Association Pattern Analysis Applications in Bioinformatics Vipin Kumar

Association Pattern Analysis – Applications in Bioinformatics Vipin Kumar University of Minnesota kumar@cs. umn. edu www. cs. umn. edu/~kumar Team Members: Michael Steinbach, Rohit Gupta, Hui Xiong, Gaurav Pandey, Blayne Field, Meenal Chhabra, Beth Zirbes Research supported by NSF, IBM

Data Mining for Bioinformatics § Recent technological advances are helping to generate large amounts of both medical and genomic data • • High-throughput experiments/techniques - Gene and protein sequences - Gene-expression data - Biological networks and phylogenetic profiles Electronic Medical Records - IBM-Mayo clinic partnership has created a DB of 5 million patients - Single Nucleotides Polymorphisms (SNPs) § Data mining offers potential solution for analysis of large-scale data • • • Automated analysis of patients history for customized treatment Prediction of the functions of anonymous genes Identification of putative binding sites in protein structures for drugs/chemicals discovery August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 2 Protein Interaction Network
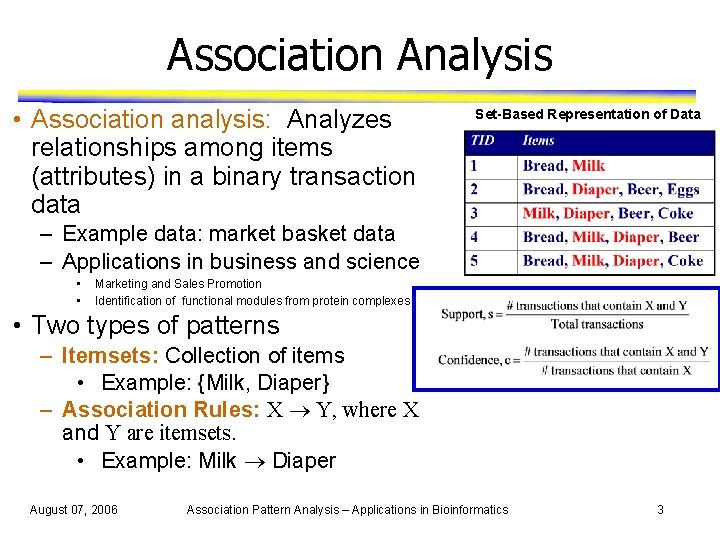
Association Analysis • Association analysis: Analyzes relationships among items (attributes) in a binary transaction data Set-Based Representation of Data – Example data: market basket data – Applications in business and science • • Marketing and Sales Promotion Identification of functional modules from protein complexes • Two types of patterns – Itemsets: Collection of items • Example: {Milk, Diaper} – Association Rules: X Y, where X and Y are itemsets. • Example: Milk Diaper August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 3

I. Application of Association Analysis: Identification of Protein Function Modules § Proteins usually do not function in isolation. They interact with other proteins either in pairs or as components of large complexes § Protein complexes can be determined using large scale experimental studies § Functional module is a group of proteins that is involved in common elementary biological function § Association analysis techniques can be used for identification of functional modules from a collection of protein complexes Protein Complex Data August 07, 2006 Protein Complexes Proteins c 1 p 1, p 2 c 2 p 1, p 3, p 4, p 5 c 3 p 2, p 3, p 4, p 6 Association Pattern Analysis – Applications in Bioinformatics 4

II. Application of Association Analysis: Personalized Medicine • Given: Patient data set that records – Phenotypic characteristics – Genetic characteristics (SNPs) – Disease • Objective: Find relationships between disease and medical and genomic characteristics • Association analysis can be used to find groups of phenotypic and genetic characteristics that are highly associated with disease Recently started project in collaboration with IBM Rochester – Drew Flaada, Fred Kullack, Tim Mullins, Carl Oberto August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 5
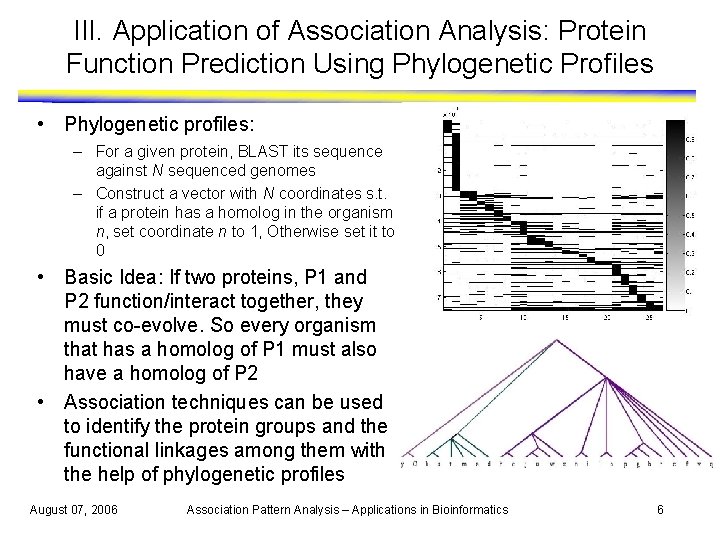
III. Application of Association Analysis: Protein Function Prediction Using Phylogenetic Profiles • Phylogenetic profiles: – For a given protein, BLAST its sequence against N sequenced genomes – Construct a vector with N coordinates s. t. if a protein has a homolog in the organism n, set coordinate n to 1, Otherwise set it to 0 • Basic Idea: If two proteins, P 1 and P 2 function/interact together, they must co-evolve. So every organism that has a homolog of P 1 must also have a homolog of P 2 • Association techniques can be used to identify the protein groups and the functional linkages among them with the help of phylogenetic profiles August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 6
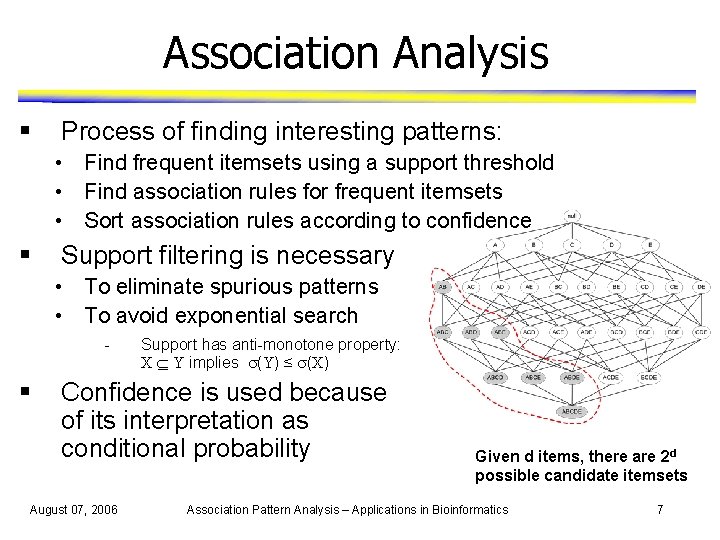
Association Analysis § Process of finding interesting patterns: • Find frequent itemsets using a support threshold • Find association rules for frequent itemsets • Sort association rules according to confidence § Support filtering is necessary • To eliminate spurious patterns • To avoid exponential search - § Support has anti-monotone property: X Y implies (Y) ≤ (X) Confidence is used because of its interpretation as conditional probability August 07, 2006 Given d items, there are 2 d possible candidate itemsets Association Pattern Analysis – Applications in Bioinformatics 7
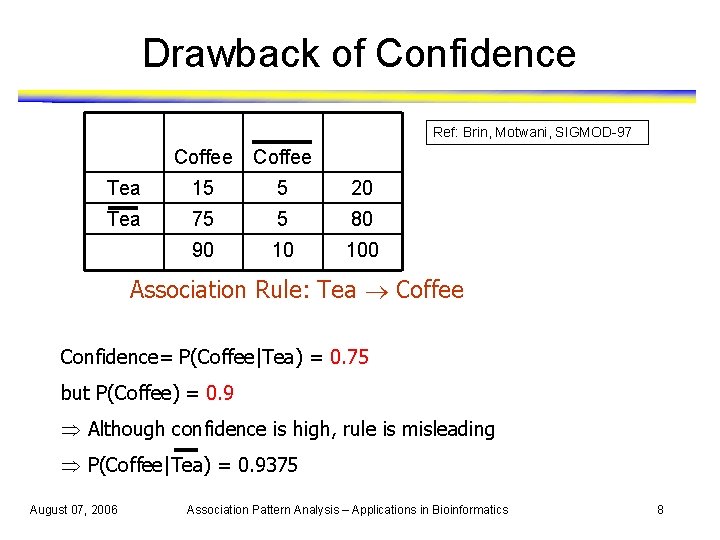
Drawback of Confidence Ref: Brin, Motwani, SIGMOD-97 Coffee Tea 15 5 20 Tea 75 5 80 90 10 100 Association Rule: Tea Coffee Confidence= P(Coffee|Tea) = 0. 75 but P(Coffee) = 0. 9 Þ Although confidence is high, rule is misleading Þ P(Coffee|Tea) = 0. 9375 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 8

There are lots of measures proposed in the literature August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 9
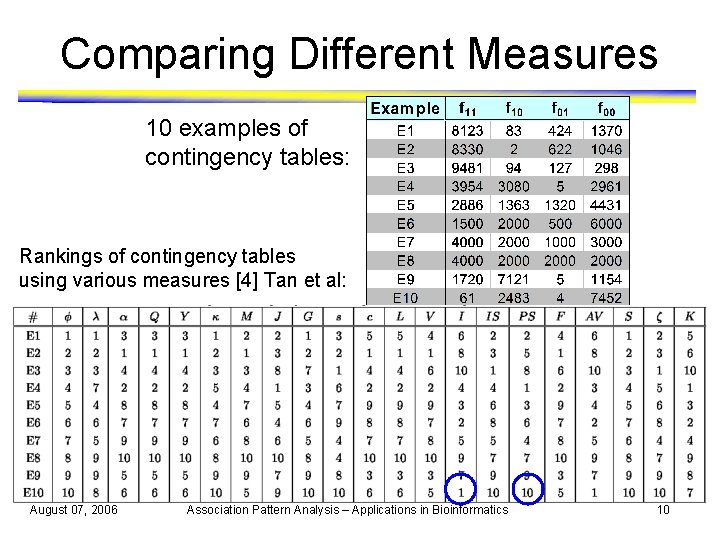
Comparing Different Measures 10 examples of contingency tables: Rankings of contingency tables using various measures [4] Tan et al: August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 10

Hyperclique Pattern § The h-confidence of a pattern P = {i 1, i 2, …, im} § Example: For a pattern P = {A, B, C}, assume that: • supp({A}) = 0. 1, supp({B}) = 0. 1, supp({C}) = 0. 06, supp({A, B, C}) = 0. 06 • hconf(A, B, C) = [supp({A, B, C})] / [max{supp({A}), supp({B}), supp({C})}] = 0. 06/0. 1 = 0. 6 § A pattern P is a hyperclique pattern if hconf(P)>=hc, where hc is a user specified minimum h-confidence threshold August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 11

Alternate Equivalent Definitions of hconfidence § Given a pattern P = {i 1, i 2, …, im} • Definition: All-Confidence Measure Omiecinski – TKDE 2003 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 12

Properties of Hyperclique Pattern § Anti-monotone § High Affinity Property • High h-confidence implies tight coupling amongst all items in the pattern § Cross support property • eliminates patterns involving items that have very different support levels § Magnitude of relationship consistent with many other measure • Jaccard, Correlation, Cosine August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 13
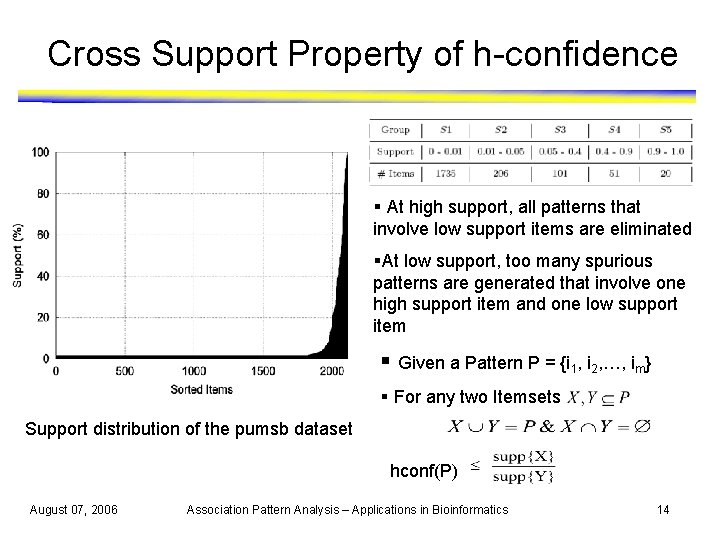
Cross Support Property of h-confidence § At high support, all patterns that involve low support items are eliminated §At low support, too many spurious patterns are generated that involve one high support item and one low support item § Given a Pattern P = {i 1, i 2, …, im} § For any two Itemsets Support distribution of the pumsb dataset hconf(P) August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 14

Consistency with other Measures § Jaccard • If an item set P = {i 1, i 2} is a size-2 hyperclique pattern, then § Correlation • Let S be a set of items and hc be the minimum h-confidence, we can form two groups: Then, any size-2 hyperclique pattern P = {A, B} has a positive correlation in each of the following cases: § Cosine • If an item set P = {i 1, i 2} is a size-2 hyperclique pattern, then August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 15
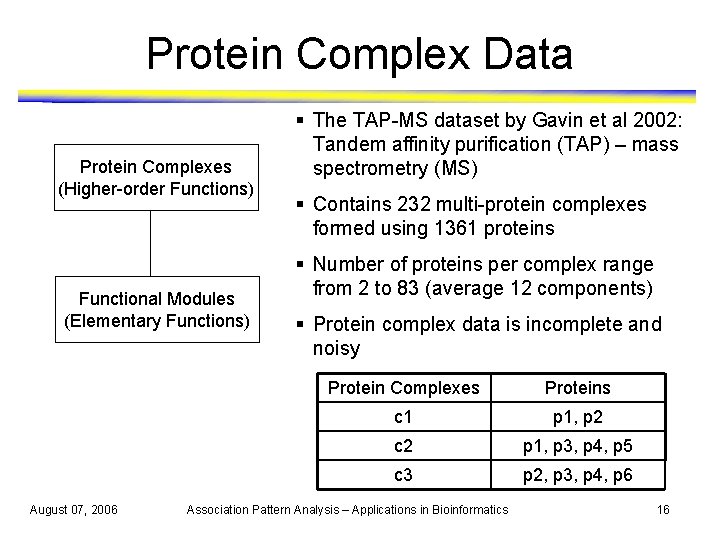
Protein Complex Data Protein Complexes (Higher-order Functions) Functional Modules (Elementary Functions) August 07, 2006 § The TAP-MS dataset by Gavin et al 2002: Tandem affinity purification (TAP) – mass spectrometry (MS) § Contains 232 multi-protein complexes formed using 1361 proteins § Number of proteins per complex range from 2 to 83 (average 12 components) § Protein complex data is incomplete and noisy Protein Complexes Proteins c 1 p 1, p 2 c 2 p 1, p 3, p 4, p 5 c 3 p 2, p 3, p 4, p 6 Association Pattern Analysis – Applications in Bioinformatics 16
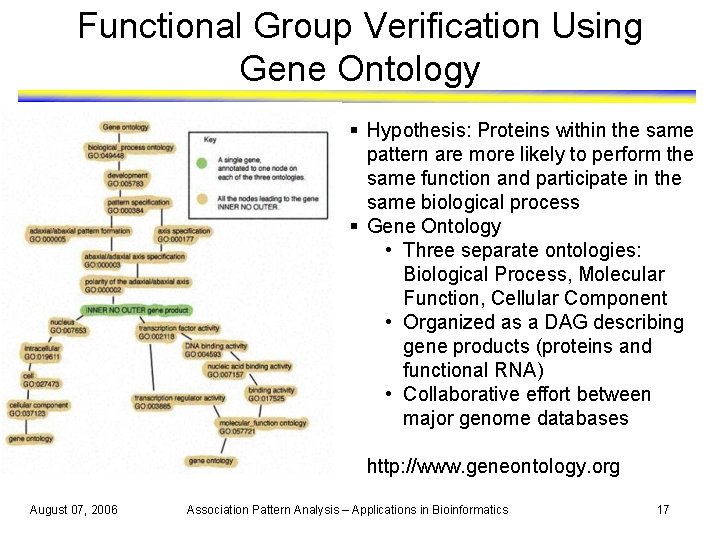
Functional Group Verification Using Gene Ontology § Hypothesis: Proteins within the same pattern are more likely to perform the same function and participate in the same biological process § Gene Ontology • Three separate ontologies: Biological Process, Molecular Function, Cellular Component • Organized as a DAG describing gene products (proteins and functional RNA) • Collaborative effort between major genome databases http: //www. geneontology. org August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 17

Hyperclique Patterns from Protein Complex Data § List of maximal hyperclique patterns at a support threshold 2 and an h-confidence threshold 60%. [1] Xiong et al. (Detailed results are at http: //cimic. rutgers. edu/~hui/pfm. html) 2 Tif 4631 2 Cdc 33 Snp 1 2 YHR 020 W Mir 1 2 Cka 1 Ckb 1 2 Ckb 2 Cka 2 2 Cop 1 Sec 27 2 Erb 1 YER 006 W 2 Ilv 1 YGL 245 W 2 Ilv 1 Sec 27 2 Ioc 3 Rsc 8 2 Isw 2 Itc 1 2 Kre 33 YJL 109 C 2 Kre 33 YPL 012 W 2 Mot 1 Isw 1 2 Npl 3 Smd 3 2 Npl 6 Isw 2 2 Npl 6 Mot 1 2 Rad 52 Rfa 1 2 Rpc 40 Rsc 8 2 Rrp 4 Dis 3 2 Rrp 40 Rrp 46 2 Cbf 5 Kre 33 3 YGL 128 C Clf 1 YLR 424 W 3 Cka 2 Cka 1 Ckb 1 3 Has 1 Nop 12 Sik 1 3 Hrr 25 Enp 1 YDL 060 W 3 Hrr 25 Swi 3 Snf 2 August 07, 2006 3 Kre 35 Nog 1 YGR 103 W 3 Krr 1 Cbf 5 Kre 33 6 Dim 1 Ltv 1 YOR 056 C YOR 145 C Enp 1 YDL 060 W 3 Nab 3 Nrd 1 YML 117 W 6 Pre 3 Pre 2 Pre 4 Pre 5 Pre 8 Pup 3 3 Nog 1 YGR 103 W YER 006 W 7 Clf 1 Lea 1 Rse 1 YLR 424 W Prp 46 Smd 2 Snu 114 3 Bms 1 Sik 1 Rpp 2 b 7 Pre 1 Pre 7 Pre 2 Pre 4 Pre 5 Pre 8 Pup 3 3 Rpn 10 Rpt 3 Rpt 6 7 Blm 3 Pre 10 Pre 2 Pre 4 Pre 5 Pre 8 Pup 3 3 Rpn 11 Rpn 12 Rpn 8 8 Clf 1 Prp 4 Smb 1 Snu 66 YLR 424 W Prp 46 Smd 2 Snu 114 3 Rpn 12 Rpn 8 Rpn 10 8 Pre 2 Pre 4 Pre 5 Pre 8 Pup 3 Pre 6 Pre 9 Scl 1 3 Rpn 9 Rpt 3 Rpt 5 10 Cdc 33 Dib 1 Lsm 4 Prp 31 Prp 6 Clf 1 Prp 4 Smb 1 Snu 66 YLR 424 W 3 Rpn 9 Rpt 3 Rpt 6 12 Dib 1 Lsm 4 Prp 31 Prp 6 Clf 1 Prp 4 Smb 1 Snu 66 YLR 424 W Prp 46 Smd 2 Snu 114 3 Brx 1 Sik 1 YOR 206 W 3 Sik 1 Kre 33 YJL 109 C 3 Taf 145 Taf 90 Taf 60 4 Fyv 14 Krr 1 Sik 1 YLR 409 C 4 Mrpl 35 Mrpl 8 YML 025 C Mrpl 3 4 Rpn 12 Rpn 8 Rpt 3 Rpt 6 6 Luc 7 Rse 1 Smd 3 Snp 1 Snu 71 Smd 2 12 Emg 1 Imp 3 Imp 4 Kre 31 Mpp 10 Nop 14 Sof 1 YMR 093 W YPR 144 C Krr 1 YDR 449 C Enp 1 13 Ecm 2 Hsh 155 Prp 19 Prp 21 Snt 309 YDL 209 C Clf 1 Lea 1 Rse 1 YLR 424 W Prp 46 Smd 2 Snu 114 13 Brr 1 Mud 1 Prp 39 Prp 40 Prp 42 Smd 1 Snu 56 Luc 7 Rse 1 Smd 3 Snp 1 Snu 71 Smd 2 39 Cus 1 Msl 1 Prp 3 Prp 9 Sme 1 Smx 2 Smx 3 Yhc 1 YJR 084 W Brr 1 Dib 1 Ecm 2 Hsh 155 Lsm 4 Mud 1 Prp 19 Prp 21 Prp 39 5 Ada 2 Gcn 5 Rpo 21 Spt 7 Taf 60 Prp 42 Prp 6 Smd 1 Snt 309 Snu 56 Srb 2 YDL 209 C Clf 1 Lea 1 6 YLR 033 W Ioc 3 Npl 6 Rsc 2 Itc 1 Rpc 40 Luc 7 Prp 4 Rse 1 Smb 1 Smd 3 Snp 1 Snu 66 Snu 71 YLR 424 W 5 Rpn 6 Rpt 2 Rpn 12 Rpn 3 Rpn 8 Association Pattern Analysis – Applications in Bioinformatics 18

Summary § Number of hypercliques: • Size-2: 22, Size-3: 18, Size-4: 3, Size-5: 2 • Size-6: 4, Size-7: 3, Size-8: 2, Size-10: 1 • Size-12: 2, Size-13: 2, Size-39: 1 § In most cases, proteins identified as hypercliques found to be functionally coherent and part of same biological process evaluated using GO hierarchies August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 19
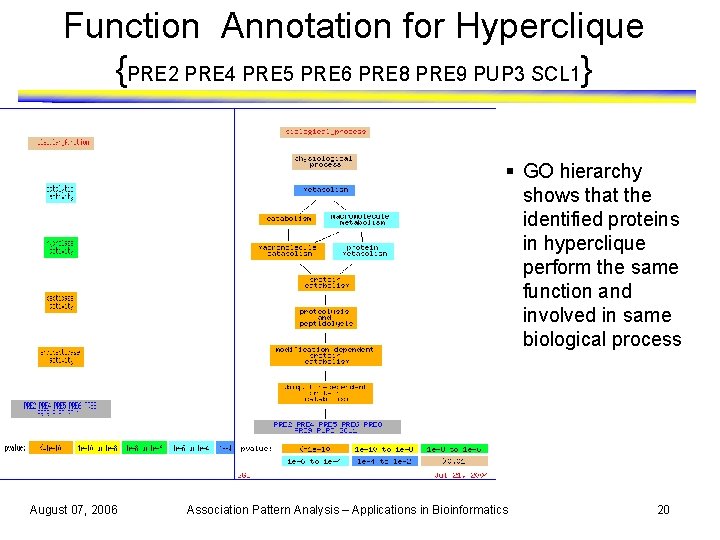
Function Annotation for Hyperclique {PRE 2 PRE 4 PRE 5 PRE 6 PRE 8 PRE 9 PUP 3 SCL 1} § GO hierarchy shows that the identified proteins in hyperclique perform the same function and involved in same biological process August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 20
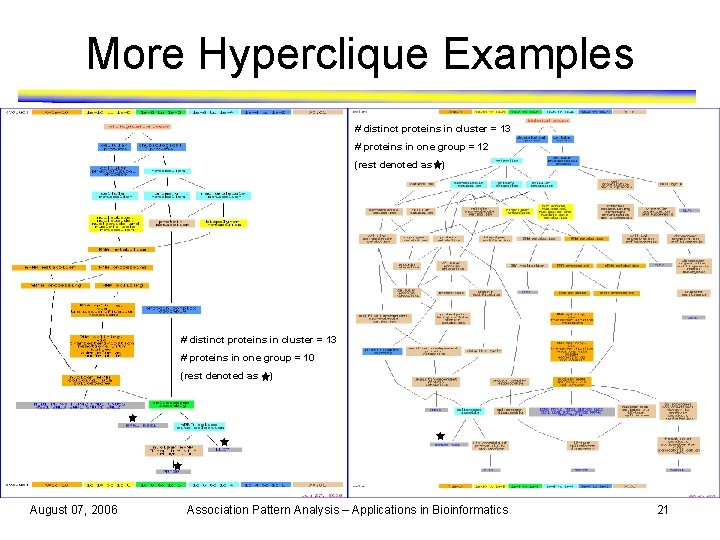
More Hyperclique Examples # distinct proteins in cluster = 13 # proteins in one group = 12 (rest denoted as ) # distinct proteins in cluster = 13 # proteins in one group = 10 (rest denoted as August 07, 2006 ) Association Pattern Analysis – Applications in Bioinformatics 21
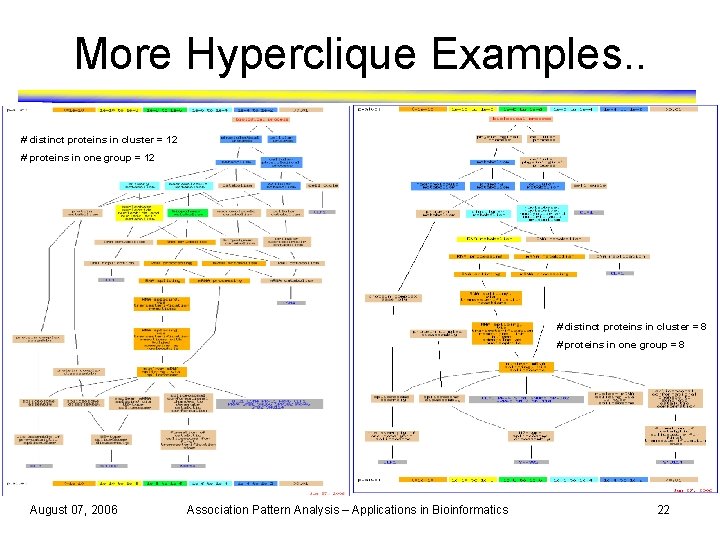
More Hyperclique Examples. . # distinct proteins in cluster = 12 # proteins in one group = 12 # distinct proteins in cluster = 8 # proteins in one group = 8 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 22
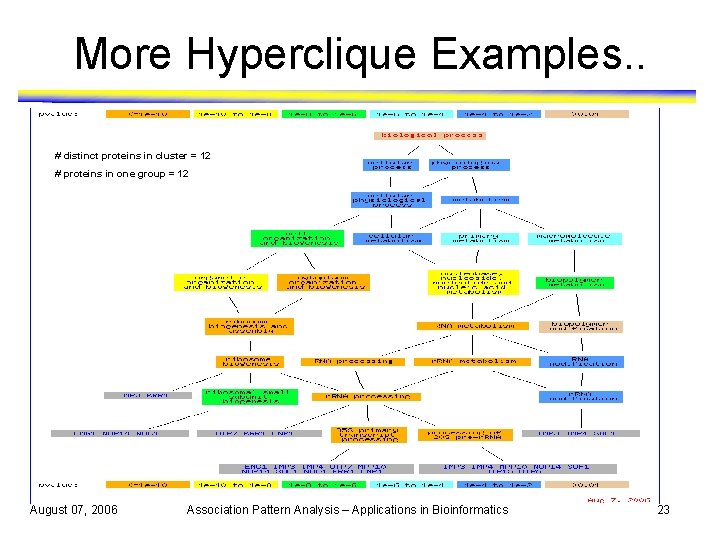
More Hyperclique Examples. . # distinct proteins in cluster = 12 # proteins in one group = 12 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 23
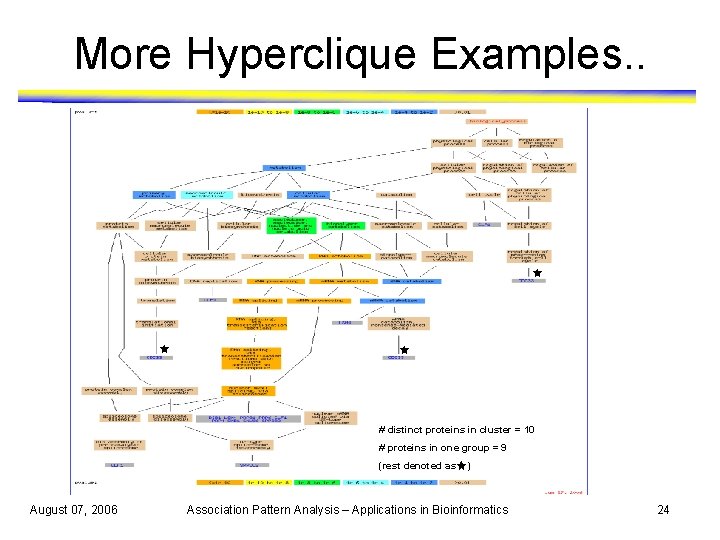
More Hyperclique Examples. . # distinct proteins in cluster = 10 # proteins in one group = 9 (rest denoted as August 07, 2006 ) Association Pattern Analysis – Applications in Bioinformatics 24

More Hyperclique Examples. . # distinct proteins in cluster = 39 # proteins in one group = 32 # proteins at node ‘m. RNA splicing’ = 37 § Only two Proteins SRB 2 and ECM 2 involved in cellular process and development got clustered together with group of proteins involved in physiological process § It is observed that 37 proteins out of 39 annotated proteins are responsible for same molecular function, m. RNA splicing via spliceosome August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 25
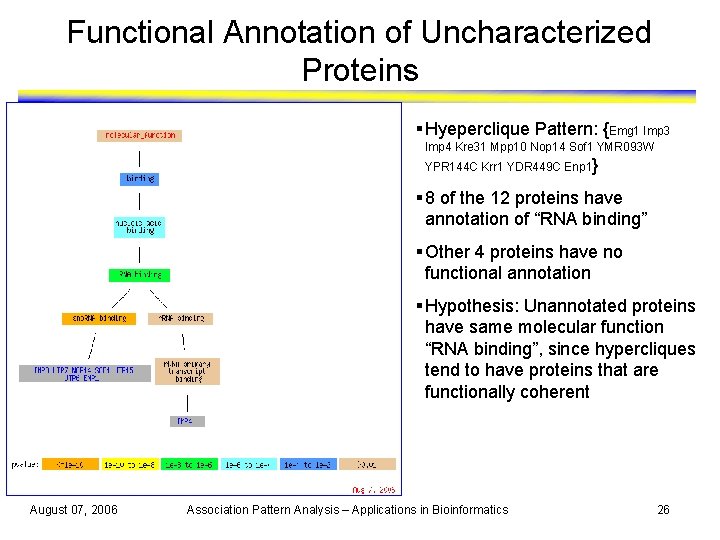
Functional Annotation of Uncharacterized Proteins § Hyeperclique Pattern: {Emg 1 Imp 3 Imp 4 Kre 31 Mpp 10 Nop 14 Sof 1 YMR 093 W YPR 144 C Krr 1 YDR 449 C Enp 1} § 8 of the 12 proteins have annotation of “RNA binding” § Other 4 proteins have no functional annotation § Hypothesis: Unannotated proteins have same molecular function “RNA binding”, since hypercliques tend to have proteins that are functionally coherent August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 26

Identification of Functional Modules Using Frequent Itemset-based Approach § Closed frequent itemset-based approach produces over 500 patterns of size 2 or more with support threshold of 2 § Number of patterns • for (h-confidence < 0. 20) = 198 • Generally very poor • for (0. 20 <= h-confidence < 0. 50) = 246 • moderate quality • for (h-confidence >= 0. 50) = 65 • Generally very good § Proteins in large size patterns (with high h-confidence) are found to be better functionally related than even proteins in small size patterns (with less hconfidence) § 2 examples illustrating this are shown August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 27
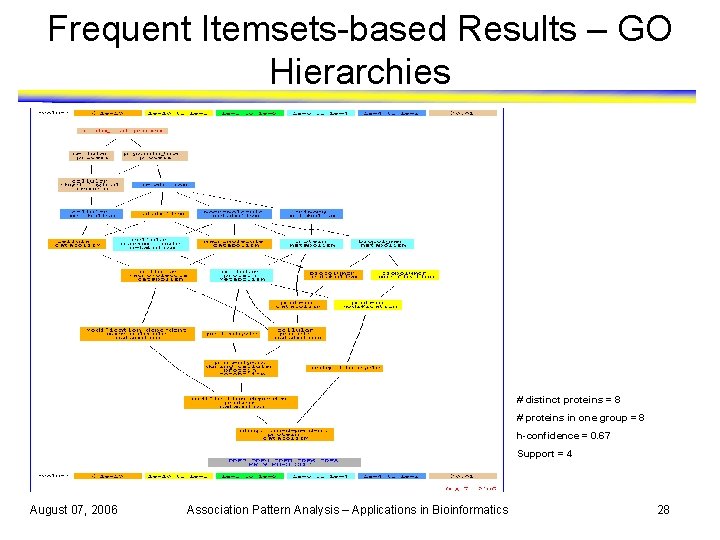
Frequent Itemsets-based Results – GO Hierarchies # distinct proteins = 8 # proteins in one group = 8 h-confidence = 0. 67 Support = 4 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 28
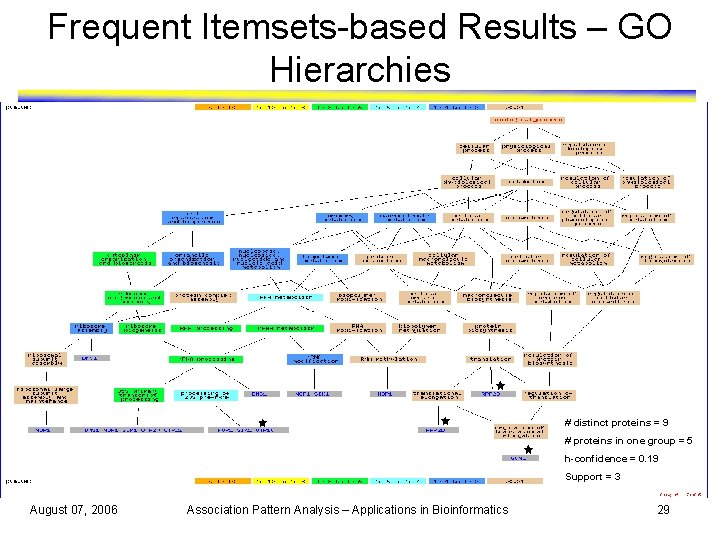
Frequent Itemsets-based Results – GO Hierarchies # distinct proteins = 9 # proteins in one group = 5 h-confidence = 0. 19 Support = 3 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 29

Clustering of Protein Complex Data § Clustering software CLUTO (http: //glaros. dtc. umn. edu/gkhome/views/cluto) is used to cluster the proteins in groups • Repeated bisection method is used as the base method for clustering • Cosine similarity measure is used to find similarity between proteins § Parameter to define the maximum number of clusters that could be obtained is set to 100 § Best clusters (as measured by internal similarity) are usually the candidates for functional modules August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 30

Clustering Results Summary § Clusters with high internal similarity (as ranked by Cluto program) and relatively small sizes are found to be functionally coherent using GO hierarchies § It is found that large clusters with relatively low internal similarity have proteins with multiple function annotations § Few examples to illustrate this are shown August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 31
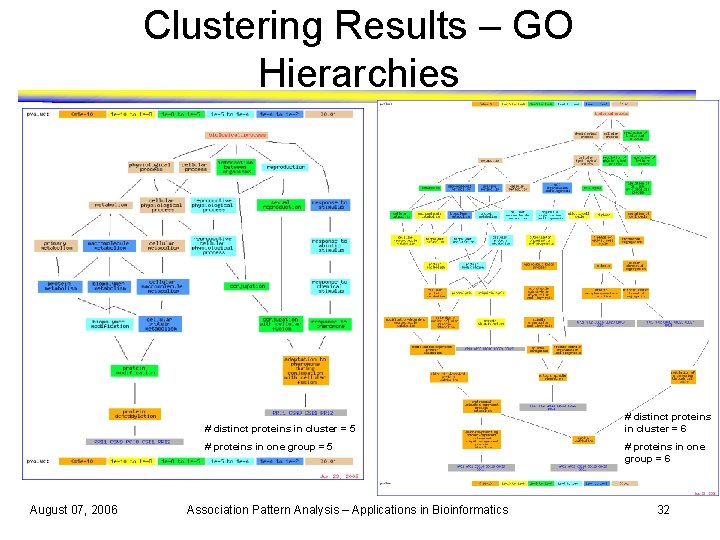
Clustering Results – GO Hierarchies # distinct proteins in cluster = 5 # proteins in one group = 5 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics # distinct proteins in cluster = 6 # proteins in one group = 6 32
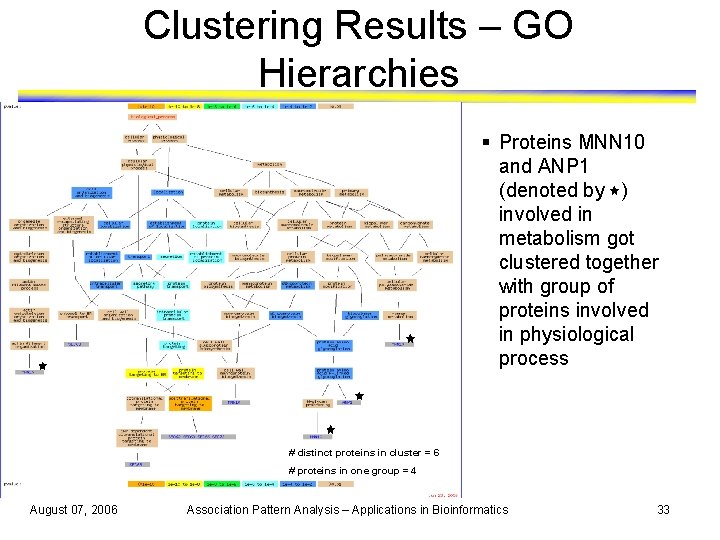
Clustering Results – GO Hierarchies § Proteins MNN 10 and ANP 1 (denoted by ) involved in metabolism got clustered together with group of proteins involved in physiological process # distinct proteins in cluster = 6 # proteins in one group = 4 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 33
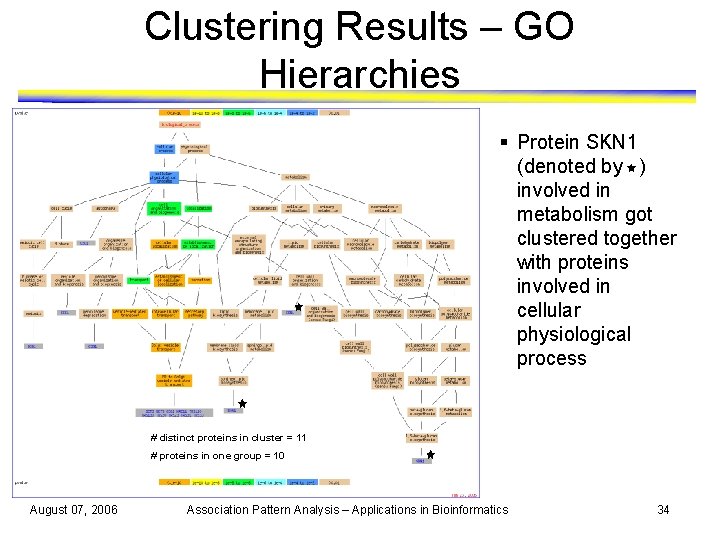
Clustering Results – GO Hierarchies § Protein SKN 1 (denoted by ) involved in metabolism got clustered together with proteins involved in cellular physiological process # distinct proteins in cluster = 11 # proteins in one group = 10 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 34
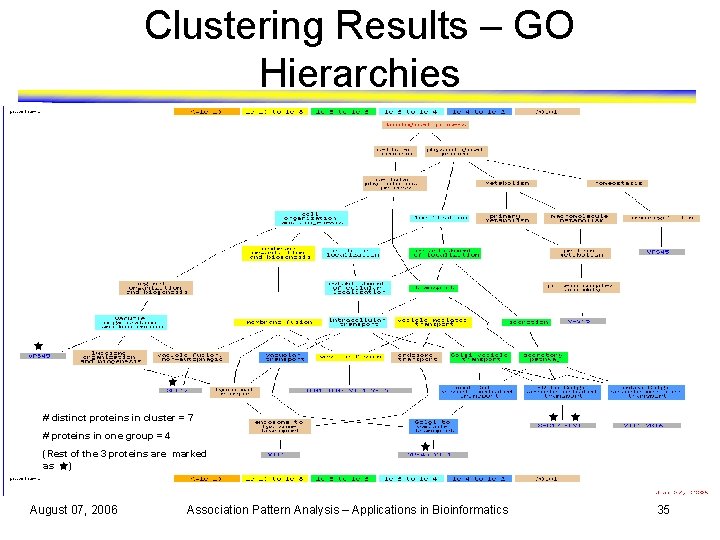
Clustering Results – GO Hierarchies # distinct proteins in cluster = 7 # proteins in one group = 4 (Rest of the 3 proteins are marked as ) August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 35
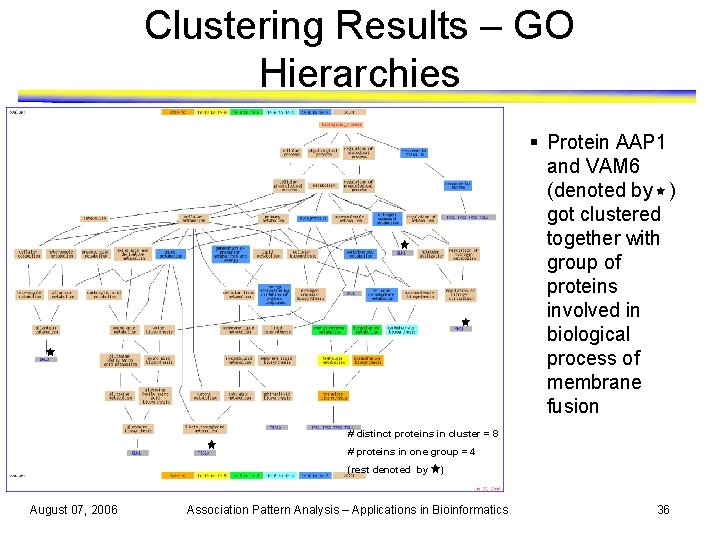
Clustering Results – GO Hierarchies § Protein AAP 1 and VAM 6 (denoted by ) got clustered together with group of proteins involved in biological process of membrane fusion # distinct proteins in cluster = 8 # proteins in one group = 4 (rest denoted by August 07, 2006 ) Association Pattern Analysis – Applications in Bioinformatics 36

Error Tolerant Itemsets (ETIs) • An error-tolerant itemset (ETI) can have a fraction of the items missing in each transaction. Example: see the data in the table – Let = 1/4. In other words, each transaction needs to have 3/4 (75%) of the items. – X = {i 1, i 2, i 3, i 4} and Y = {i 5, i 6, i 7, i 8} are both ETIs with a support of 4 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 37

ETIs to Identify Protein Functional Modules § Groups of proteins are identified as error tolerant itemsets (ETIs) § ETI relaxes the density constraints of the pattern in both dimensions § Maximum sparseness allowed: 0. 2 (along row) and 0. 25 (along column) § Minimum support: 5 protein complexes § Gene Ontology is used to validate following three identified ETIs • {CLF 1, LEA 1, PRP 46, RSE 1, SMB 1, SMD 2, SNU 114, SPP 382} • {Pre 2, Pre 4, Pre 5, Pre 6, Pre 8, Pre 9, Pup 3, Rpt 3, Scl 1} • {Rpn 10, Rpn 12, Rpn 3, Rpn 6, Rpn 8, Rpn 9, Rpt 2, Rpt 3, Rpt 6} August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 38
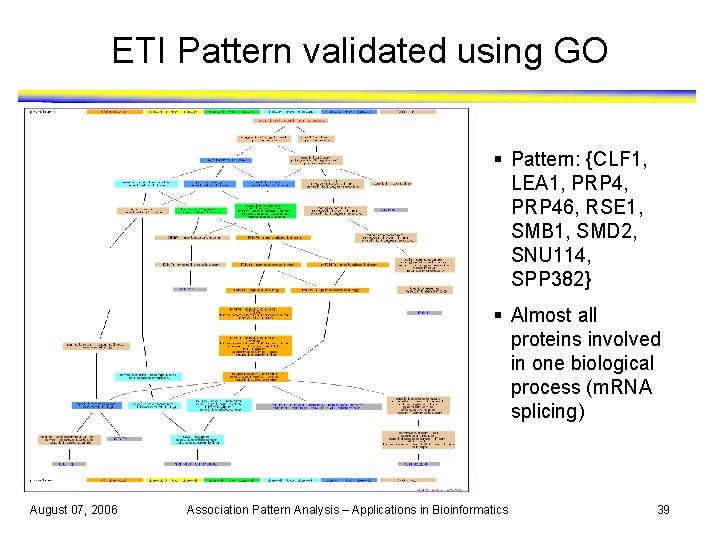
ETI Pattern validated using GO § Pattern: {CLF 1, LEA 1, PRP 4, PRP 46, RSE 1, SMB 1, SMD 2, SNU 114, SPP 382} § Almost all proteins involved in one biological process (m. RNA splicing) August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 39
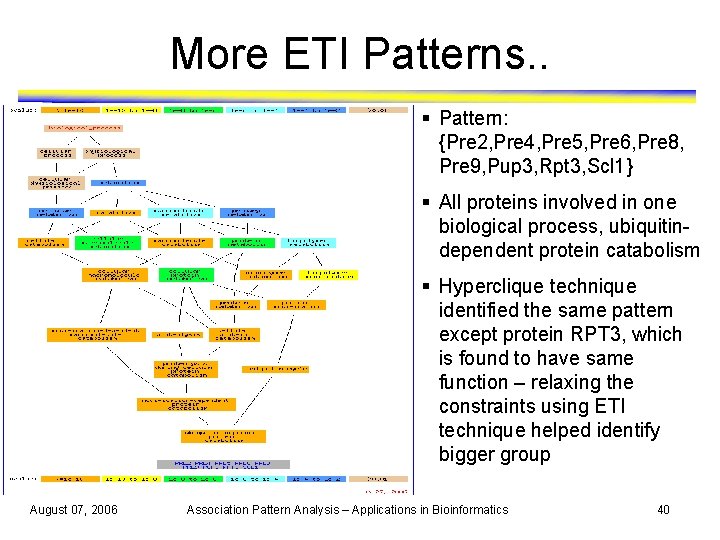
More ETI Patterns. . § Pattern: {Pre 2, Pre 4, Pre 5, Pre 6, Pre 8, Pre 9, Pup 3, Rpt 3, Scl 1} § All proteins involved in one biological process, ubiquitindependent protein catabolism § Hyperclique technique identified the same pattern except protein RPT 3, which is found to have same function – relaxing the constraints using ETI technique helped identify bigger group August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 40
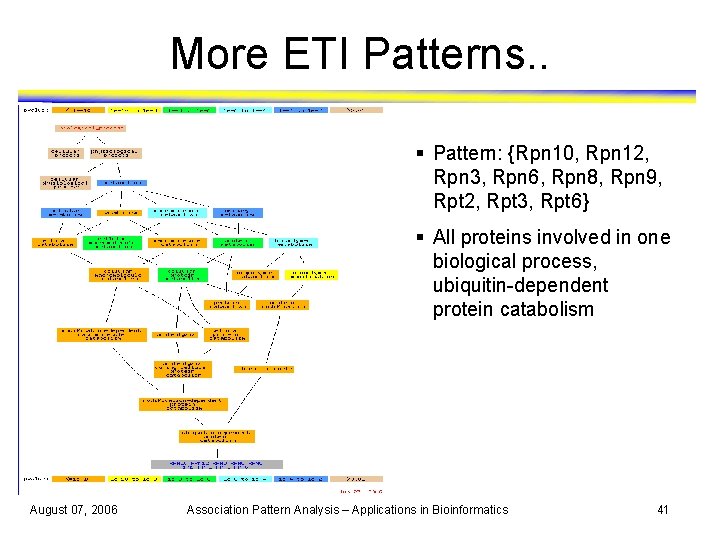
More ETI Patterns. . § Pattern: {Rpn 10, Rpn 12, Rpn 3, Rpn 6, Rpn 8, Rpn 9, Rpt 2, Rpt 3, Rpt 6} § All proteins involved in one biological process, ubiquitin-dependent protein catabolism August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 41

Concluding Remarks 1. Hyperclique and ETI patterns show great promise for identifying protein modules and for annotating uncharacterized proteins § Clustering does not perform as well as hypercliques and ETI due to a variety of reasons: • Each protein gets assigned to some cluster even if there is no right cluster for it • Modules can be overlapping • Modules can of different sizes • Data is high-dimensional August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 42
![References [1] Hui Xiong, X. He, Chris Ding, Ya Zhang, Vipin Kumar, Stephen R. References [1] Hui Xiong, X. He, Chris Ding, Ya Zhang, Vipin Kumar, Stephen R.](http://slidetodoc.com/presentation_image_h/ce4601eed84d86ae046ea5cfc256a8aa/image-43.jpg)
References [1] Hui Xiong, X. He, Chris Ding, Ya Zhang, Vipin Kumar, Stephen R. Holbrook, Identification of Functional Modules in Protein Complexes via Hyperclique Pattern Discovery, in Proc. of the Pacific Symposium on Biocomputing, (PSB 2005), 2005 [2] Pang-Ning Tan, Michael Steinbach, and Vipin Kumar, Introduction to Data Mining, Addison-Wesley April 2005 [3] Jinze Liu, Susan Paulsen, Xing Xu, Wei Wang, Andrew Nobel, Jan Prins, Mining Approximate Frequent Item sets in the Presence of Noise: Algorithms and Analysis, SIAM 2006 [4] Pang-Ning Tan, Vipin Kumar, and Jaideep Srivastava, Selecting the Right Interestingness Measure for Association Patterns, Proc of the Eighth ACM SIGKDD Int'l Conf. on Knowledge Discovery and Data Mining (SIGKDD-2002) [5] Hui Xiong, Pang-Ning Tan, and Vipin Kumar, Hyperclique Pattern Discovery, Data Mining and Knowledge Discovery Journal, accepted for publication as a regular paper, 2006 (short version appeared in proc of ICDM 2003) [6] A. Gavin et al. Functional organization of the yeast proteome by systematic analysis of protein complexes, Nature, 415: 141 -147, 2002 August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 43

Organizing Committee http: //www. siam. org/meetings/sdm 07/ Steering Committee Chair Vipin Kumar, University of Minnesota Conference Co-Chairs Chid Apte, IBM Research David Skillicorn, Queen’s University Program Co-Chairs Srinivasan Parthasarathy, Ohio State University Bing Liu, University of Illinois at Chicago Tutorial Chair Pang-Ning Tan, Michigan State University Workshop Co-Chairs Michael Berry, University of Tennessee Philip Chan, Florida Institute of Technology Publicity Chair Hui Xiong, Rutgers University August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 44

August 07, 2006 Association Pattern Analysis – Applications in Bioinformatics 45
- Slides: 45