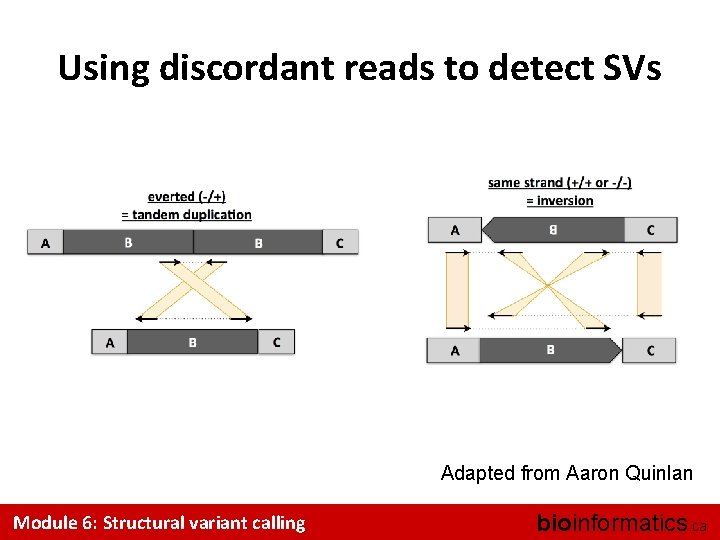
Using discordant reads to detect SVs Adapted from Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
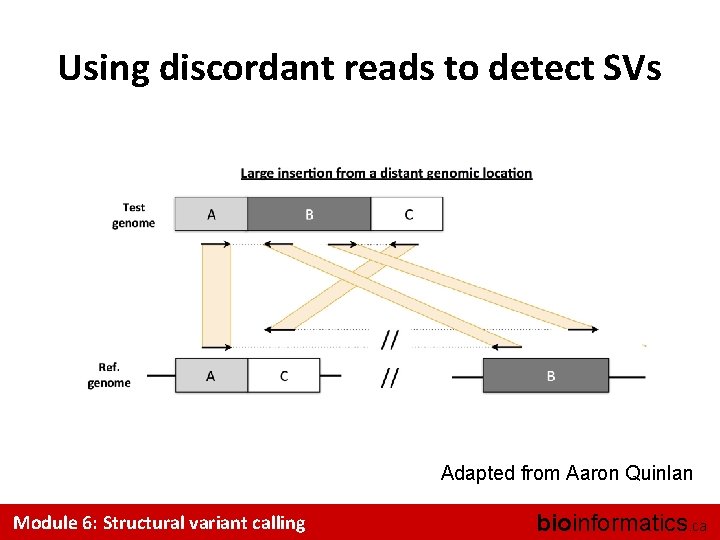
Using discordant reads to detect SVs Adapted from Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca

Read-pair tools • • Break. Dancer Variation. Hunter Mo. DIL GASV-PRO DELLY LUMPY Genome. STRi. P… Module 6: Structural variant calling bioinformatics. ca
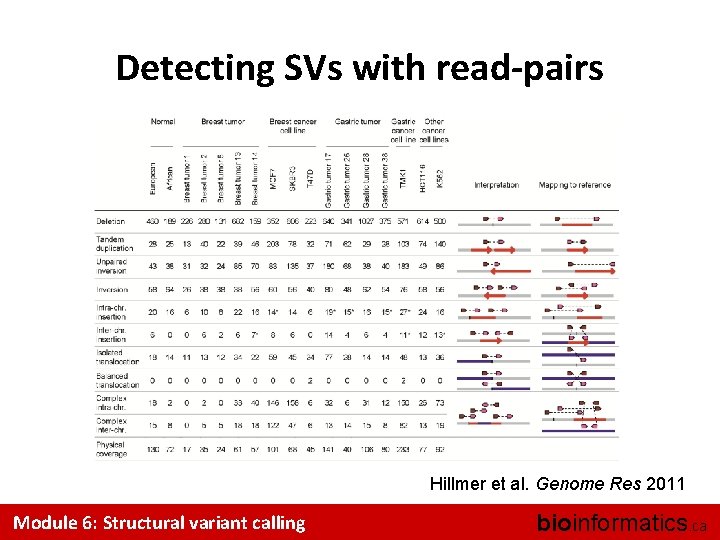
Detecting SVs with read-pairs Hillmer et al. Genome Res 2011 Module 6: Structural variant calling bioinformatics. ca
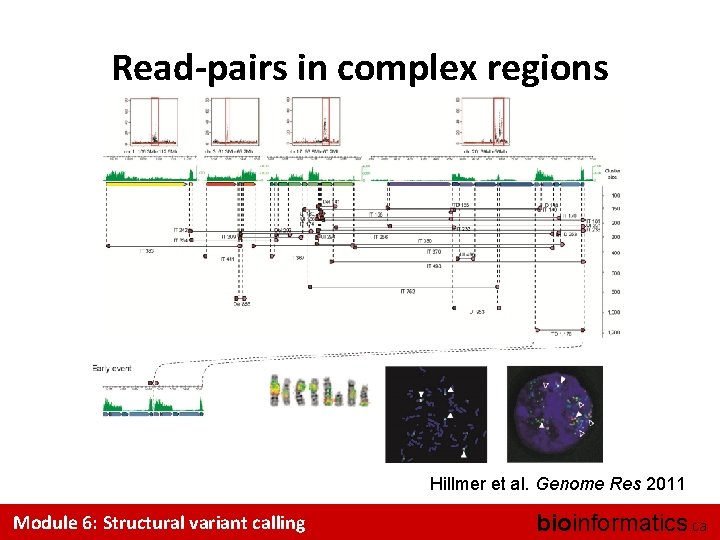
Read-pairs in complex regions Hillmer et al. Genome Res 2011 Module 6: Structural variant calling bioinformatics. ca

Read-pair summary • Weaknesses – Difficult to interpret read-pairs in repetitive regions – Difficult to fully characterize highly rearranged regions – High rate of false positives • Strengths: – Most classes of variation can, in principle, be detected Module 6: Structural variant calling bioinformatics. ca

Strategies for calling SVs from NGS data 2. Baker Nat Methods 2012 Module 6: Structural variant calling bioinformatics. ca

Read-depth Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
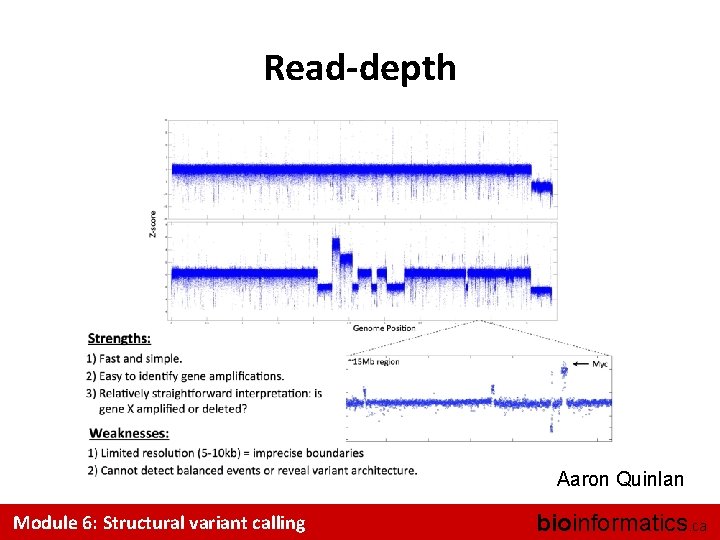
Read-depth Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
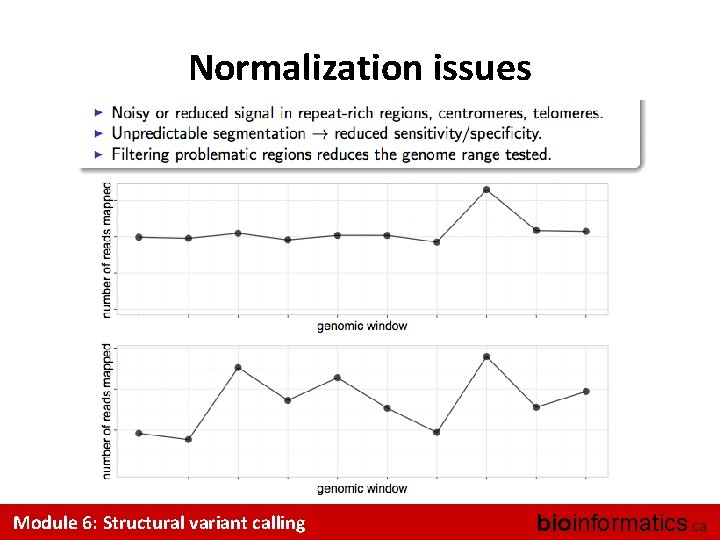
Normalization issues Module 6: Structural variant calling bioinformatics. ca
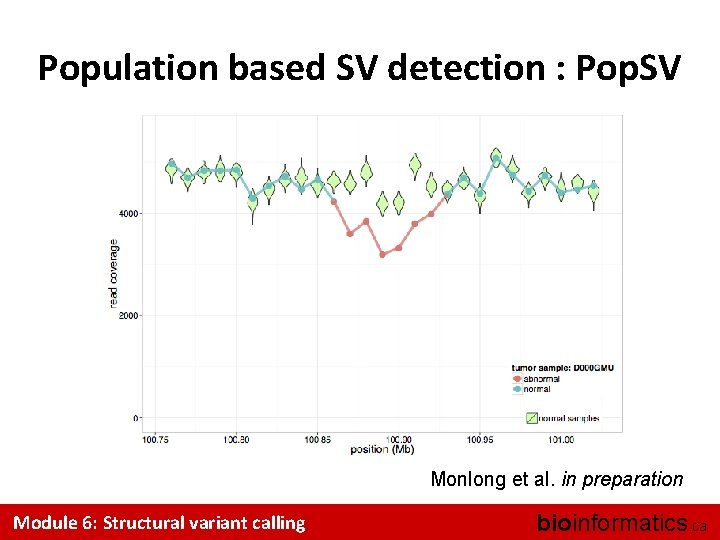
Population based SV detection : Pop. SV Monlong et al. in preparation Module 6: Structural variant calling bioinformatics. ca

Read depth tools • • • Read. Depth RDXplorer cnv. Seq CNVer Copy. Seq Genome. STRi. P CNVnator Pop. SV … Module 6: Structural variant calling bioinformatics. ca

Read-depth summary • Weaknesses – Relatively low resolution (normally ~10 Kb) – Cannot detect balanced rearrangements (e. g. , inversions), or transposon insertions • Strengths: – Determines DNA copy number (unlike most other methods) – Provides useful information even with low coverage, albeit at low resolution Module 6: Structural variant calling bioinformatics. ca

Strategies for calling SVs from NGS data 3. Baker Nat Methods 2012 Module 6: Structural variant calling bioinformatics. ca

Split reads Rausch et al. Bioinformatics 2012 Module 6: Structural variant calling bioinformatics. ca

Split read tools • • • Pindel DELLY LUMPY PRISM Mobster … Module 6: Structural variant calling bioinformatics. ca

Split reads summary • Weaknesses – Requires sufficient coverage – Can have false positives especially in repetitive regions • Strengths: – Can be added to read-pairs methods – Base-pair resolution of breakpoints Module 6: Structural variant calling bioinformatics. ca

Strategies for calling SVs from NGS data 4. Baker Nat Methods 2012 Module 6: Structural variant calling bioinformatics. ca
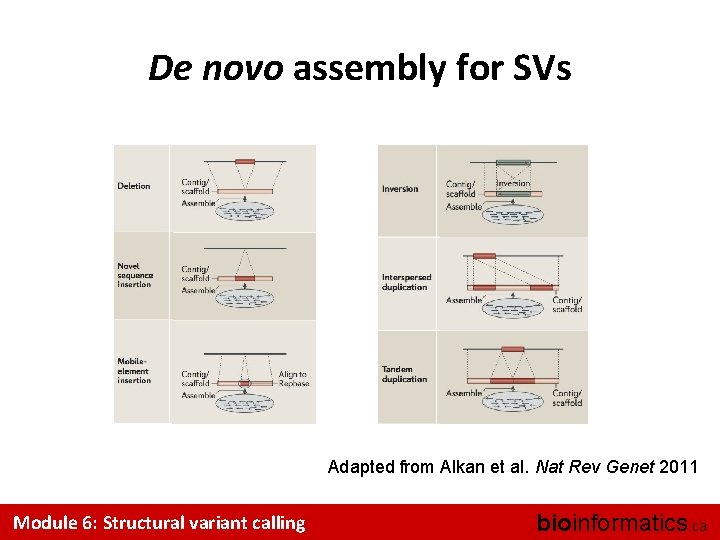
De novo assembly for SVs Adapted from Alkan et al. Nat Rev Genet 2011 Module 6: Structural variant calling bioinformatics. ca

De novo assembly tools for SVs • • • Cortex SGA DISCOVAR ABy. SS Ray … Module 6: Structural variant calling bioinformatics. ca

De novo assembly for SVs summary • Weaknesses – Computationally very intensive – Hard to resolve repetitive and complex regions • Strengths: – Base-pair resolution of breakpoints – All classes of variation can, in principle, be detected Module 6: Structural variant calling bioinformatics. ca
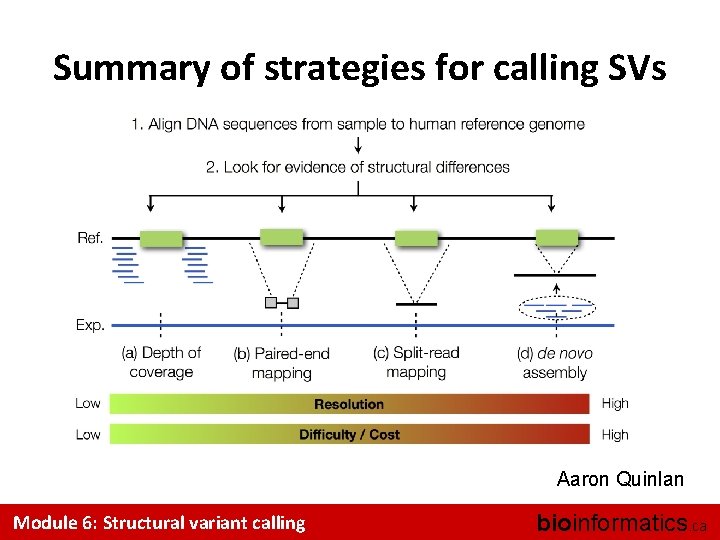
Summary of strategies for calling SVs Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
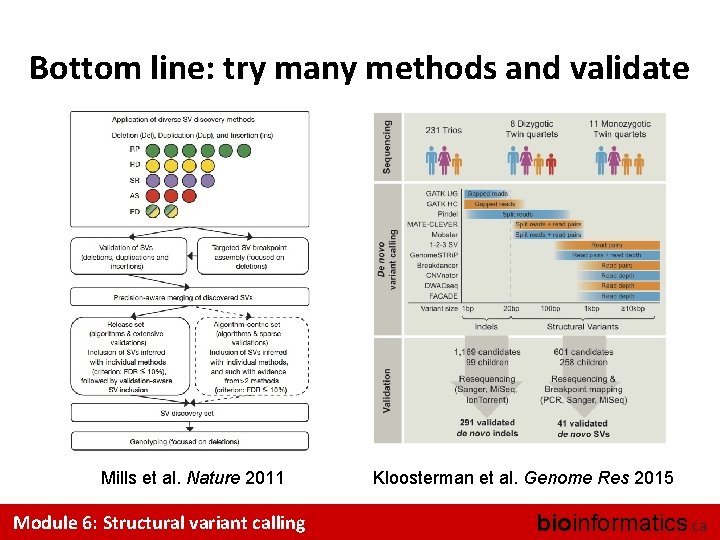
Bottom line: try many methods and validate Mills et al. Nature 2011 Module 6: Structural variant calling Kloosterman et al. Genome Res 2015 bioinformatics. ca

Visual validation: a deletion Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
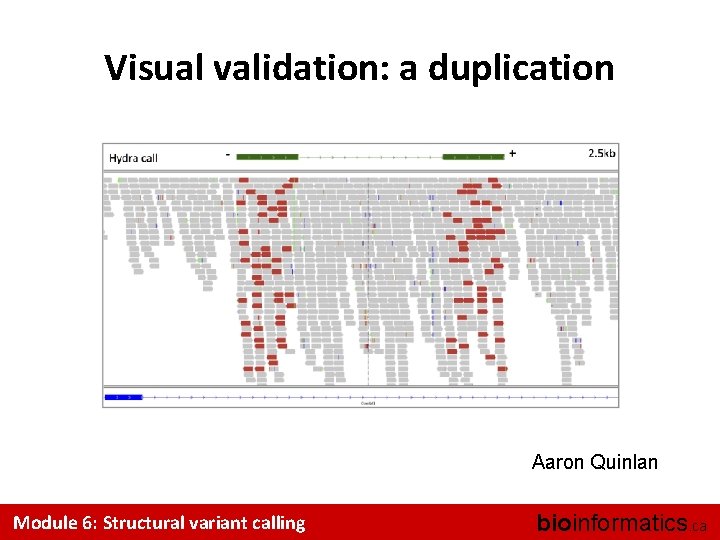
Visual validation: a duplication Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
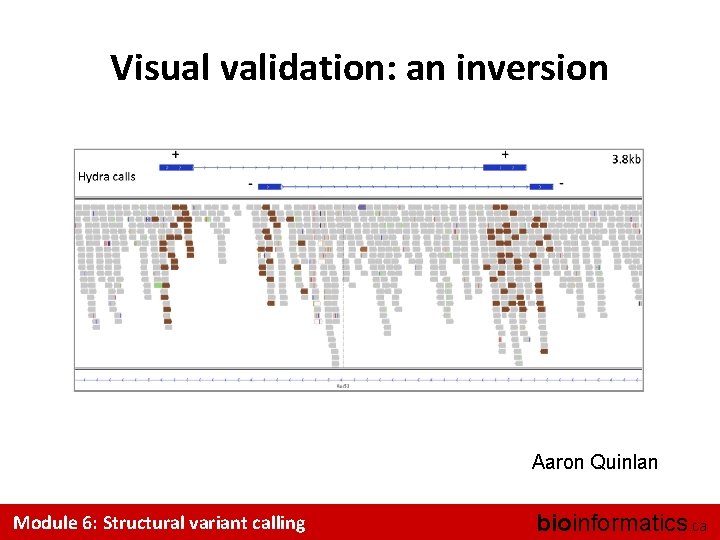
Visual validation: an inversion Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca

Visual validation: an insertion (in the reference) Aaron Quinlan Module 6: Structural variant calling bioinformatics. ca
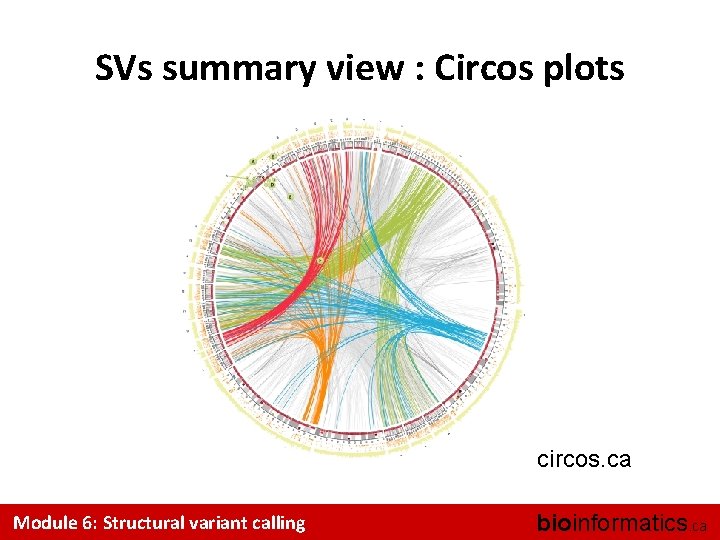
SVs summary view : Circos plots circos. ca Module 6: Structural variant calling bioinformatics. ca

Lab time! Module 6: Structural variant calling bioinformatics. ca

We are on a Coffee Break & Networking Session Module 6: Structural variant calling bioinformatics. ca
- Slides: 47