Applications of Molecular Cytogenetics Dr Mohammed Alqahtani CSLTCG
























- Slides: 38


Applications of Molecular Cytogenetics Dr Mohammed Alqahtani CSLT(CG), CLSp(CG), RT, MBA, Ph. D Genomic Medicine Unit Founder & Director Center of Excellence in Genomic Medicine Research Founder & Director

Lecture Objectives • Understand how molecular cytogenetic techniques can be used to identify clinically relevant chromosome abnormalities • Be aware of the different types of molecular techniques that can be used to identify and clarify chromosome rearrangements

• Molecular Cytogenetic Techniques Powerful complement to conventional cytogenetic analysis of: – aneuploidy – structural rearrangements – submicroscopic rearrangements • microdeletions/duplications • subtelomere rearrangements

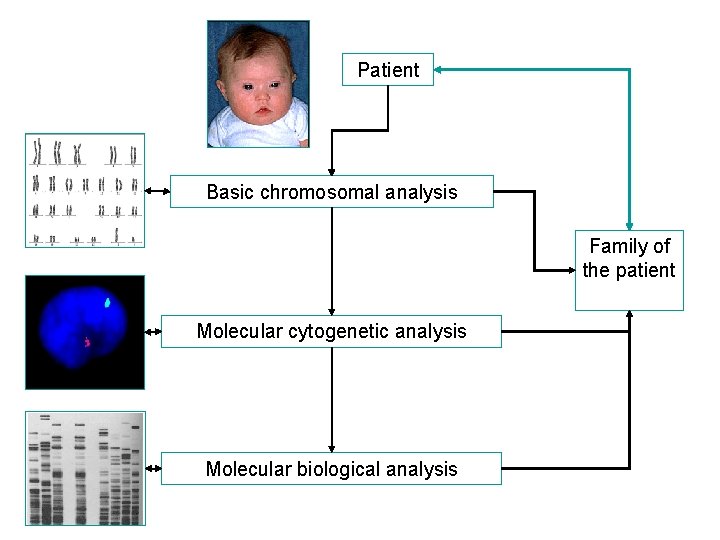
Patient Basic chromosomal analysis Family of the patient Molecular cytogenetic analysis Molecular biological analysis

Molecular cytogenetic examinations PCR hybridization • In most of cases interphase cells could be used for analysis (with exception of whole chromosome painting probes and M-FISH) • Examples of methods: – in situ hybridization and its modifications (CGH, M-FISH, fiber FISH atd. ) – Gene chips, resp. array CGH, DNA microarray etc. – PRINS, PCR in situ – quantitative fluorescent PCR, real time PCR – methods based on amplification of probe attached to target sequence (MLPA, MAPH)

Molecular Cytogenetics Era • • 1988 FISH 1992 Comparative Genomic Hybridization 1994 Reverse FISH 1996 Spectral Karyotyping, M-FISH 1999 M-Band analysis 2002 Fiber FISH 2002 Primed in situ labeling (PRINS) 2002 Microarray

Molecular Cytogenetic testing • POSTNATAL Stat Blood Routine Blood Skin Biopsy Product of Conception

• PRENATAL Amniotic Fluid Chorionic Villus Sampling Fetal Cord Blood • CANCER GENETICS Bone Marrow Oncology Blood Solid Tumor Lymph Node Pleural Effusion Core Biopsy

Molecular Application • • • FISH CGH PCR Real Time PCR DNA Sequencing Microarray

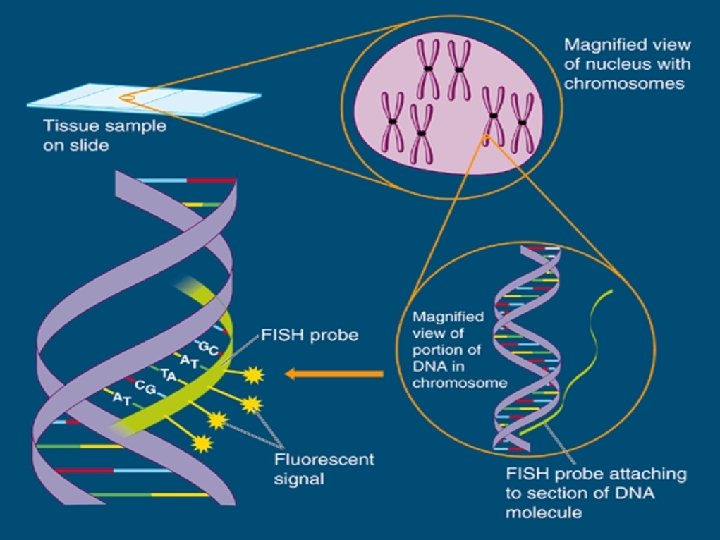
Fluorescence In Situ Hybridization (FISH)

FISH • A technique that hybridizes a DNA nucleic acid probe to a target DNA sequence contained within a cell nucleus. • A variety of specimen types can by analyzed using FISH. The intact cells are attached to a microscope slide using standard cytogenetic methods.

(FISH) TO RULE OUT: Ø Chromosome Microdeletion Detection Ø Interphase Chromosome Enumeration Ø Gene Rearrangements (ie, bcr/abl, PML/RARA) Ø Cryptic Chromosomal Rearrangements Ø Marker Chromosome Identification Ø Chromosome Breakpoint Mapping

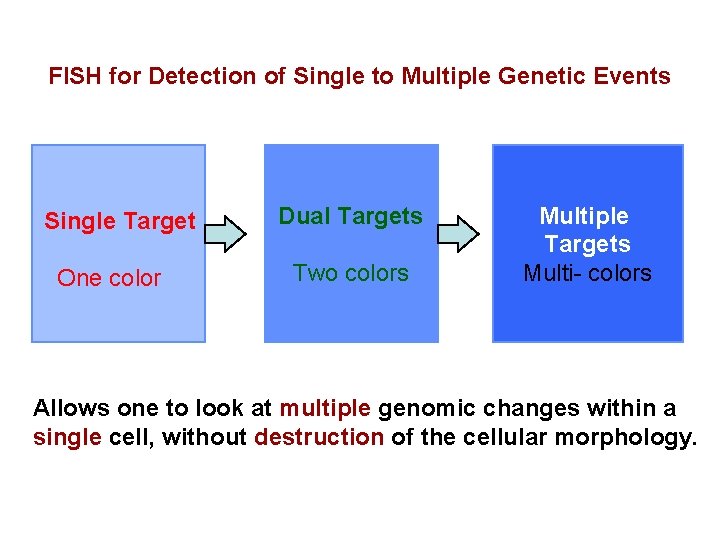
FISH for Detection of Single to Multiple Genetic Events Single Target One color Dual Targets Two colors Multiple Targets Multi- colors Allows one to look at multiple genomic changes within a single cell, without destruction of the cellular morphology.

Probes • Probe is a nucleic acid that – can be labeled with a marker which allows identification and quantitation – will hybridize to another nucleic acid on the basis of base complementarity

Probes Types of labeling • Direct & Indirect • Radioactive (32 P, 35 S, 14 C, 3 H) • Fluorescent • FISH: fluorescent in situ hybridization • Biotinylated (avidin-streptavidin)

Probe • A part of DNA (or RNA) that is complementary to certain sequence on target DNA (i. e. DNA of the patient) • Plasmid, phage DNA, cosmid (or combination of phage and plasmid DNA), YAC • PCR-product (amplification of certain segment of chromosomal DNA)

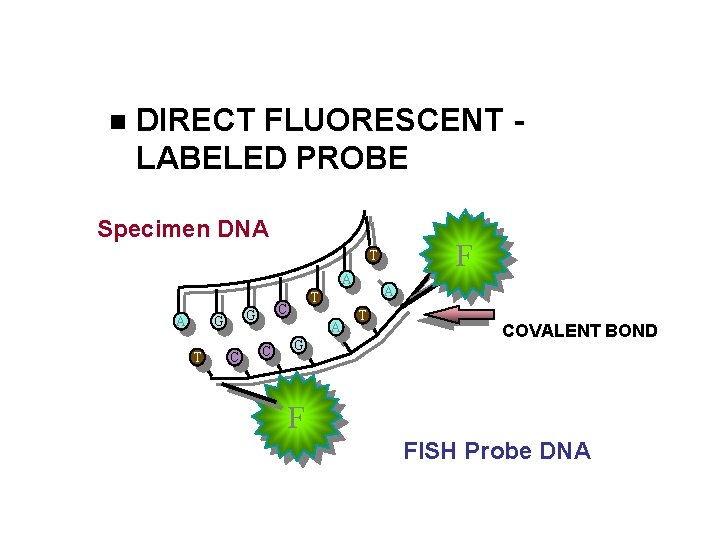
DIRECT FLUORESCENT LABELED PROBE Specimen DNA F T A A T C G G C A T G T COVALENT BOND F FISH Probe DNA


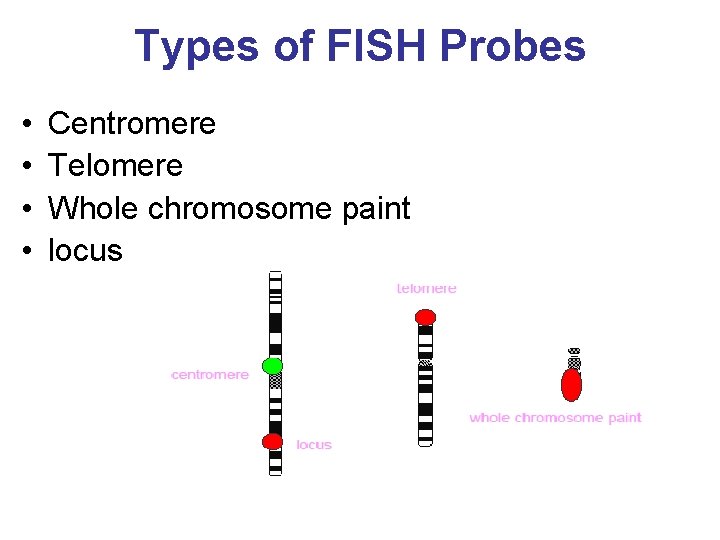
Types of FISH Probes • • Centromere Telomere Whole chromosome paint locus

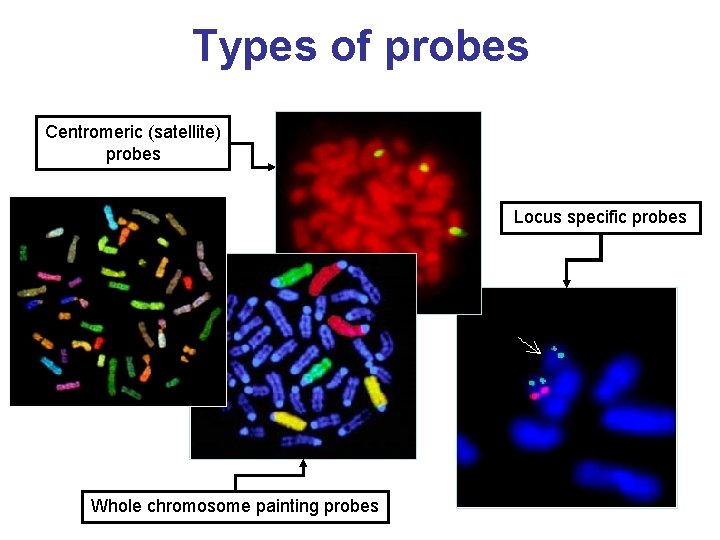
Types of probes Centromeric (satellite) probes Locus specific probes Whole chromosome painting probes

Types of probes • Telomeric probes have specificity for a single human chromosome arm. They contain a locus estimated to be within 300 kb of the end of the chromosome. • WCP Chromosome Painting Probes the hybridized probe fluoresces with bright intensity along the length of chromosome • CEP Chromosome Enumerator Probes (centromere area) – Most are Alpha and Satellite III Probes – Centromere regions stained brighter - means they are rich in A-T bonds

Types of probes • LSI Locus Specific Identifiers – Deletion Probes – Translocation Probes – Gene Detection & Localization – Gene Amplification Probes

In which conditions we have to indicate FISH analysis? • The material doesn't contain metaphase chromosomes – Unsuccessful cultivation – It isn't possible to cultivate the tissue from patient (preimplantation analysis, rapid prenatal examinations, examinations of solid tumors or autopsy material) • Analysis of complicated chromosomal rearrangements

In which conditions we have to indicate FISH analysis? • Identification of marker chromosomes • Analysis of low-frequency mosaic • Diagnosis of submicroscopic (cryptic) chromosomal rearrangements – Microdeletion syndromes – Amplification of oncogenes and microdeletion of tumor-suppressor genes in malignancies

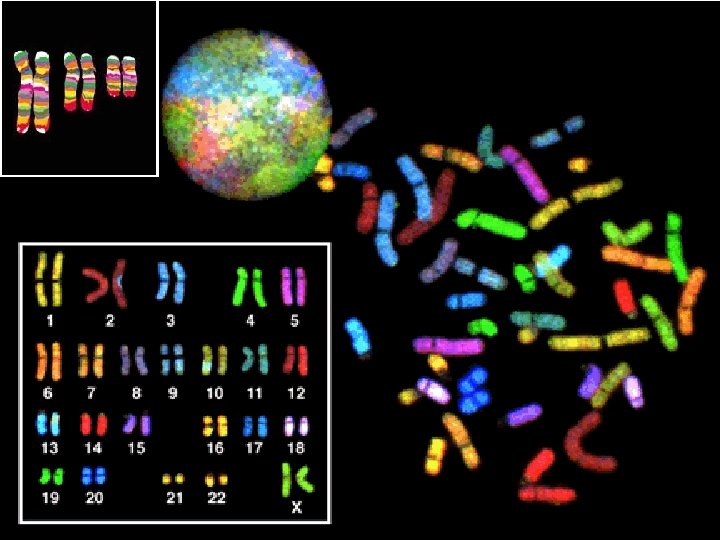
Multi Color FISH • Multicolor FISH can provide “colorized” information relative to chromosome rearrangements, especially useful in specimens where chromosome preparations are less than optimal for standard cytogenetic banding analysis.


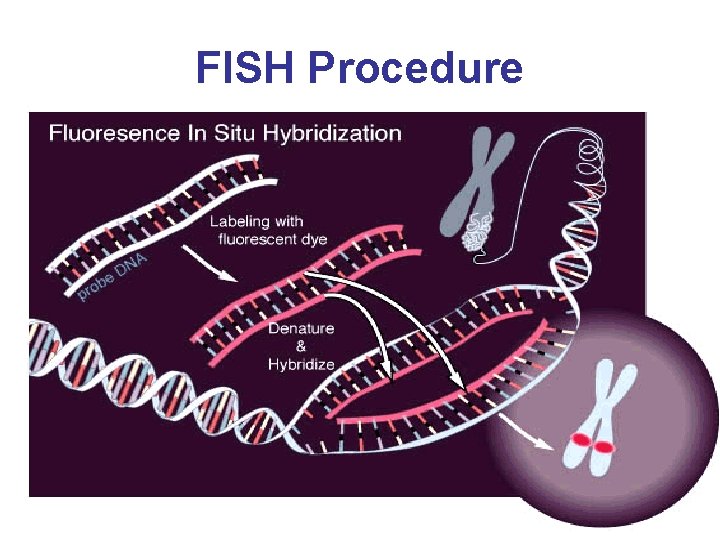
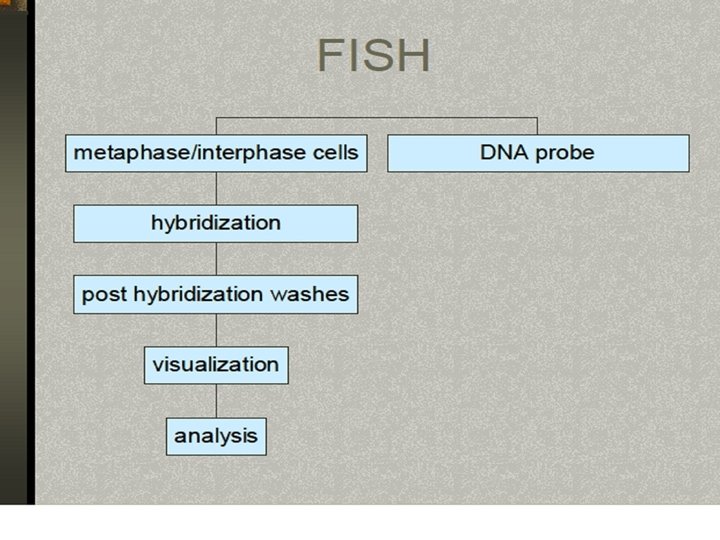
FISH Procedure • • • Denature the chromosomes Denature the probe Hybridization Fluorescence staining Examine slides or store in the dark

FISH Procedure

Direct Label FISH Technology

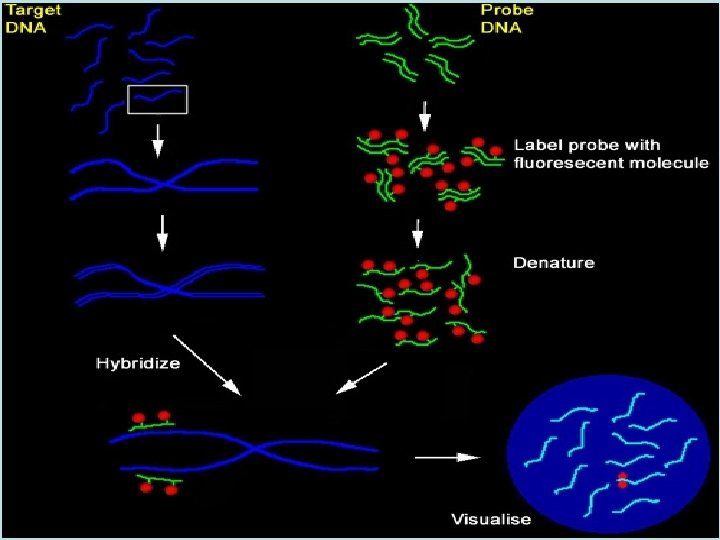
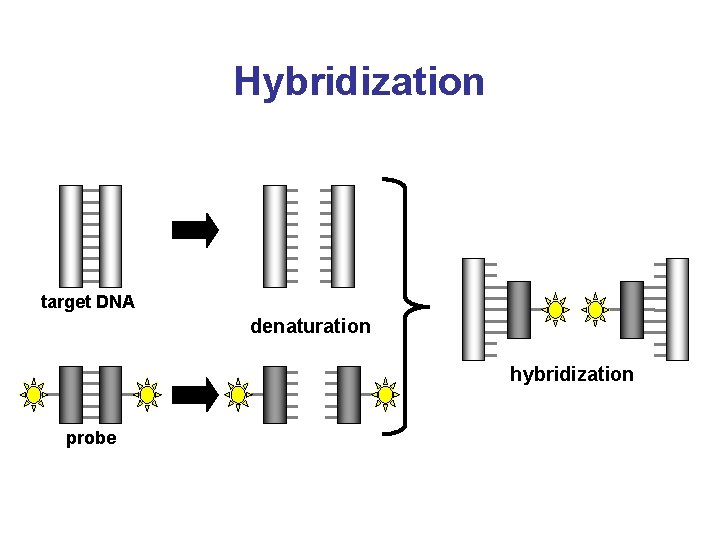
Hybridization target DNA denaturation hybridization probe

Hybridization • Nucleic acid hybridization is the formation of a duplex between two complementary sequences • Intermolecular hybridization: between two polynucleotide chains which have complementary bases – DNA-DNA – DNA-RNA – RNA-RNA • Annealing is another term used to describe the hybridization of two complementary molecules

Automated Hybridization HYBrite™ • The probe and target DNA are denatured together. • Faster, easier, and safer hybridization.


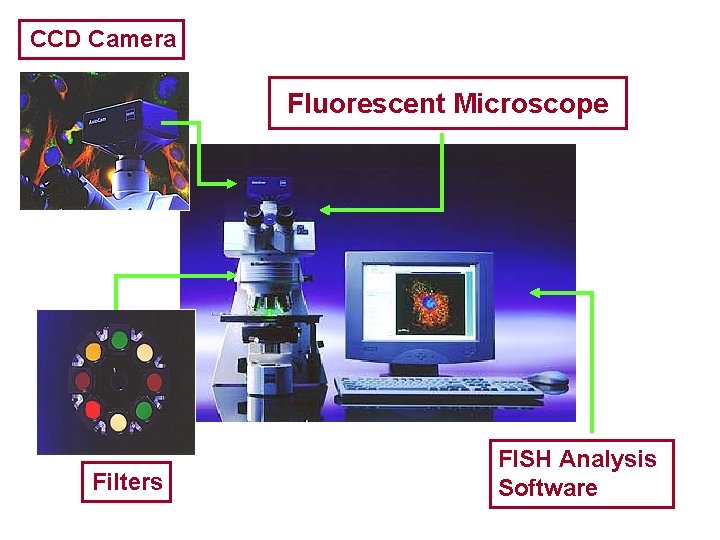
Visualization of the Probe • DNA probe is labeled with a colored fluorescent molecule. • This fluorescent molecule remains attached to the DNA during the hybridization process • The molecule emits a particular color when viewed through a fluorescence microscope that is equipped with the appropriate filter sets.

CCD Camera Fluorescent Microscope Filters FISH Analysis Software


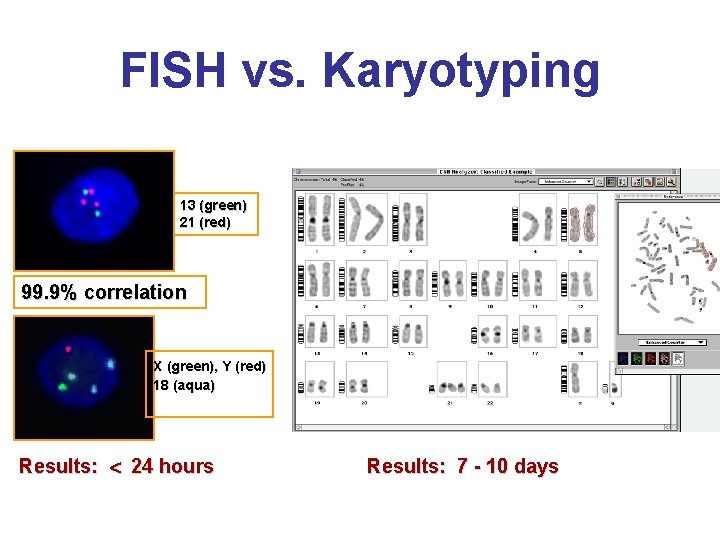
FISH vs. Karyotyping 13 (green) 21 (red) 99. 9% correlation X (green), Y (red) 18 (aqua) Results: 24 hours Results: 7 - 10 days