Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2
Module 3 Clinical Data Integration Anna Lapuk, Ph. D Bioinformatics for Cancer Genomics May 27 – 31 st, 2013

Module 3 overview • Part I – Clinical Data and Biomarkers • Part II – Statistical aspects • Lab: – Survival analysis Module 3: Clinical Data Integration bioinformatics. ca

Learning Objectives • To understand how clinical information is used • To understand the process of genomic data integration with clinical information • To review the current advances on the biomarker/clinical applications development • To learn how to evaluate the biomarker and perform survival analysis • Be able to analyze tumour cohorts with regard to association of molecular subgroups with outcome. Module 3: Clinical Data Integration bioinformatics. ca

What is the clinical data Module 3: Clinical Data Integration bioinformatics. ca
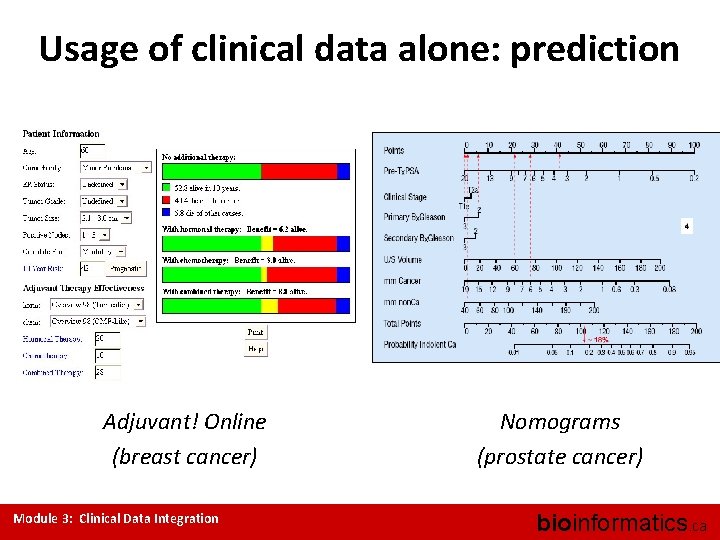
Usage of clinical data alone: prediction Adjuvant! Online (breast cancer) Module 3: Clinical Data Integration Nomograms (prostate cancer) bioinformatics. ca
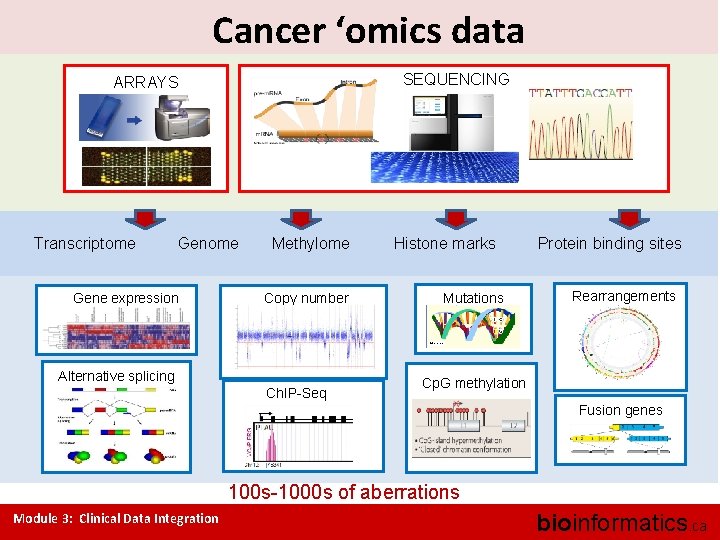
Cancer ‘omics data SEQUENCING ARRAYS Transcriptome Gene expression Methylome Copy number Alternative splicing Ch. IP-Seq Histone marks Mutations Protein binding sites Rearrangements s Cp. G methylation Fusion genes 100 s-1000 s of aberrations Module 3: Clinical Data Integration bioinformatics. ca
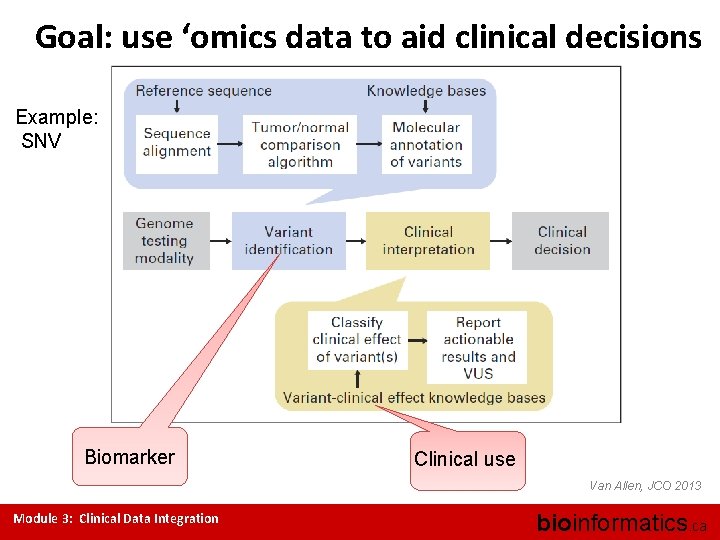
Goal: use ‘omics data to aid clinical decisions Example: SNV Biomarker Clinical use Van Allen, JCO 2013 Module 3: Clinical Data Integration bioinformatics. ca

Biomarkers & therapeutic targets (NCI) • Biomarker is a biological molecule (or a set thereof) found in blood, other body fluids, or tissues that is a sign of a normal or abnormal process, or of a condition or disease. Also called molecular marker and signature molecule. • Therapeutic target is a biological molecule, an enzyme, receptor or other protein that can be modified by an external stimulus (drug). The implication is that a molecule is "hit" by a signal and its behavior is thereby changed. Module 3: Clinical Data Integration bioinformatics. ca

Biomarker features • Biomarker comes from alterations: – – Germline/somatic mutation Genomic amplification/deletion Transcriptional change Post-transcriptional modification • Biomarker types: – – – Proteins Nucleic acids (m. RNA, mi. RNA, non-coding RNA) Cells (Circulating Tumour Cells) Peptides Individual molecules vs Sets (signatures): • Gene expression (n genes) • Proteomic (n proteins) • Metabolomic (n metabolites) Module 3: Clinical Data Integration bioinformatics. ca

Biomarker features (cont’d) • Biomarker screening: – Circulation (blood, serum, plasma) – Excretions/secretions (stool, urine, etc. ) – Tissues (biopsy + imaging) • Biomarkers may be also therapeutic targets, but not always: – HER 2 (breast cancer): biomarker and therapeutic target – PSA (prostate cancer): biomarker. AR – target. – KRAS mutations (colorectal cancer): biomarker. EGFR – target. Module 3: Clinical Data Integration bioinformatics. ca

Clinical use of biomarkers • To diagnose or subclassify the disease state => diagnostic – BCR-ABL fusion leukemia (Philadelphia chromosome) • To make prognosis about a clinical outcome (survival or recurrence) => prognostic – Oncotype. Dx gene expression (estimates the risk of breast cancer recurrence) • To predict the activity of a therapy => predictive – HER 2 and herceptin (predicts response in breast cancer) • To identify a subgroup of patients for whom therapy has shown benefit => companion diagnostic markers – BRAF V 600 E mutation and BRAF inhibitor (confers sensitivity in melanoma) Module 3: Clinical Data Integration bioinformatics. ca

Examples of use Module 3: Clinical Data Integration bioinformatics. ca
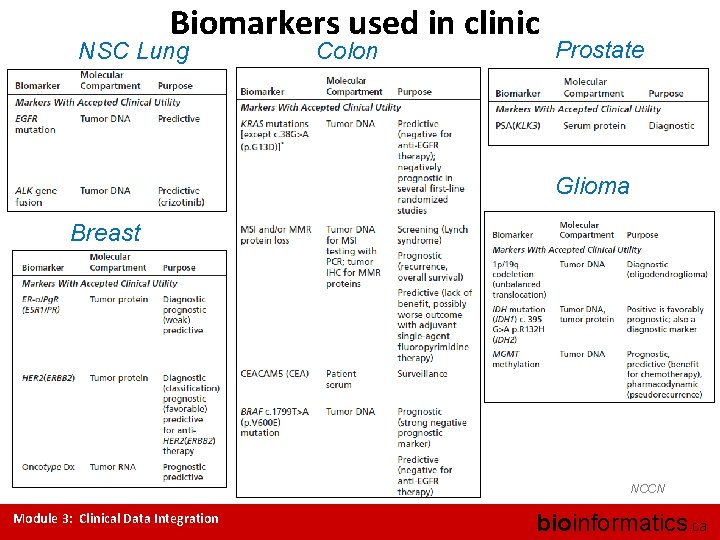
Biomarkers used in clinic NSC Lung Colon Prostate Glioma Breast Glioma NCCN Module 3: Clinical Data Integration bioinformatics. ca
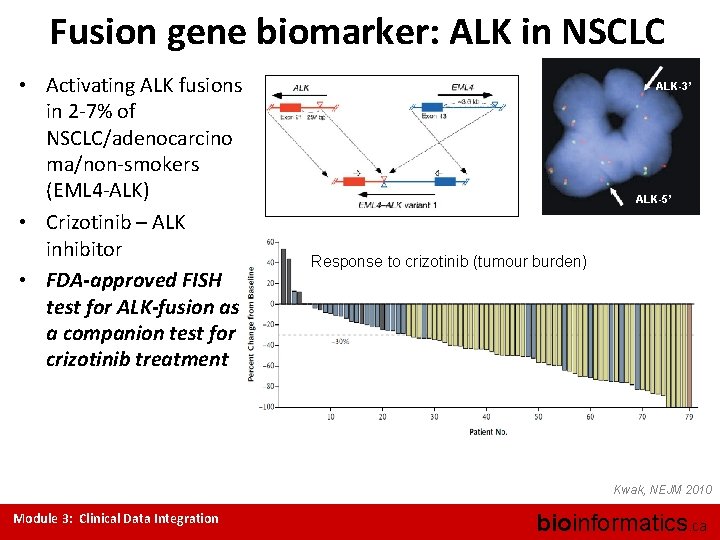
Fusion gene biomarker: ALK in NSCLC • Activating ALK fusions in 2 -7% of NSCLC/adenocarcino ma/non-smokers (EML 4 -ALK) • Crizotinib – ALK inhibitor • FDA-approved FISH test for ALK-fusion as a companion test for crizotinib treatment ALK-3’ ALK-5’ Response to crizotinib (tumour burden) Kwak, NEJM 2010 Module 3: Clinical Data Integration bioinformatics. ca
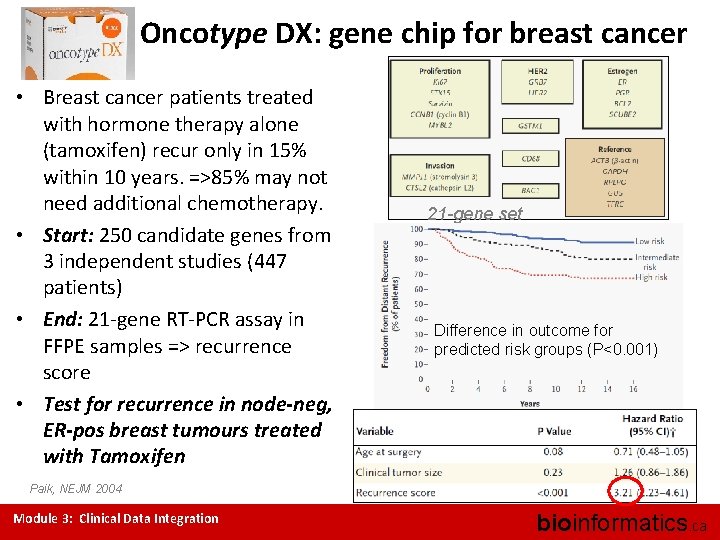
Oncotype DX: gene chip for breast cancer • Breast cancer patients treated with hormone therapy alone (tamoxifen) recur only in 15% within 10 years. =>85% may not need additional chemotherapy. • Start: 250 candidate genes from 3 independent studies (447 patients) • End: 21 -gene RT-PCR assay in FFPE samples => recurrence score • Test for recurrence in node-neg, ER-pos breast tumours treated with Tamoxifen 21 -gene set Difference in outcome for predicted risk groups (P<0. 001) Paik, NEJM 2004 Module 3: Clinical Data Integration bioinformatics. ca
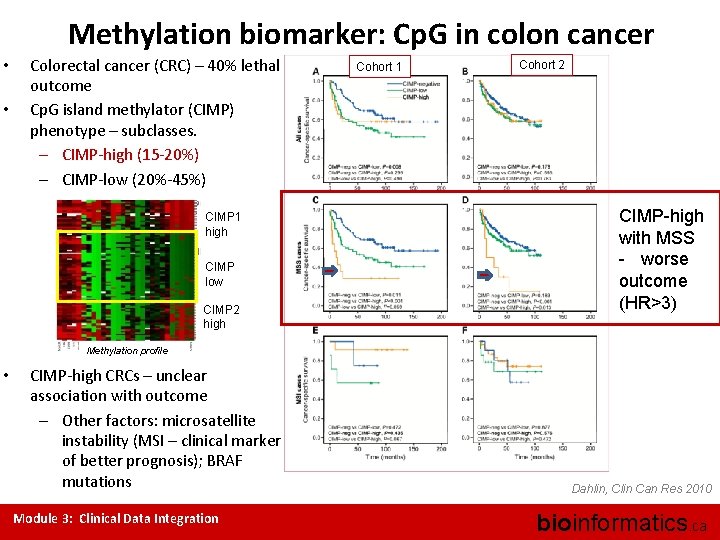
Methylation biomarker: Cp. G in colon cancer • • Colorectal cancer (CRC) – 40% lethal outcome Cp. G island methylator (CIMP) phenotype – subclasses. – CIMP-high (15 -20%) – CIMP-low (20%-45%) CIMP 1 high CIMP low CIMP 2 high Cohort 1 Cohort 2 CIMP-high with MSS - worse outcome (HR>3) Methylation profile • CIMP-high CRCs – unclear association with outcome – Other factors: microsatellite instability (MSI – clinical marker of better prognosis); BRAF mutations Module 3: Clinical Data Integration Dahlin, Clin Can Res 2010 bioinformatics. ca
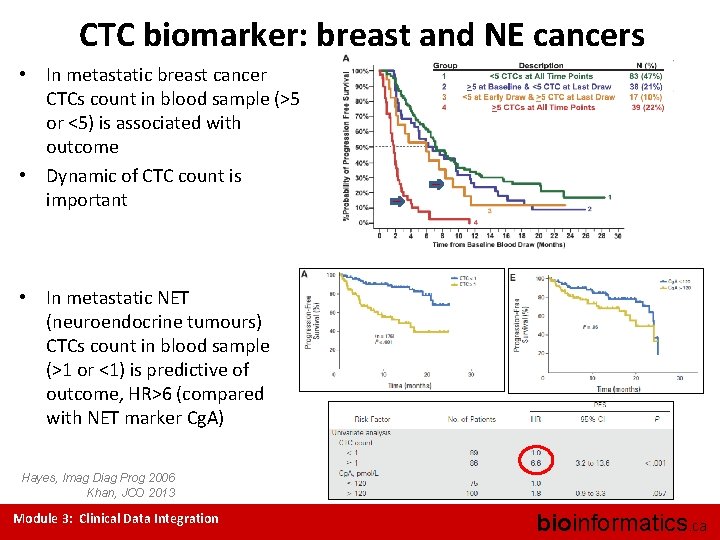
CTC biomarker: breast and NE cancers • In metastatic breast cancer CTCs count in blood sample (>5 or <5) is associated with outcome • Dynamic of CTC count is important • In metastatic NET (neuroendocrine tumours) CTCs count in blood sample (>1 or <1) is predictive of outcome, HR>6 (compared with NET marker Cg. A) Hayes, Imag Diag Prog 2006 Khan, JCO 2013 Module 3: Clinical Data Integration bioinformatics. ca

Biomarker development • Identification. Discovery approach to identify biomarkers that are different between cohorts of tumours using variety of technologies – Microarrays/ sequencing/ mass spectrometry. – Important: careful study design to avoid bias in biomarker discovery (matched cases and controls)! • Validation. – Analytical validity. • Biomarker assay: reproducibility, sensitivity, specificity. – Clinical validity • How reliably the biomarker divides the populations into 2 groups of different outcomes. Important: validation should be done on independent cohorts of tumours! – Clinical utility • Does the biomarker able to improve the clinical decision-making. Depends on the strength of association of biomarker with outcome, the size of the effect, particular disease and overall benefits, risks and economics. Example: marker identifies 2 subgroups of tumours with very different survival. However, no treatment options are available => no clinical utility. Module 3: Clinical Data Integration bioinformatics. ca
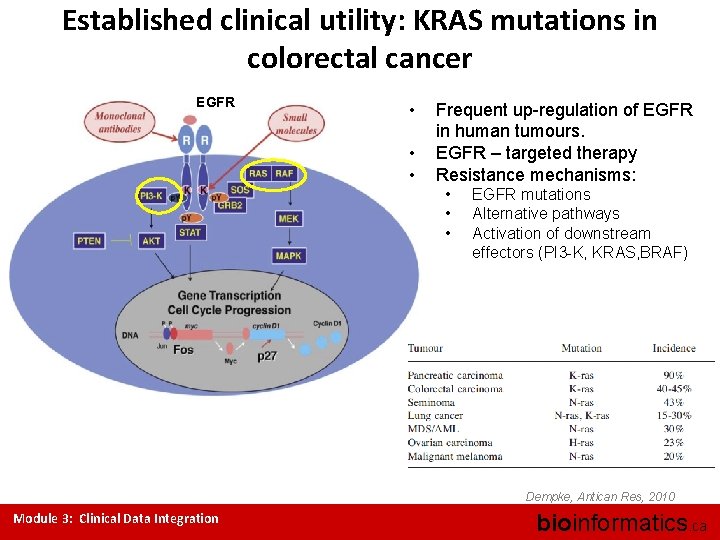
Established clinical utility: KRAS mutations in colorectal cancer EGFR • • • Frequent up-regulation of EGFR in human tumours. EGFR – targeted therapy Resistance mechanisms: • • • EGFR mutations Alternative pathways Activation of downstream effectors (PI 3 -K, KRAS, BRAF) Dempke, Antican Res, 2010 Module 3: Clinical Data Integration bioinformatics. ca
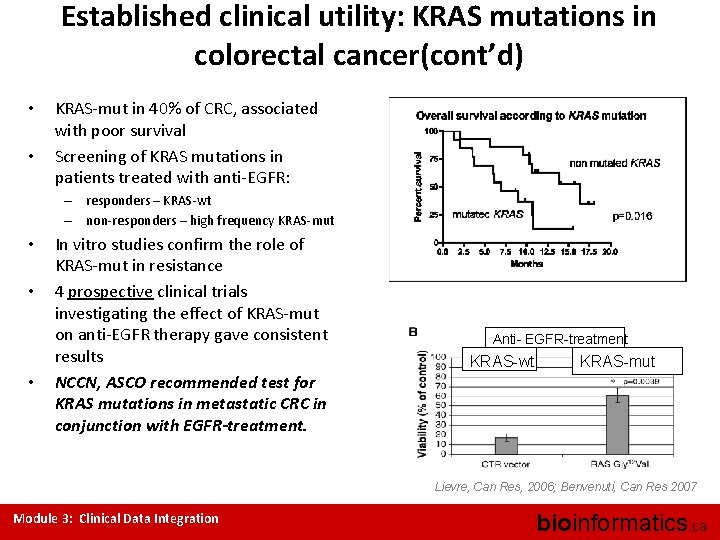
Established clinical utility: KRAS mutations in colorectal cancer(cont’d) • • KRAS-mut in 40% of CRC, associated with poor survival Screening of KRAS mutations in patients treated with anti-EGFR: – responders – KRAS-wt – non-responders – high frequency KRAS-mut • • • In vitro studies confirm the role of KRAS-mut in resistance 4 prospective clinical trials investigating the effect of KRAS-mut on anti-EGFR therapy gave consistent results NCCN, ASCO recommended test for KRAS mutations in metastatic CRC in conjunction with EGFR-treatment. Anti- EGFR-treatment KRAS-wt KRAS-mut Lievre, Can Res, 2006; Benvenuti, Can Res 2007 Module 3: Clinical Data Integration bioinformatics. ca
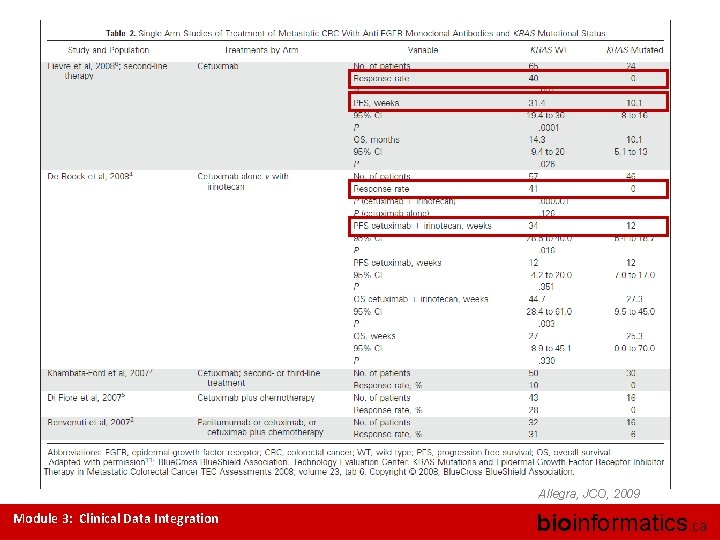
Allegra, JCO, 2009 Module 3: Clinical Data Integration bioinformatics. ca
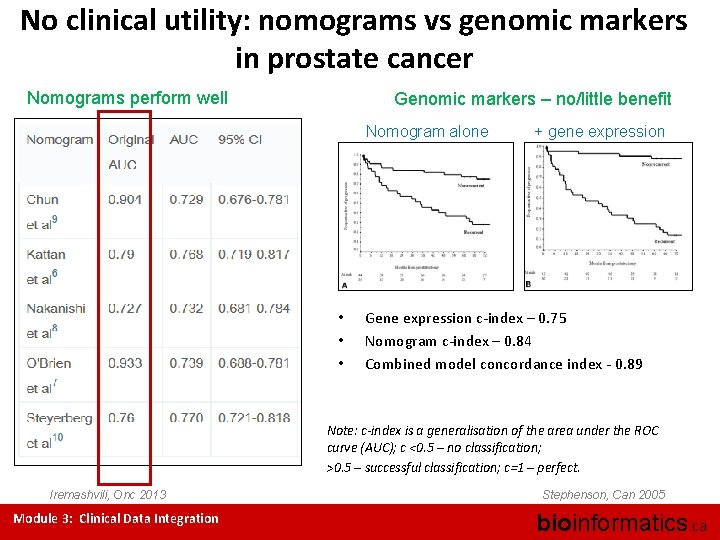
No clinical utility: nomograms vs genomic markers in prostate cancer Nomograms perform well Genomic markers – no/little benefit Nomogram alone • • • + gene expression Gene expression c-index – 0. 75 Nomogram c-index – 0. 84 Combined model concordance index - 0. 89 Note: c-index is a generalisation of the area under the ROC curve (AUC); c <0. 5 – no classification; >0. 5 – successful classification; c=1 – perfect. Iremashvili, Onc 2013 Module 3: Clinical Data Integration Stephenson, Can 2005 bioinformatics. ca
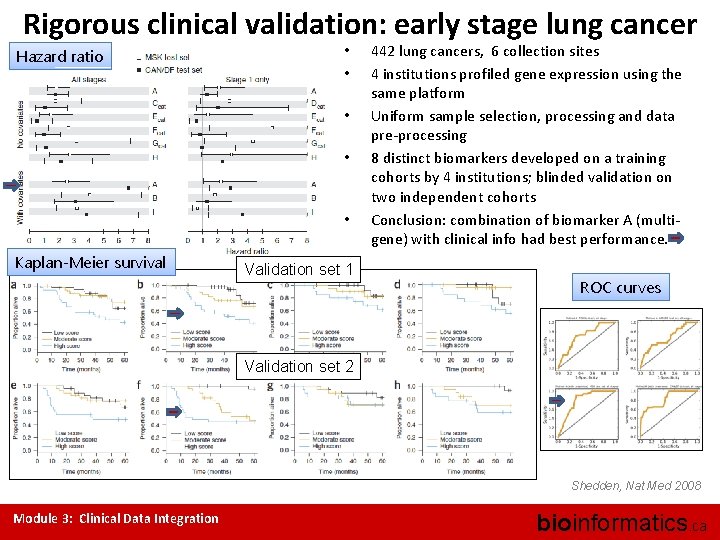
Rigorous clinical validation: early stage lung cancer Hazard ratio • • • Kaplan-Meier survival Validation set 1 442 lung cancers, 6 collection sites 4 institutions profiled gene expression using the same platform Uniform sample selection, processing and data pre-processing 8 distinct biomarkers developed on a training cohorts by 4 institutions; blinded validation on two independent cohorts Conclusion: combination of biomarker A (multigene) with clinical info had best performance. ROC curves Validation set 2 Shedden, Nat Med 2008 Module 3: Clinical Data Integration bioinformatics. ca
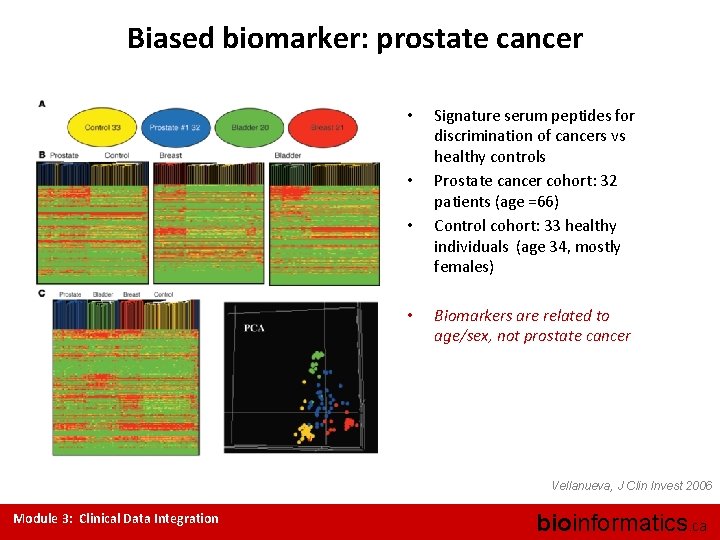
Biased biomarker: prostate cancer • • Signature serum peptides for discrimination of cancers vs healthy controls Prostate cancer cohort: 32 patients (age =66) Control cohort: 33 healthy individuals (age 34, mostly females) Biomarkers are related to age/sex, not prostate cancer Vellanueva, J Clin Invest 2006 Module 3: Clinical Data Integration bioinformatics. ca

Statistical aspects of biomarker development Module 3: Clinical Data Integration bioinformatics. ca
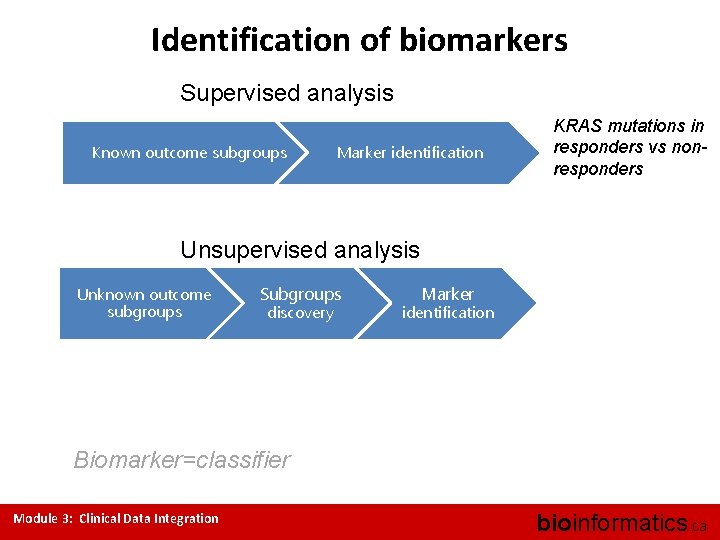
Identification of biomarkers Supervised analysis Known outcome subgroups Marker identification KRAS mutations in responders vs nonresponders Unsupervised analysis Unknown outcome subgroups Subgroups discovery Marker identification Biomarker=classifier Module 3: Clinical Data Integration bioinformatics. ca
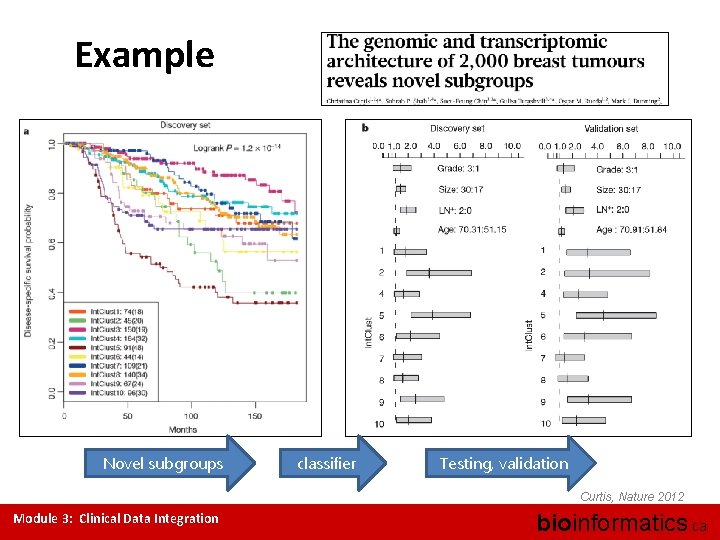
Example Novel subgroups classifier Testing, validation Curtis, Nature 2012 Module 3: Clinical Data Integration bioinformatics. ca
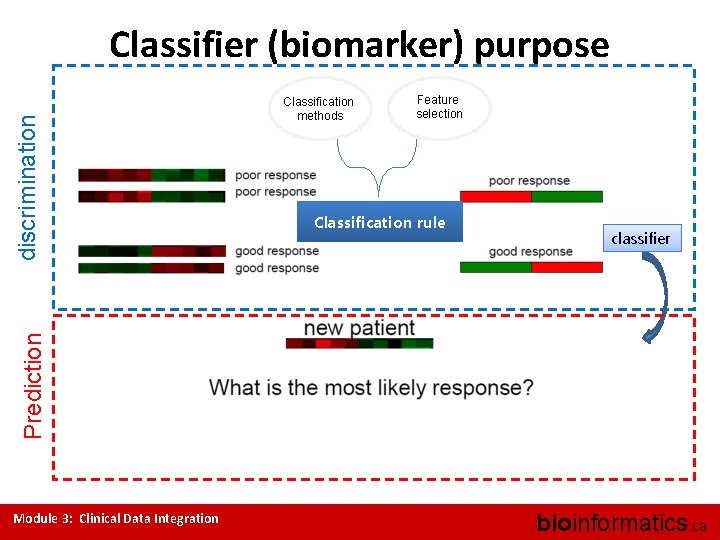
Classification methods Feature selection Classification rule classifier Prediction discrimination Classifier (biomarker) purpose Module 3: Clinical Data Integration bioinformatics. ca
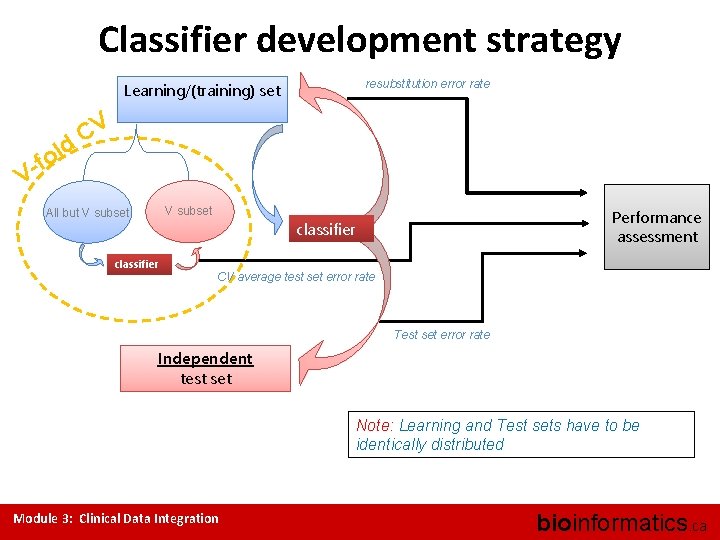
Classifier development strategy resubstitution error rate Learning/(training) set ld o -f CV V V subset All but V subset classifier Performance assessment classifier CV average test set error rate Test set error rate Independent test set Note: Learning and Test sets have to be identically distributed Module 3: Clinical Data Integration bioinformatics. ca

Classifier performance assessment • How accurate is classifier (confusion matrix, accuracy) • How well classifier worked on learning set (resubstitution error rate) • How well classifier worked on test set (test set error rate) • Cross validation • How do different classifiers compare (ROC curves) Module 3: Clinical Data Integration bioinformatics. ca

Confusion matrix Actual diagnose (pathology) (patient is positive/negative for cancer) Fractionpo positive negative Positive True positive False positive negative False negative True negative Sensitivity (true pos rate) Specificity (true neg rate) s Prediction using biomarkers Fractionneg Positive predictive value Negative predictive value Accuracy ACC = (TP + TN) / (P + N) or ACC=Sensitivity*Fractionpos + specificity*Fractionneg Module 3: Clinical Data Integration bioinformatics. ca
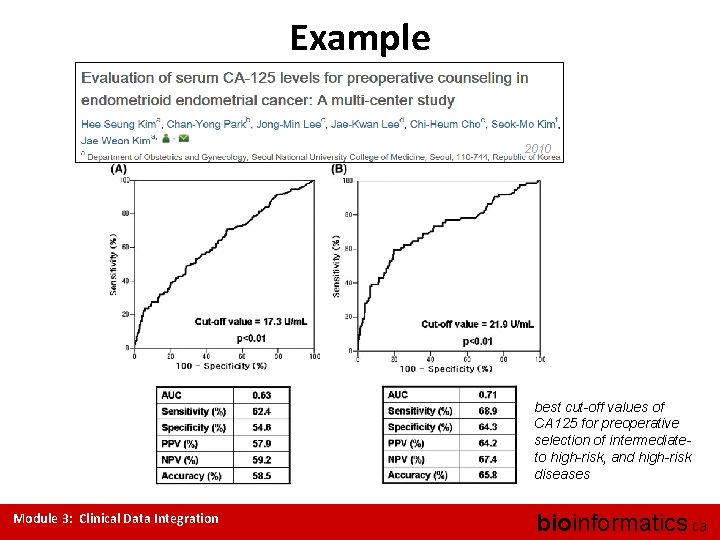
Example 2010 best cut-off values of CA 125 for preoperative selection of intermediateto high-risk, and high-risk diseases Module 3: Clinical Data Integration bioinformatics. ca
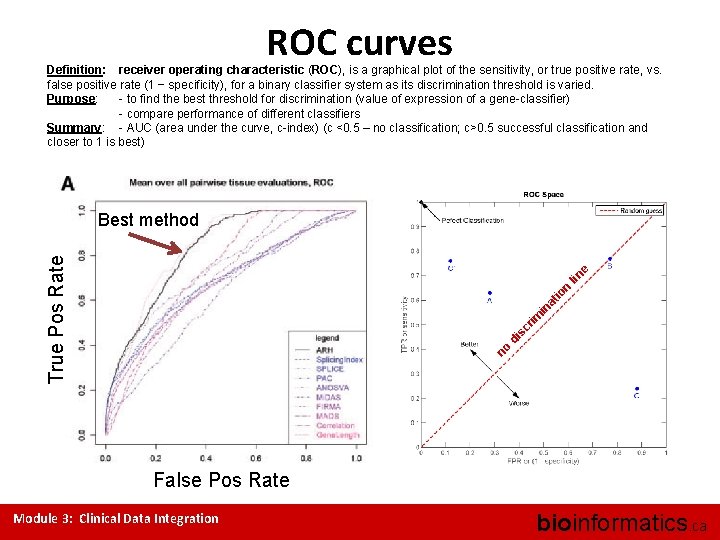
ROC curves Definition: receiver operating characteristic (ROC), is a graphical plot of the sensitivity, or true positive rate, vs. false positive rate (1 − specificity), for a binary classifier system as its discrimination threshold is varied. Purpose: - to find the best threshold for discrimination (value of expression of a gene-classifier) - compare performance of different classifiers Summary: - AUC (area under the curve, c-index) (c <0. 5 – no classification; c>0. 5 successful classification and closer to 1 is best) no di sc rim in at io n lin e True Pos Rate Best method False Pos Rate Module 3: Clinical Data Integration bioinformatics. ca

Survival data – special case Survival times – time to a given end point Survival analysis Module 3: Clinical Data Integration bioinformatics. ca
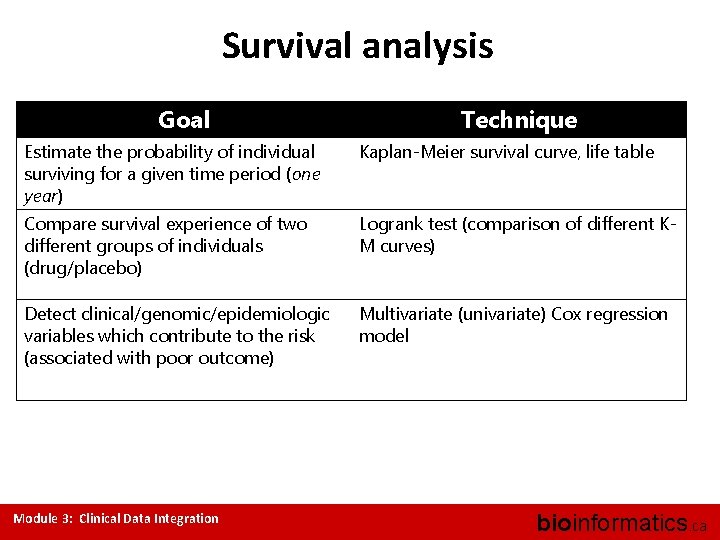
Survival analysis Goal Technique Estimate the probability of individual surviving for a given time period (one year) Kaplan-Meier survival curve, life table Compare survival experience of two different groups of individuals (drug/placebo) Logrank test (comparison of different KM curves) Detect clinical/genomic/epidemiologic variables which contribute to the risk (associated with poor outcome) Multivariate (univariate) Cox regression model Module 3: Clinical Data Integration bioinformatics. ca
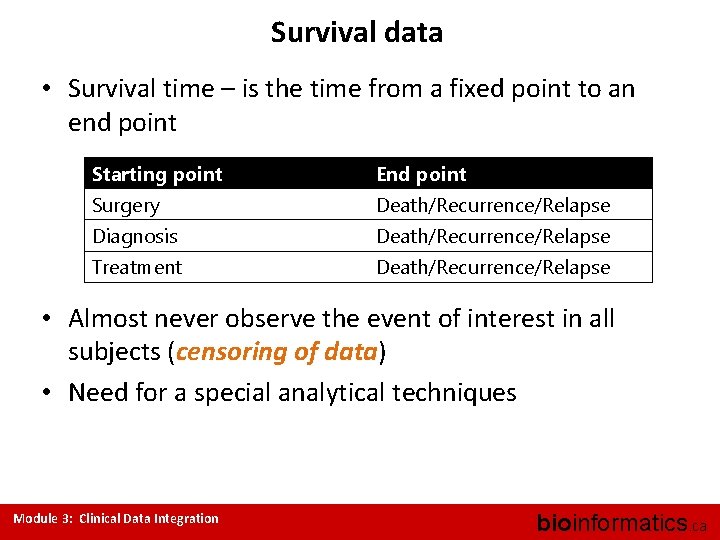
Survival data • Survival time – is the time from a fixed point to an end point Starting point End point Surgery Death/Recurrence/Relapse Diagnosis Death/Recurrence/Relapse Treatment Death/Recurrence/Relapse • Almost never observe the event of interest in all subjects (censoring of data) • Need for a special analytical techniques Module 3: Clinical Data Integration bioinformatics. ca

Censored observations • Arise whenever the dependent variable of interest represents the time to a terminal event, and the duration of the study is limited in time. • Incomplete observation - the event of interest did not occur at the time of the analysis. Event of Interest Censored observation Death of the disease Still alive Survival of marriage Still married Drop-out-time from school Still in school • Type I and II censoring (time fixed/proportion of subjects fixed) • Right and left censoring Module 3: Clinical Data Integration bioinformatics. ca
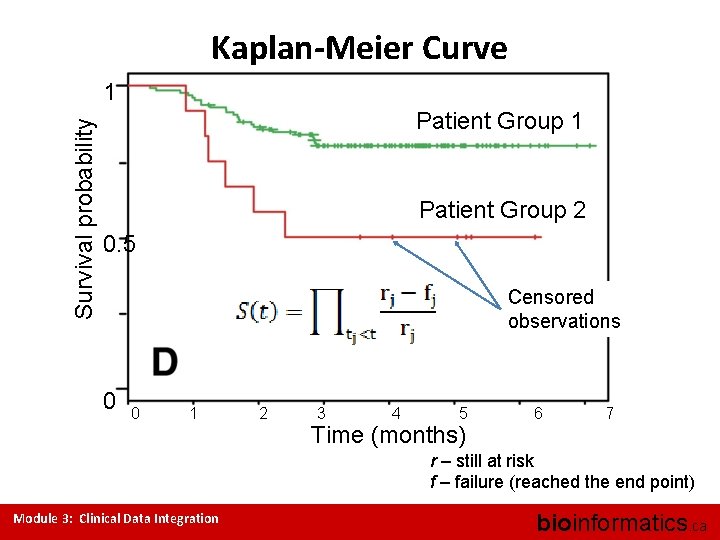
Kaplan-Meier Curve Survival probability 1 Patient Group 2 0. 5 Censored observations 0 0 1 2 3 4 5 Time (months) 6 7 r – still at risk f – failure (reached the end point) Module 3: Clinical Data Integration bioinformatics. ca
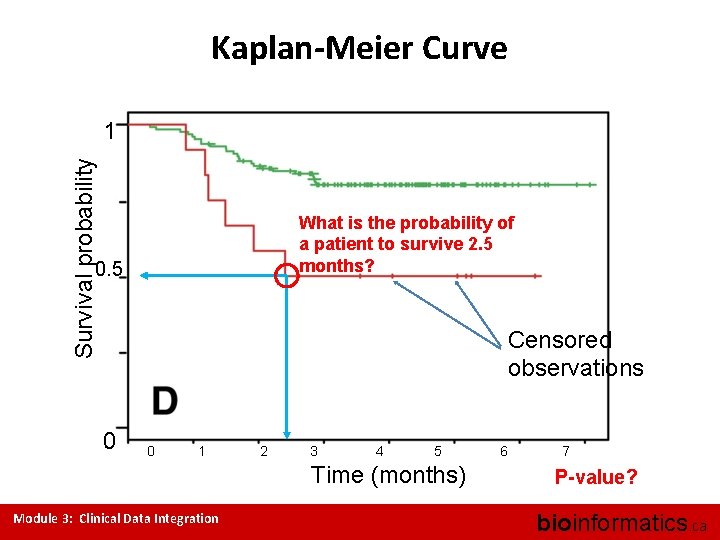
Kaplan-Meier Curve Survival probability 1 What is the probability of a patient to survive 2. 5 months? 0. 5 Censored observations 0 0 1 2 3 4 5 Time (months) Module 3: Clinical Data Integration 6 7 P-value? bioinformatics. ca

Logrank test: compare survival experience of two different groups of individuals Log-rank k - groups of patients to compare O – observed proportion (summed over time points) E – expected proportion (summed over time points) V – variance of (O-E) (summed over time points) Then compare with the χ2 distribution with (k-1) degrees of freedom and get the p-value Module 3: Clinical Data Integration (Doesn’t tell how different) bioinformatics. ca

Hazard ratio compares two groups differing in treatments or prognostic variables etc. Measures relative survival in two groups based on the complete period studied. R=0. 43 – relative risk (hazard) of poor outcome under the condition of group 1 is 43% of that of group 2. R= 2. 0 then the rate of failure in group 1 is twice the rate in the group 2. (tells how different) Module 3: Clinical Data Integration bioinformatics. ca

Cox-proportional hazard model Used to investigate the effect of several variables on survival experience. Multivariable proportional hazards regression model described by D. R. Cox for modeling survival times. It is also called proportional hazards model because it estimates the ratio of the risks (hazard ratio or relative hazard). There are multiple predictor variables (such as prognostic markers whose individual contribution to the outcome is being assessed in the presence of the others) and the outcome variable. Module 3: Clinical Data Integration bioinformatics. ca

Hazard function Prognostic index (PI) • X 1. . . Xp – independent variable of interest • b 1. . . bp – regression coefficients to be estimated • Assumption: the effect of variables is constant over time and additive in a particular scale • (Similarly to K-M) Hazard function is a risk of dying after a given time assuming survival thus far. • Cumulative function • H 0(t) – cumulative baseline or underlying function. • Probability of surviving to time t is S(t) = exp[-H(t)] for every individual with given values of the variables in the model we can estimate this probability. Module 3: Clinical Data Integration bioinformatics. ca
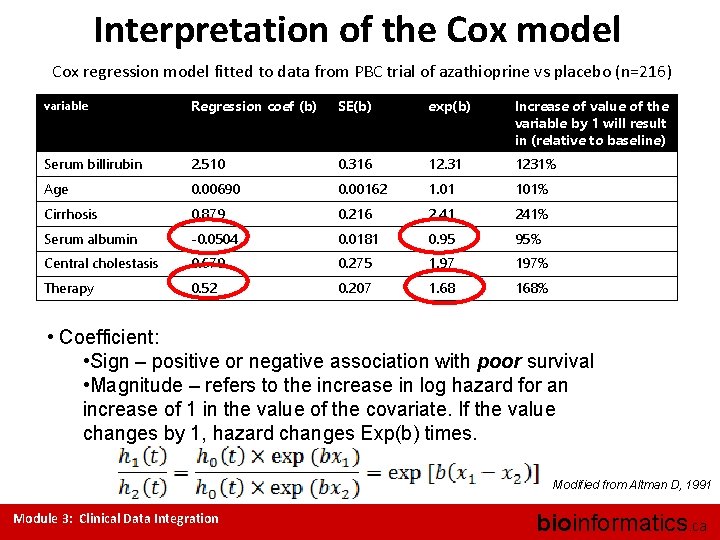
Interpretation of the Cox model Cox regression model fitted to data from PBC trial of azathioprine vs placebo (n=216) variable Regression coef (b) SE(b) exp(b) Increase of value of the variable by 1 will result in (relative to baseline) Serum billirubin 2. 510 0. 316 12. 31 1231% Age 0. 00690 0. 00162 1. 01 101% Cirrhosis 0. 879 0. 216 2. 41 241% Serum albumin -0. 0504 0. 0181 0. 95 95% Central cholestasis 0. 679 0. 275 1. 97 197% Therapy 0. 52 0. 207 1. 68 168% • Coefficient: • Sign – positive or negative association with poor survival • Magnitude – refers to the increase in log hazard for an increase of 1 in the value of the covariate. If the value changes by 1, hazard changes Exp(b) times. Modified from Altman D, 1991 Module 3: Clinical Data Integration bioinformatics. ca
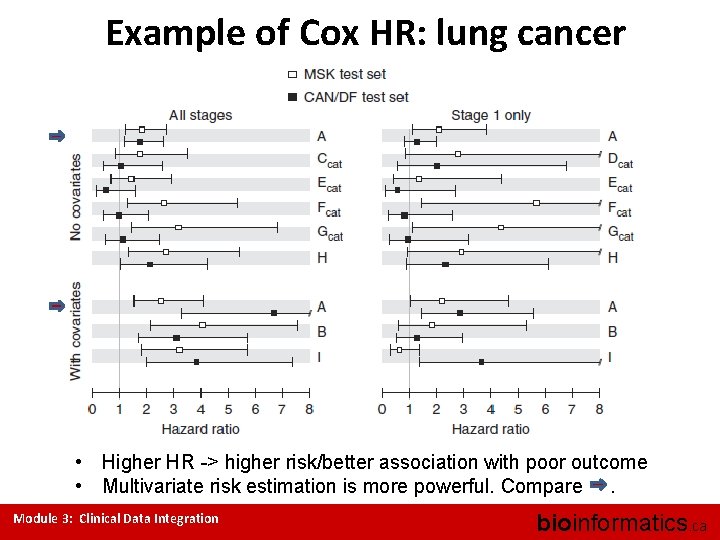
Example of Cox HR: lung cancer • Higher HR -> higher risk/better association with poor outcome • Multivariate risk estimation is more powerful. Compare. Module 3: Clinical Data Integration bioinformatics. ca

We are on a Coffee Break & Networking Session Module 3: Clinical Data Integration bioinformatics. ca
- Slides: 48