Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2

Module 2 From Pre-Processing to Gene-Lists Paul Boutros Microarray Data Analysis June 4 -5, 2012

Pre-Processing What exactly is pre-processing (aka normalization)? Why do we do it? Module 2 bioinformatics. ca
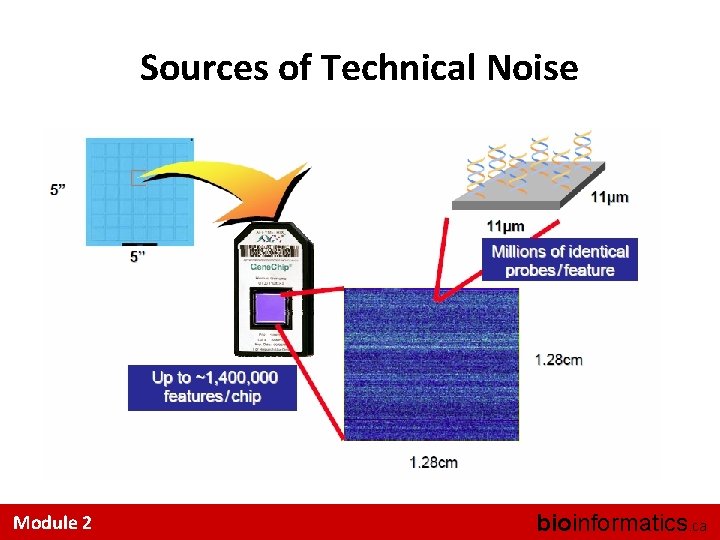
Sources of Technical Noise Where does technical noise come from? Module 2 bioinformatics. ca

More Sources of Technical Noise Module 2 bioinformatics. ca

Any step in the experimental pipeline can introduce artifactual noise • • Array design Array manufacturing Sample quality Sample identity sequence effects? Sample processing Hybridization conditions ozone? Scanner settings Pre-Processing tries to remove these systematic effects Module 2 bioinformatics. ca

Affymetrix Pre-Processing Steps 1. Background Correction 2. Normalization 3. Probe-Specific Adjustment 4. Summarizing multiple Probes into a single Probe. Set Let’s look at two common approaches Module 2 bioinformatics. ca

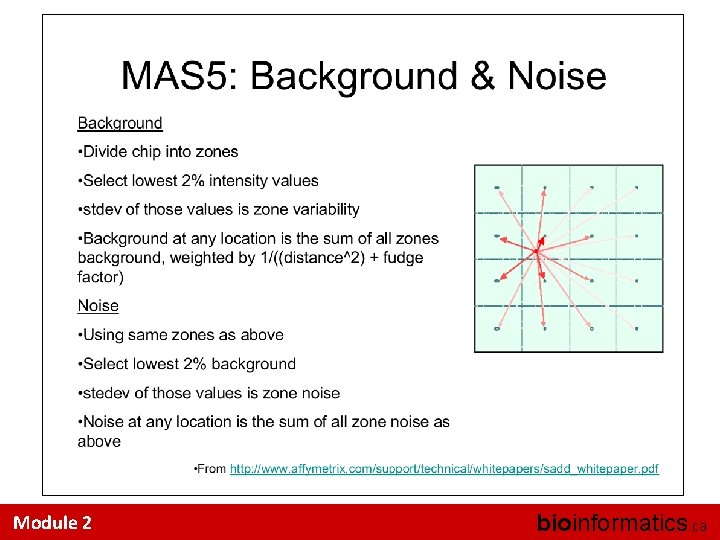
Approach #1: MAS 5 • Affymetrix put significant effort into developing good data pre-processing approaches • MAS 5 was an attempt to develop a “standard” technique for 3’ expression arrays • The flaws of MAS 5 led to an influx of research in this area. • The algorithm is best-described in an Affymetrix white-paper, and is actually quite challenging to reproduce exactly in R. Module 2 bioinformatics. ca

MAS 5 Model Observations = True Signal + Random Noise + Probe Effects Assumptions? Module 2 bioinformatics. ca
Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

What is RMA? RMA = Robust Multi-Array Why do we use a “robust” method? Robust summaries really improve over the standard ones by down weighing outliers and leaving their effects visible in residuals. Why do we use “array”? To put each chip’s values in the context of a set of similar values. Module 2 bioinformatics. ca

What is RMA? It is a log scale linear additive model Assumes all the chips have the same background distribution Does not use the mismatch probe (MM) data from the microarray experiments Why? Module 2 bioinformatics. ca

What is RMA? Mismatch probes (MM) definitely have information - about both signal and noise - but using it without adding more noise is a challenge We should be able to improve the background correction using MM, without having the noise level blow up: topic of current research (GCRMA) Ignoring MM decreases accuracy but increases precision Module 2 bioinformatics. ca

Methodology Quantile Normalization – the goal of this method is to make the distribution of probe intensities for each array in a set of arrays the same. This method is motivated by the idea that a Q-Q plot shows that the distribution of two data vectors is the same if the plot is a straight diagonal line and not the same if it is anything else. Module 2 bioinformatics. ca
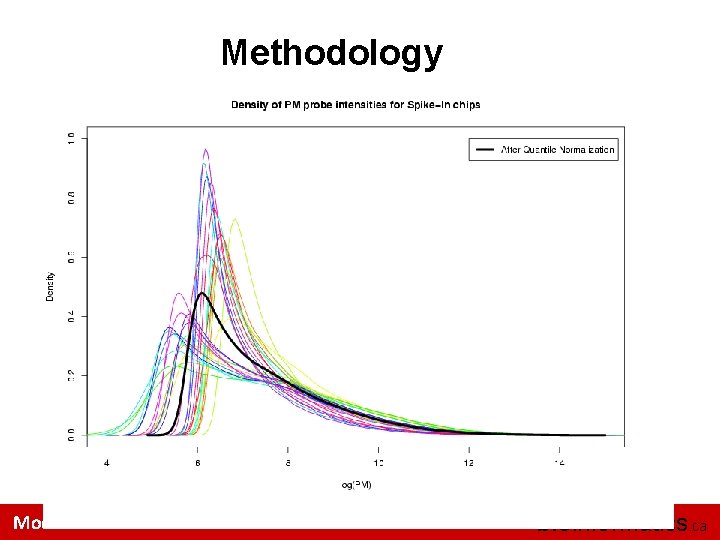
Methodology Module 2 bioinformatics. ca

Methodology Summarization: combining multiple probe intensities of each probeset to produce expression values An additive linear model is fit to the normalized data to obtain an expression measure for each probe on the Gene. Chip Yij = aj + βi + εij Module 2 bioinformatics. ca

Methodology Yij = aj + βi + εij Yij denotes the background-corrected normalized probe value corresponding to the ith Gene. Chip and the jth probe within the probeset [log 2(PM-BG)*ij] aj is the probe affinity jth probe βi is the chip effect for the ith Gene. Chip (log scale expression level) εij is the random error term Module 2 bioinformatics. ca

Methodology Yij = aj + βi + εij Estimate aj ( probe affinity) and βi (chip effect) using a robust method: • Tukey’s Median polish (quick) - fits iteratively, successively removing row and column medians, and accumulating the terms, until the process stabilizes. The residuals are what is left at the end Module 2 bioinformatics. ca

RMA vs MAS 5 • RMA sacrifices accuracy for precision • RMA is generally not appropriate for clinical settings • RMA provides higher sensitivity/specificity in some tests • RMA reduces variance (critical for small-n studies) • RMA is better accepted by journals and reviewers Module 2 bioinformatics. ca

One key detail has been omitted so far: How do we know if our preprocessing actually worked? Module 2 bioinformatics. ca

Outline • Assessing Pre-Processing Results • Univariate Statistical Approaches • Probe. Set Remapping Module 2 bioinformatics. ca

Can we determine how well our preprocessing worked? Or if our data looks good? Module 2 bioinformatics. ca

Let’s See Some “Bad” Data Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Those Three Were From A Spike-In Experiment Done by Affymetrix Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Module 2 bioinformatics. ca

Those Last Three Were From An Experiment We Did On Rat Liver Samples Module 2 bioinformatics. ca

Were Those Bad Samples? • Lots of evident spatial artifacts • But in practice all samples were carried forward into analysis • And validation (RT-PCR) confirmed the overall study results for many genes Module 2 bioinformatics. ca

Eye-ball Assessments Are Hard • A couple of useful tricks: – Look at the distributions • Did quantile normalization work (for RMA)? – Look at the inter-sample correlations • Is one sample a strong outlier? – Look at the 3’ 5’ trend across a Probe. Set I know of no accepted, systematic QA/QC methods Module 2 bioinformatics. ca
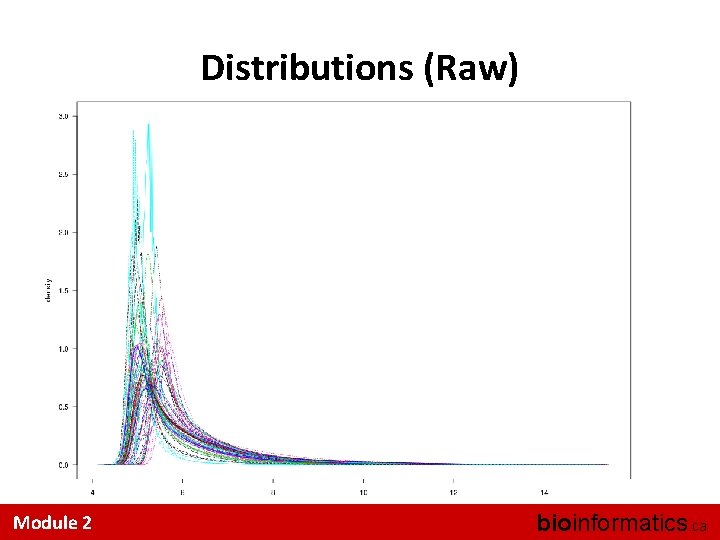
Distributions (Raw) Module 2 bioinformatics. ca
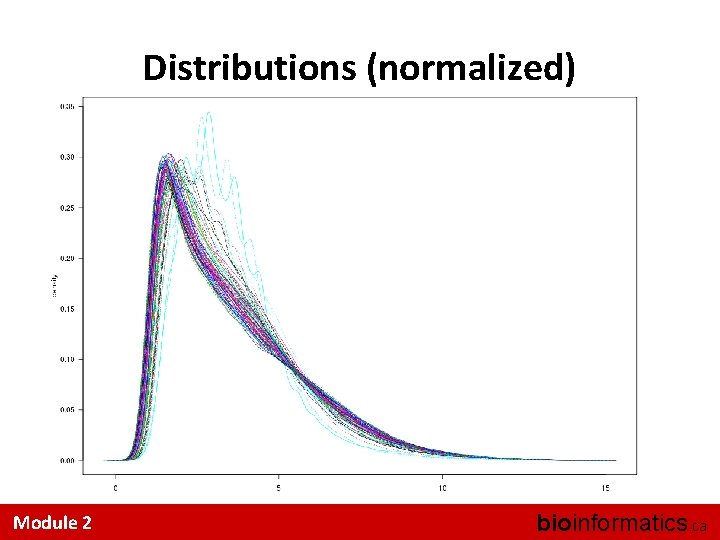
Distributions (normalized) Module 2 bioinformatics. ca
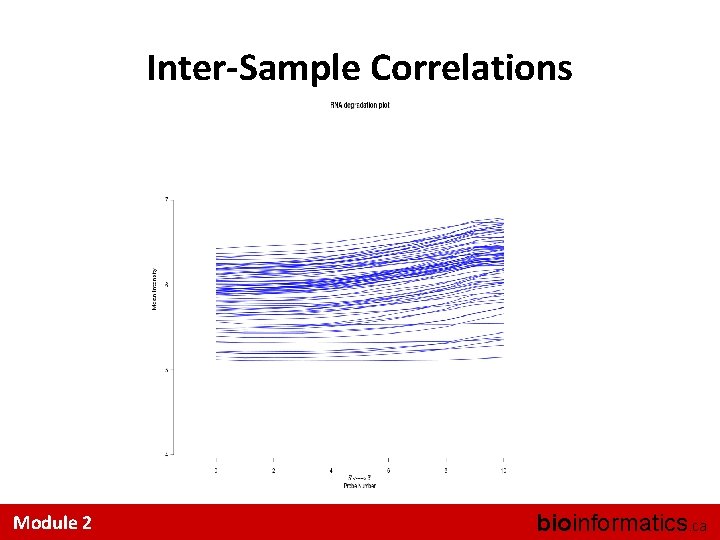
Inter-Sample Correlations Module 2 bioinformatics. ca
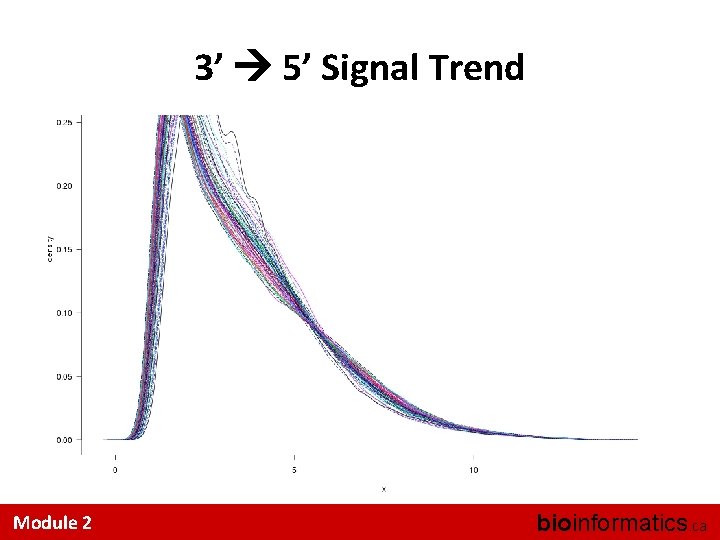
3’ 5’ Signal Trend Module 2 bioinformatics. ca

What Do You Do If You Find a Bad Array? • Repeat it? • Drop the sample? • Include it but account for the “noise” in another way? Module 2 bioinformatics. ca

In This Case • We excluded a series of outlier samples • We believed these samples had been badly degraded because their were derived from FFPE blocks Module 2 bioinformatics. ca
Final Distribution Module 2 bioinformatics. ca
Final Heatmap Module 2 bioinformatics. ca

Outline • Assessing Pre-Processing Results • Univariate Statistical Approaches • Probe. Set Remapping Module 2 bioinformatics. ca

Statistics Reminder Very generally, statistics can be divided into two major branches: Estimation • Measures of centrality • Measures of error Mean, Median, Std-Dev Module 2 Significance-Testing • P-values • Goodness of fit tests T-Test, F-Test, ANOVA bioinformatics. ca

Statistics Reminder #2 What is a P-Value? • • • Imagine we are playing a dice game I roll a 5 You need to roll a 6 to win What is the chance that you will win? 1 in 6 P = 1 / 6 = 0. 167 The probability that you will win is 0. 167 You have a 16. 7% chance of winning I have a 83. 3% chance of winning Module 2 bioinformatics. ca

Significance Testing Questions 1. Are these two groups different? 2. Do these two things synergize? 3. Does treatment affect patient outcome? Module 2 bioinformatics. ca

Distributional Assumptions • A parametric test makes assumptions about the underlying distribution of the data. • A non-parametric test makes no assumptions about the underlying distribution, but may make other assumptions! Module 2 bioinformatics. ca

Two-Sample Analyses • Also called univariate analysis – Requires two conditions – Treating them as a binary variable • E. g. treatment or control • Probably the most common experimental design • Standard approaches: – – Module 2 T-test Wilcoxon rank-sum test T-test variants Permutation tests bioinformatics. ca

T-tests • What are the assumptions of the t-test? • When would you feel comfortable using a t-test? Module 2 bioinformatics. ca

T-Test Alternative: Wilcoxon Rank-Sum • Also called: – U-test – Mann-Whitney (U) test • Some argue that for continuous microarray data there is rarely a good reason to use this test: – Low n: tests of normality are not very powerful – High n: the central limit theorem provides support • If the sample is normal, asymptotic efficiency is 0. 95 Module 2 bioinformatics. ca

T-Test Alternative: Moderated Statistics • A series of highly complex methods based on Bayesian statistical methodologies • Gordon Smyth’s limma R package is by far the most widely used implementation of this technique This term is “shrunk” by borrowing power across all genes. This increases effective power. Module 2 bioinformatics. ca

T-Test Alternative: Permutation Tests • SAM is the classic method – Most people suggest not using SAM today • Empirically estimate the null distribution Iterate Start with many samples Module 2 Randomly Sample bioinformatics. ca

Problems with Significance Testing • What happens if there are NO changes? • Imagine: – You analyzed 1, 000 clinical samples – 20, 000 genes in the genome – P < 0. 05 • What if… somebody comes and randomizes all your data? Module 2 bioinformatics. ca

You had a lot of Data 20, 000 genes / array 1, 000 patients 20, 000 data points All Randomized Genes are mixed up together Patients are mixed together What happens if you analyze this data? There should be NO real hits anymore! Module 2 bioinformatics. ca

What will you actually find? Array: 20, 000 genes Threshold: p < 0. 05 20, 000 x 0. 05 = 1000 False Positives This is called “multiple testing”. There is a solution Module 2 bioinformatics. ca
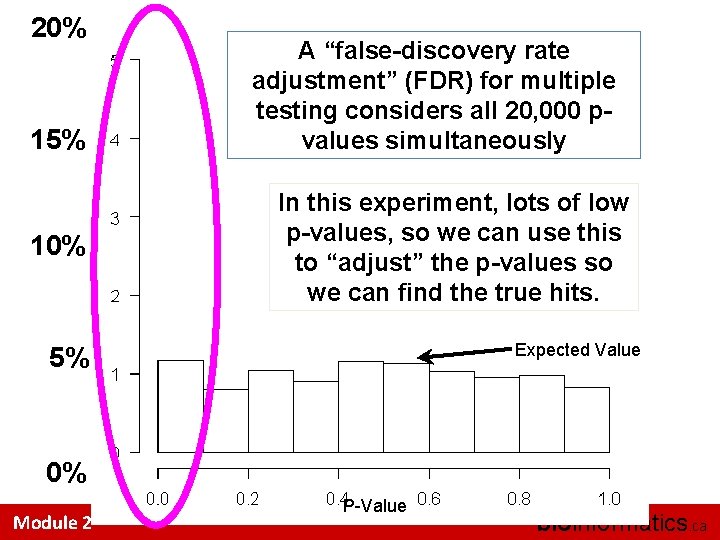
20% 15% A “false-discovery rate adjustment” (FDR) for multiple testing considers all 20, 000 pvalues simultaneously 10% In this experiment, lots of low p-values, so we can use this to “adjust” the p-values so we can find the true hits. 5% Expected Value 0% Module 2 P-Value bioinformatics. ca
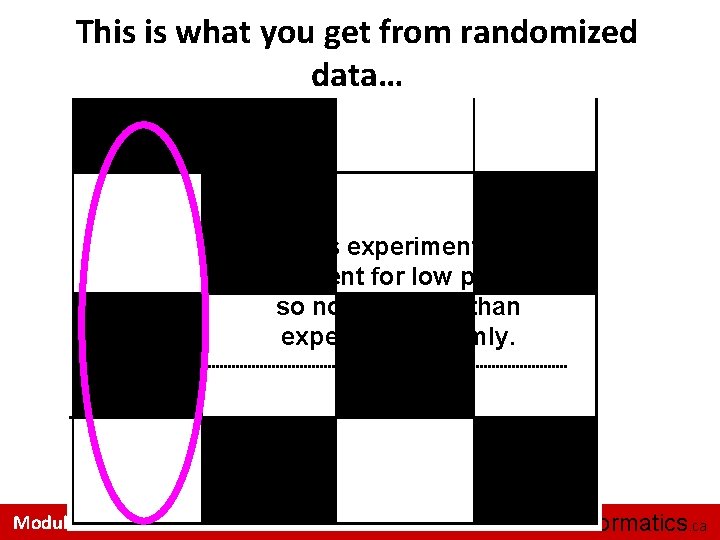
This is what you get from randomized data… In this experiment, NO enrichment for low p-values, so no more hits than expected randomly. Module 2 bioinformatics. ca

Outline • Pre-Processing Matters! • Assessing Pre-Processing Results • Univariate Statistical Approaches • Probe. Set Remapping Module 2 bioinformatics. ca

Arrays Can Become Outdated • Gene definitions change • The reference genome sequence gets finished • Novel splice variants are found • Errors are made in the initial design and remain present in all arrays made Module 2 bioinformatics. ca
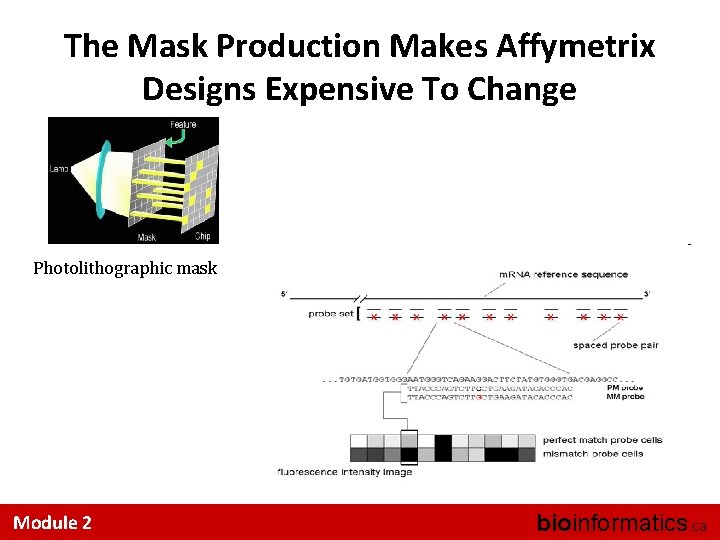
The Mask Production Makes Affymetrix Designs Expensive To Change Photolithographic mask Module 2 bioinformatics. ca
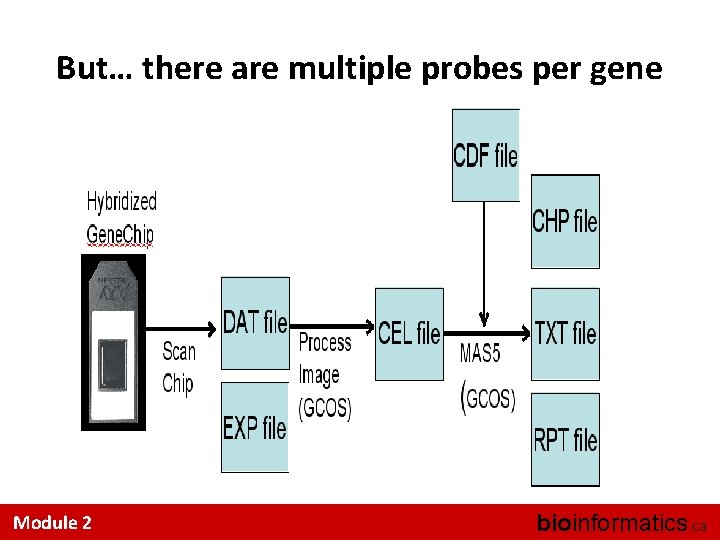
But… there are multiple probes per gene Module 2 bioinformatics. ca

We Can Change Those Mappings! Hybridized Chip Module 2 bioinformatics. ca

CDF File • Chip Definition File • This file maps Probes (positions) into Probe. Sets • We can update those mappings – Ignore deprecated or cross-hybridizing probes – Merge multiple probes that recognize the same gene – Account for entirely new genes that were not known at the time of array-design Module 2 bioinformatics. ca

Sequence Mappings Are Slow • Requires aligning millions of 25 bp probes against the transcriptome and identifying the best match for each • Fortunately, other groups have done this for us, and regularly update their mappings Module 2 bioinformatics. ca
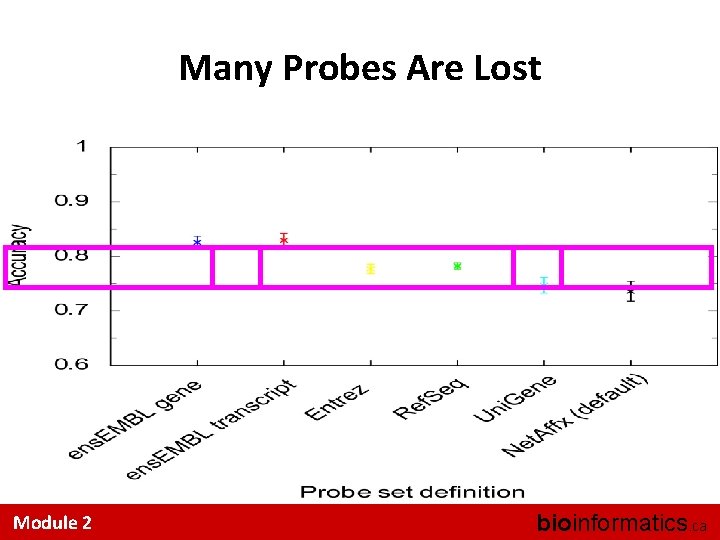
Many Probes Are Lost Module 2 bioinformatics. ca
But There Is Also A Major Benefit Increased validation rates using RT-PCR (~10%) Module 2 Sandberg et al BMC Bioinformatics 2007 bioinformatics. ca

After the break • Learning how to make QA/QC plots in R • Compare univariate statistical analysis techniques • Apply an alternative Probe. Set remapping • Contrast the effects of pre-processing Module 2 bioinformatics. ca

We are on a Coffee Break & Networking Session Module 2 bioinformatics. ca
- Slides: 71