Algorithm based on approaches n Global Sequence Alignment

Algorithm based on approaches n § Global Sequence Alignment Local Sequence Alignment

Global Sequence Alignment n n n n Goal? to discover the overall relationship between sequences How? The alignments span the entire length of the sequences being compared All characters in both sequences participate in the alignment Useful mostly for finding closely related sequences where sequences are expected to be similar across their entire lengths Algorithm used- the Needleman-Wunsch (Needle)

Needleman-Wunsch Algorithm n n Based on the dynamic programming approach to obtain an optimal global alignment of two sequences solving complex problems by breaking them down into simpler subproblems In general, to solve a given problem, we need to solve different parts of the problem (subproblems), then combine the solutions of the subproblems to reach an overall solution. Often, many of these subproblems are really the same. The dynamic programming approach seeks to solve each subproblem only once, thus reducing the number of computations

3 steps in Needleman-Wuncsh Algorithm 1. Initialization- 2 sequences of length x and y 2. Matrix fill- the value for each position of the are considered for the alignment as input data. The program creates a matrix with (x+1) columns and (y+1) rows. matrix Mi, j is the information about the maximum global alignment score that position computed with the help of recurrence relations specific to this algorithm.

3. Traceback - This step determines the actual alignment(s) that result in the maximum score. It begins, in the M(x+1), (y+1) position in the matrix that has the maximal score.

The formulae…. M i, j = MAXimum [M i-1, j-1 + Sxi, yi (match/mismatch in the diagonal), M i, j-1 + w (gap in sequence 1), M i-1, j + w (gap in sequence 2) ]

Example n n Given 2 sequences Seq 1 GAATTCAGTTA Seq 2 GGATCGA Compute the above alignment using Needleman-Wuncsh Algorithm

Step 1 - Initialization n n Sequence 1: G A A T T C A G T T Sequence 2: G G A T C G A The scoring scheme assumed is Si, j = 1 for the match score if the residue at position i of sequence 1 is the same as the residue at position j of the sequence Si, j = 0 for the mismatch score w = 0 for the gap penalty

Step 2 – Matrix Fill n n n The positions of the first row (i. e. , M 0, j ) and the first column (i. e. , Mi, 0) are filled with 0’s, as in these cases each residue in the sequence is actually compared with nothing actual scoring is done starting from M 1, 1, the upper left hand corner of the matrix M and then the row and column corresponding to that position is filled. This way the maximal score Mi, j for each position in the matrix is filled. To find Mi, j you need to know the score for the left Mi-1, j, above Mi, j-1, and the top left diagonal Mi-1, j-1 matrix positions to i, j.

G A A T T C A G T T A 0 0 0 G 0 1 G 0 A 0 T 0 C 0 G 0 A 0 Matrix with values filled in the 0 th row and column as well as position 1, 1

G A A T T C A G T T A 0 0 0 G 0 1 1 1 G 0 1 A 0 1 T 0 1 C 0 1 G 0 1 A 0 1 Matrix with values filled in the 1 st row and column

G A A T T C A G T T A 0 0 0 G 0 1 1 1 G 0 1 1 1 1 2 2 A 0 1 2 2 2 2 2 3 T 0 1 2 2 3 3 3 3 C 0 1 2 2 3 3 3 4 4 4 G 0 1 2 2 3 3 3 4 4 5 5 5 A 0 1 2 3 3 4 5 5 5 6 Matrix with maximum score value
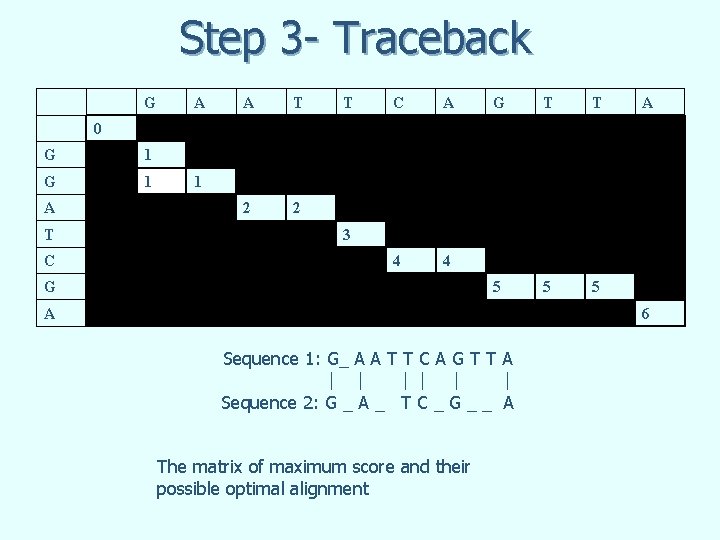
Step 3 - Traceback G A A T 2 2 T C A 4 4 G T T 5 5 5 A 0 G 1 A T C 1 3 G A 6 Sequence 1: G_ A A T T C A G T T A Sequence 2: G _ A _ T C _ G _ _ A The matrix of maximum score and their possible optimal alignment

Local Sequence Alignment n n n n To detect regions of high similarity between sequences Flexible, the alignment scores can be high because fragments of sequences are considered Application: Sequences of different lengths are compared Long sequences containing both the coding and noncoding regions are compared Proteins from different protein families are compared to find conserved domains Sequence comparison using global alignment does not give expected score, but there are clues that let you think that the sequences have similar parts.

Algorithm based on types of sequence alignment Pairwise Sequence Alignment n. Multiple Sequence Alignment n

Pairwise Sequence Alignment n n n Concerned with finding the best matching in local or global alignments of protein or DNA/RNA data Purpose: to find related homologous of a gene or gene product in a database on known examples Application: the identification of sequences of unknown structure or function, study of molecular evolution

Multiple Sequence Alignment n n is an extension of pairwise alignment. Here an unknown sequence is matched with several known sequences find common regions between the sequences at once, without making pairwise alignments first. There are several approaches, one of the most popular being the progressive alignment strategy used by the Clustal family of programs.

MSA-ctd’ n n n This is used to build phylogenetic trees as well as to build sequence profiles which are used to search sequence databases for more distant relatives. Here we will learn multiple alignment with heuristic algorithms. Others- Phylogenetic trees and hidden Markov Model

MSA with heuristic algorithm n n n Heuristic algorithms are faster algorithms that are based on assumptions and approximations Unlike dynamic programming, these algorithms do not make all possible pairwise comparisons to all of the database sequences, and thus they are not so expensive learning to solve a solution by trying is the approach of such algorithms The algorithms learn by experiences as done by rule-ofthumb or by ‘trial-and-error’ methods. Thus based on successive approximations, heuristics algorithms solve similarity search and alignment problems

Ctd’ n n n methods that are devised to search a small fraction of a dynamic programming matrix by looking at all the high scoring alignments. heuristic algorithms compromise on sensivity There are cases where such algorithms sometimes miss the best scoring alignment. Even the selectivity of these algorithms is comparable to the searches by dynamic programs. Both sensitivity and selectivity of heuristic algorithms are due to the incorporation of certain statistical parameters into the following two programs.

Ctd’ n n The two best known heuristic algorithms are BLAST and FASTA. These are two most commonly employed computational tools for scanning protein and DNA databases for similarity to a query sequence. Both programs do the following using different approach.

Ctd’ n n n Identify very short, exact matches between the query sequence and the database sequence(s). Extend the best short hits from the first step to look for longer stretches of similarity. Optimize the best hits with some form of dynamic programming.

Application of MSA When multiple sequence alignment applied and why it is used? n n n n larger group of protein distantly related members useful to reveal conserved residues or motifs in output. studying c. DNA clones, it is common practice to sequence them analysis of population define protein families making a phylogenetic tree the regulatory region
- Slides: 23