Dynamic Programming Algorithms II Mohammed Aledhari Bioinformatics data

Dynamic Programming Algorithms II Mohammed Aledhari Bioinformatics/ data mining Thursday May 23 rd, 2013

Outline • • • Local Sequence Alignment with Gap Penalties Multiple Alignment Gene Prediction Statistical Approaches to Gene Prediction Similarity-Based Approaches to Gene Prediction • Conclusion • References

The Local Alignment Problem • Goal: Find the best local alignment between two strings • Input : Strings v, w and scoring matrix δ • Output : Alignment of substrings of v and w whose alignment score is maximum among all possible alignment of all possible substrings

Local Alignments: Why? • Two genes in different species may be similar over short conserved regions and dissimilar over remaining regions. • Example: – Homeobox genes have a short region called the homeodomain that is highly conserved between species. – A global alignment would not find the homeodomain because it would try to align the ENTIRE sequence
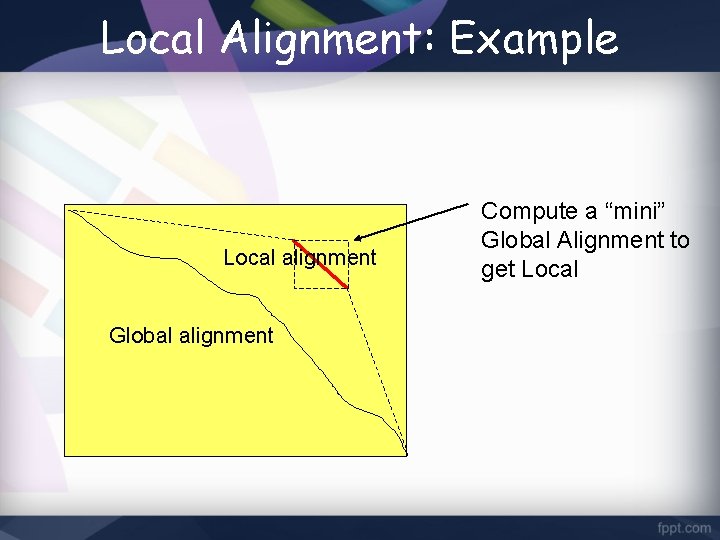
Local Alignment: Example Local alignment Global alignment Compute a “mini” Global Alignment to get Local

Local Alignment: Free Rides Yeah, a free ride! Vertex (0, 0) The dashed edges represent the free rides from (0, 0) to every other node.

Affine Gap Penalties • In nature, a series of k indels often come as a single event rather than a series of k single nucleotide events: This is more likely. Normal scoring would give the same score This is less for both alignments likely.

Adding “Affine Penalty” Edges to the Edit Graph There are many such edges! Adding them to the graph increases the running time of the alignment algorithm by a factor of n (where n is the number of vertices) So the complexity increases from O(n 2) to O(n 3)

Multiple Alignment versus Pairwise Alignment • Up until now we have only tried to align two sequences. • What about more than two? And what for? • A faint similarity between two sequences becomes significant if present in many • Multiple alignments can reveal subtle similarities that pairwise alignments do not reveal

Generalizing the Notion of Pairwise Alignment • Alignment of 2 sequences is represented as a 2 -row matrix • In a similar way, we represent alignment of 3 sequences as a 3 -row matrix A T _ G C G _ A _ C G T _ A A T C A C _ A • Score: more conserved columns, better alignment
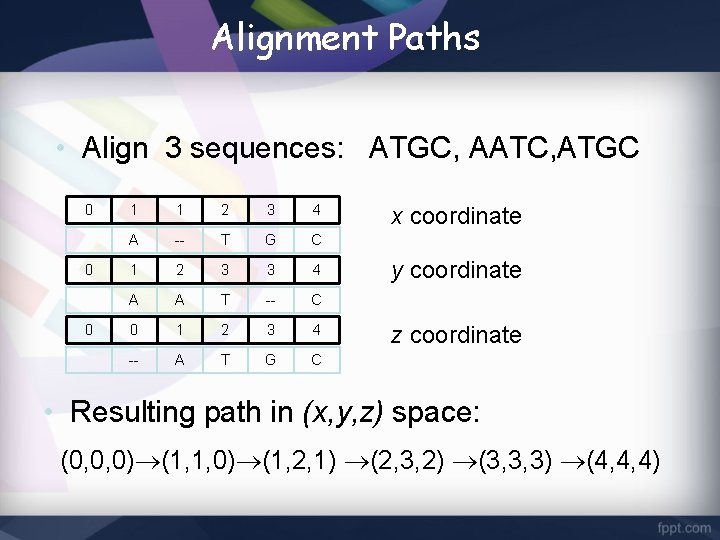
Alignment Paths • Align 3 sequences: ATGC, AATC, ATGC 0 0 0 1 1 2 3 4 A -- T G C 1 2 3 3 4 A A T -- C 0 1 2 3 4 -- A T G C x coordinate y coordinate z coordinate • Resulting path in (x, y, z) space: (0, 0, 0) (1, 1, 0) (1, 2, 1) (2, 3, 2) (3, 3, 3) (4, 4, 4)
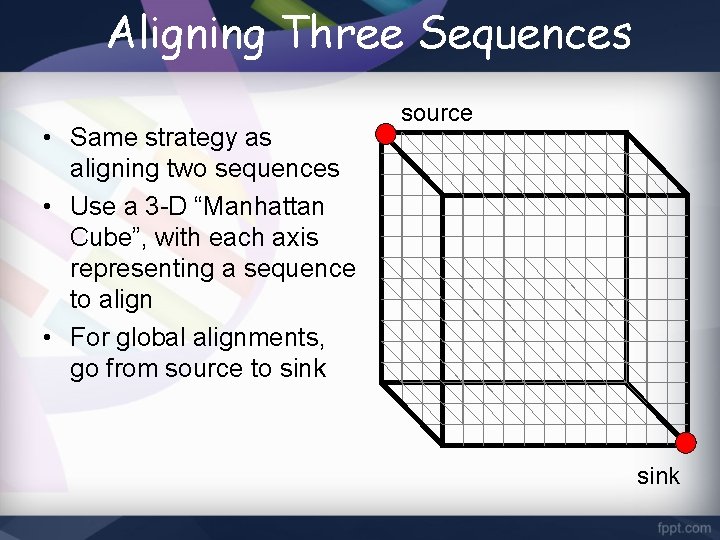
Aligning Three Sequences • Same strategy as aligning two sequences • Use a 3 -D “Manhattan Cube”, with each axis representing a sequence to align • For global alignments, go from source to sink source sink
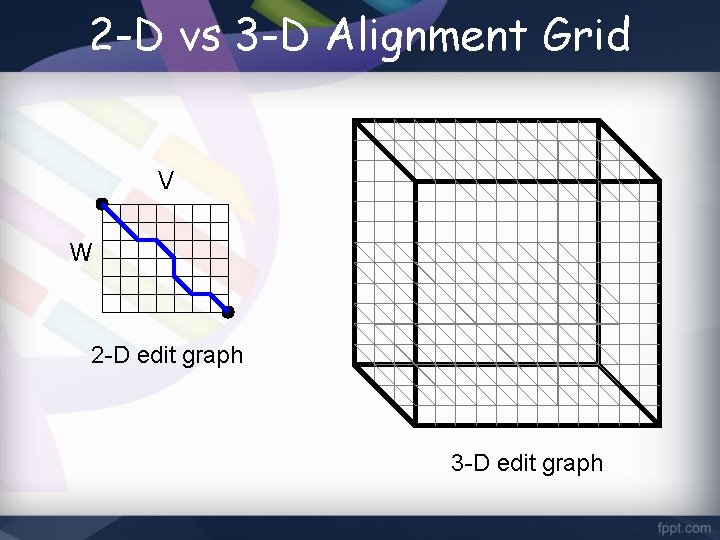
2 -D vs 3 -D Alignment Grid V W 2 -D edit graph 3 -D edit graph
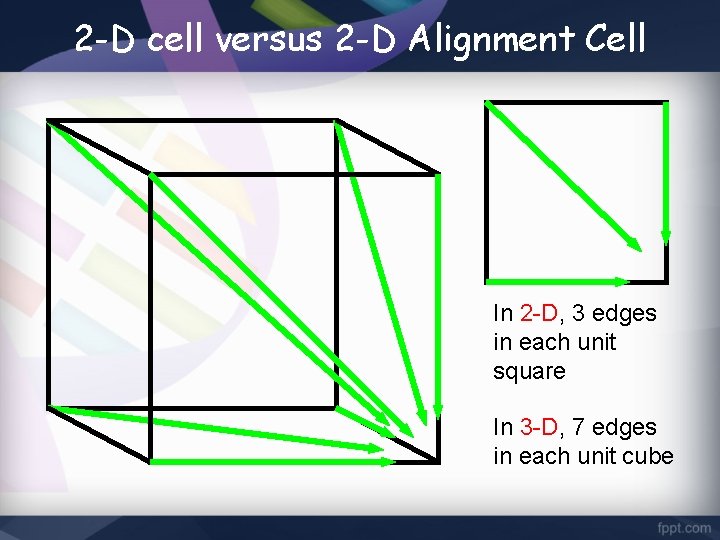
2 -D cell versus 2 -D Alignment Cell In 2 -D, 3 edges in each unit square In 3 -D, 7 edges in each unit cube
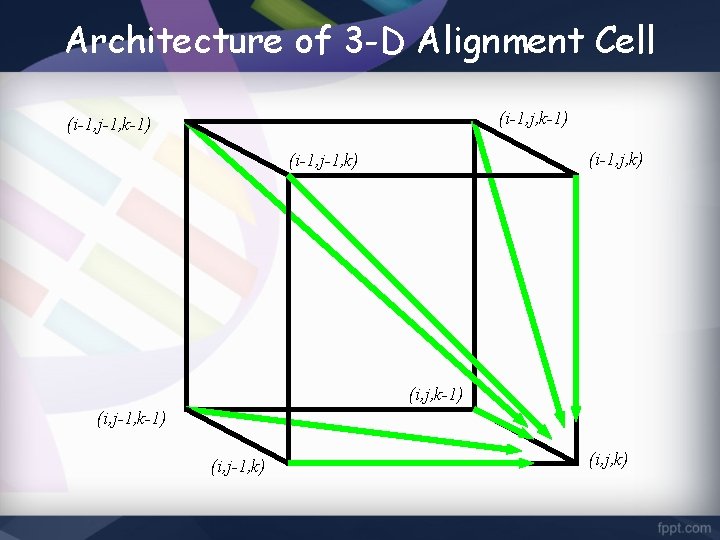
Architecture of 3 -D Alignment Cell (i-1, j, k-1) (i-1, j-1, k-1) (i-1, j, k) (i-1, j-1, k) (i, j, k-1) (i, j-1, k) (i, j, k)

Multiple Alignment: Running Time • For 3 sequences of length n, the run time is 7 n 3; O(n 3) • For k sequences, build a k-dimensional Manhattan, with run time (2 k-1)(nk); O(2 knk) • Dynamic programming approach for alignment between two sequences is easily extended to k sequences but it is impractical due to exponential running time

Greedy Approach: Example • Consider these 4 sequences s 1 s 2 s 3 s 4 GATTCA GTCTGA GATATT GTCAGC

Greedy Approach: Example (cont’d) • There are = 6 possible alignments s 2 s 4 GTCTGA GTCAGC (score = 2) s 1 s 4 GATTCA-G—T-CAGC(score = 0) s 1 s 2 GAT-TCA G-TCTGA (score = 1) s 2 s 3 G-TCTGA GATAT-T (score = -1) s 1 s 3 GAT-TCA GATAT-T (score s 3 s 4 GAT-ATT G-TCAGC (score = -1) = 1)

Greedy Approach: Example (cont’d) s 2 and s 4 are closest; combine: s 2 s 4 GTCTGA GTCAGC s 2, 4 GTCt/a. Ga/c. A (profile) new set of 3 sequences: s 1 s 3 s 2, 4 GATTCA GATATT GTCt/a. Ga/c
Gene Prediction • Gene Prediction is the process of detection of the location of open reading frames (ORFs) and delineation of the structures of introns as well as exons if the genes of interest are of eukaryotic origin. • The ultimate goal is to describe all the genes computationally with near 100% accuracy

Gene • Genes are the functional and physical unit of heredity passed from parent to offspring. • Gene: A sequence of nucleotides coding for protein • Gene Prediction Problem: Determine the beginning and end positions of genes in a genome • Genes are pieces of DNA, and most genes contain the information for making a specific protein.
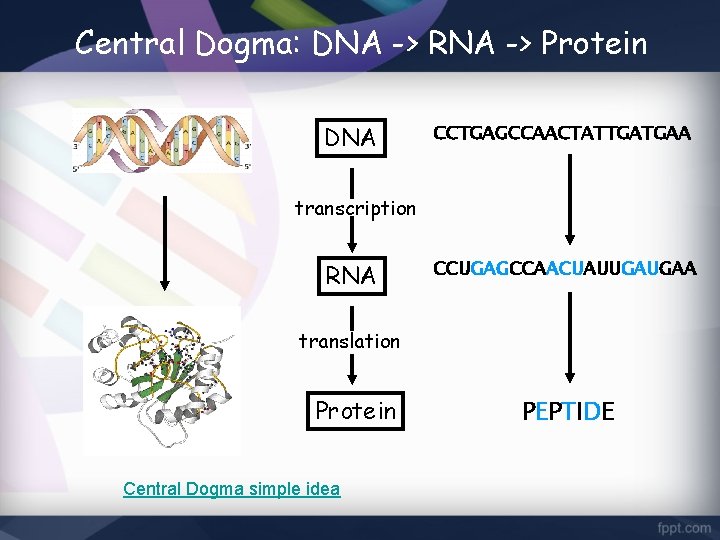
Central Dogma: DNA -> RNA -> Protein DNA CCTGAGCCAACTATTGATGAA transcription RNA CCUGAGCCAACUAUUGAUGAA translation Protein Central Dogma simple idea PEPTIDE
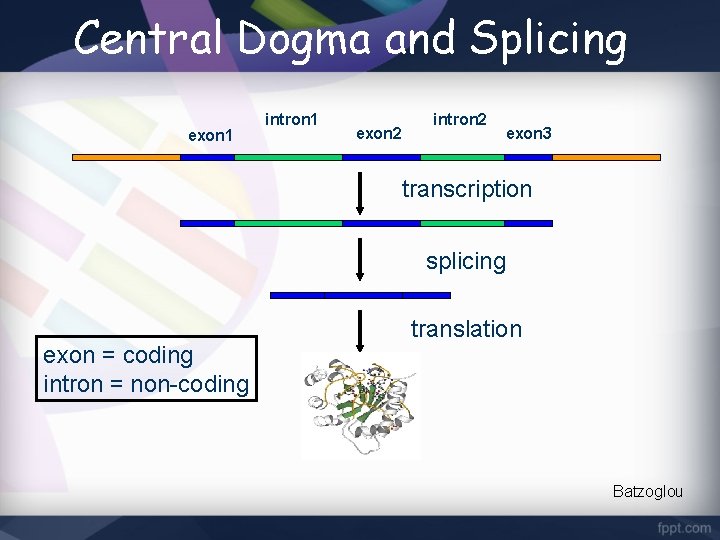
Central Dogma and Splicing exon 1 intron 1 exon 2 intron 2 exon 3 transcription splicing exon = coding intron = non-coding translation Batzoglou

Coding v/s Noncoding Coding region Noncoding region Coding regions are the parts of DNA which will give rise to a mature messenger RNA that will be translated into the specific amino acids of the protein product Noncoding regions are the parts of DNA which do not encode protein sequences. They may or may not be transcribed into RNA. E. g. : t. RNA, r. RNA, s. RNA genes

Two Approaches to Gene Prediction • Statistical: coding segments (exons) have typical sequences on either end and use different subwords than non-coding segments (introns). • Similarity-based: many human genes are similar to genes in mice, chicken, or even bacteria. Therefore, already known mouse, chicken, and bacterial genes may help to find human genes.

Gene Prediction Methods • Gene Prediction represents one of the most difficult problems in the field of pattern recognition, particularly in the case of eukaryotes • The principle difficulties are: o Detection of initiation site (AUG) o Alternative start codons o Gene overlap o Undetected small proteins

Gene Prediction Methods ACGTACTACGTACGATCGATCGACTGATCGATCG TACGTAGCGACTGACTGATCGACTACG TAGCTGCAGTCGACTGACTA Ab-initio methods Homology based methods

Ab-initio Methods • Predicts gene based on the given sequence alone. • Consists of two types of models: o Markov based models o Dynamic Programming

A brief introduction of HMMs • Hidden Markov models (HMMs) are discrete Markov processes where every state generates an observation at each time step. • A hidden Markov model (HMM) is statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states.

From Markov Model to HMM • HMMs are discrete Markov processes where each state also emits an observation according to some probability distribution, we need to augment our model. • Parameters – Initial state probabilities: πi – State transition probabilities: aij – Emission probabilities: ei(k) Markov Model Hidden Markov Model Each state emits an observation with 100% probability Each state emits an observation according to a certain probability distribution 30
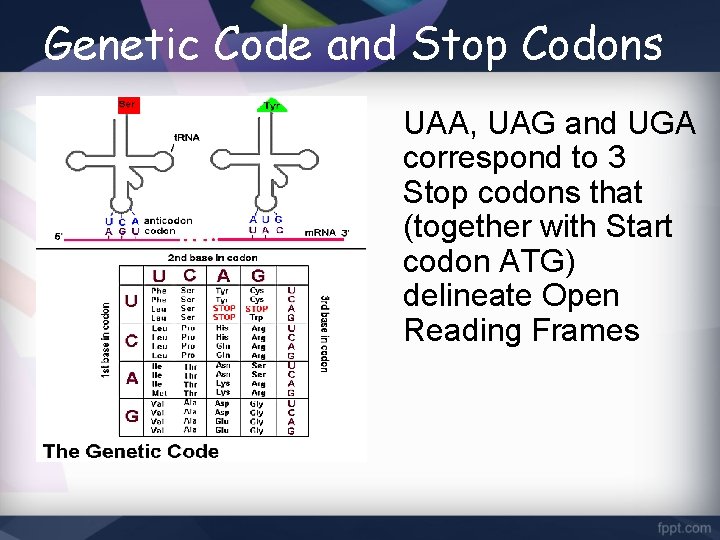
Genetic Code and Stop Codons UAA, UAG and UGA correspond to 3 Stop codons that (together with Start codon ATG) delineate Open Reading Frames
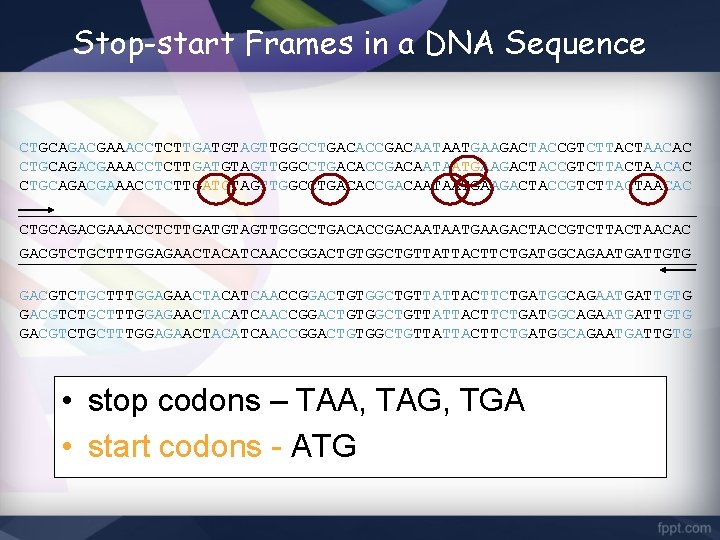
Stop-start Frames in a DNA Sequence CTGCAGACGAAACCTCTTGATGTAGTTGGCCTGACACCGACAATAATGAAGACTACCGTCTTACTAACAC GACGTCTGCTTTGGAGAACTACATCAACCGGACTGTGGCTGTTATTACTTCTGATGGCAGAATGATTGTG • stop codons – TAA, TAG, TGA • start codons - ATG

Codon Usage in Human Genome

Gene Prediction Tools • • GENSCAN/Genome Scan Twin. Scan Glimmer Gen. Mark

The GENSCAN Algorithm • Algorithm is based on probabilistic model of gene structure similar to Hidden Markov Models (HMMs). • GENSCAN uses a training set in order to estimate the HMM parameters, then the algorithm returns the exon structure using maximum likelihood approach standard to many HMM algorithms (Viterbi algorithm). – Biological input: Codon bias in coding regions, gene structure (start and stop codons, typical exon and intron length, presence of promoters, presence of genes on both strands, etc) – Covers cases where input sequence contains no gene, partial gene, complete gene, multiple genes.

GENSCAN Limitations – Does not use similarity search to predict genes. – Does not address alternative splicing. – Could combine two exons from consecutive genes together

Genome. Scan • Incorporates similarity information into GENSCAN: predicts gene structure which corresponds to maximum probability conditional on similarity information • Algorithm is a combination of two sources of information – Probabilistic models of exons-introns – Sequence similarity information

Twin. Scan • Aligns two sequences and marks each base as gap ( - ), mismatch (: ), match (|), resulting in a new alphabet of 12 letters: Σ {A-, A: , A |, C-, C: , C |, G-, G: , G |, T-, T: , T|}. • Run Viterbi algorithm using emissions ek(b) where b ∊ {A-, A: , A|, …, T|}.

Twin. Scan (cont’d) • The emission probabilities are estimated from human/mouse gene pairs. – Ex. e. I(x|) < e. E(x|) since matches are favored in exons, and e. I(x-) > e. E(x-) since gaps (as well as mismatches) are favored in introns. – Compensates for dominant occurrence of poly-A region in introns

Glimmer • Gene Locator and Interpolated Markov Model. ER • Finds genes in bacterial DNA • Uses interpolated Markov Models

The Glimmer Algorithm • Made of 2 programs – Build. IMM • Takes sequences as input and outputs the Interpolated Markov Models (IMMs) – Glimmer • Takes IMMs and outputs all candidate genes • Automatically resolves overlapping genes by choosing one, hence limited • Marks “suspected to truly overlap” genes for closer inspection by user

Gen. Mark • Based on non-stationary Markov chain models • Results displayed graphically with coding vs. noncoding probability dependent on position in nucleotide sequence

Useful internet gene prediction resources • http: //www. nslij-genetics. org/gene/ • http: //dot. imgen. bcm. tmc. edu: 9331/seqsearch/gene-search. html • http: //genome. cs. mtu. edu/aat. html • http: //bioweb. pasteur. fr/seqanal/interfaces/cdssimple. html • http: //genomic. sanger. ac. uk/gf/gf. shtml • http: //searchlauncher. bcm. tmc. edu: 9331/seqsearch/gene-search. html

Conclusion Local sequence alignment, Alignment with gap penalties, Multiple alignment, Gene prediction, Statistical approaches to gene prediction, Similarity-Based approaches to gene prediction

References • http: //bix. ucsd. edu/bioalgorithms/ (text book website) • http: //biochem 218. stanford. edu/ • http: //www. cs. washington. edu/education/c ourses/cse 527/09 au/ • http: //stellar. mit. edu/S/course/6/fa 09/6. 047 /index. html

Questions & comments Thank you
- Slides: 46