Multiple Sequence Alignments The Global Alignment problem AGTGCCCTGGAACCCTGACGGTGGGTCACAAAACTTCTGGA
Multiple Sequence Alignments
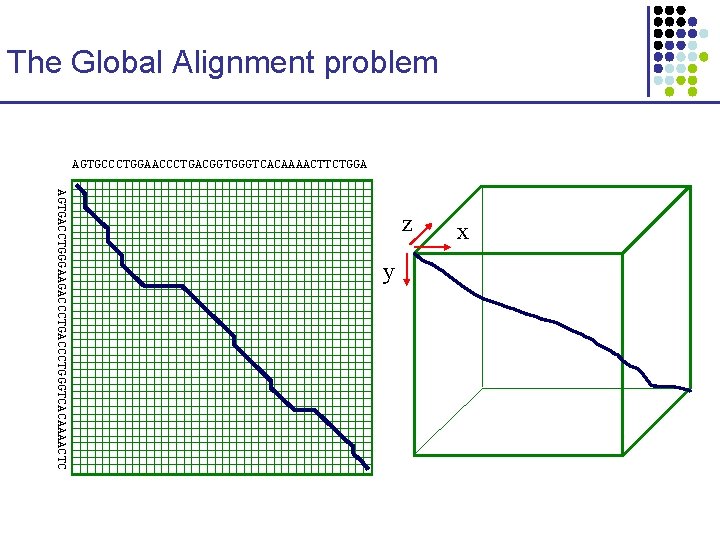
The Global Alignment problem AGTGCCCTGGAACCCTGACGGTGGGTCACAAAACTTCTGGA AGTGACCTGGGAAGACCCTGGGTCACAAAACTC z y x

Definition • Given N sequences x 1, x 2, …, x. N: § Insert gaps (-) in each sequence xi, such that • All sequences have the same length L • Score of the global map is maximum • A faint similarity between two sequences becomes significant if present in many • Multiple alignments can help improve the pairwise alignments

Scoring Function: Sum Of Pairs Definition: Induced pairwise alignment A pairwise alignment induced by the multiple alignment Example: x: y: z: AC-GCGG-C AC-GC-GAG GCCGC-GAG Induces: x: ACGCGG-C; y: ACGC-GAC; x: AC-GCGG-C; z: GCCGC-GAG; y: AC-GCGAG z: GCCGCGAG
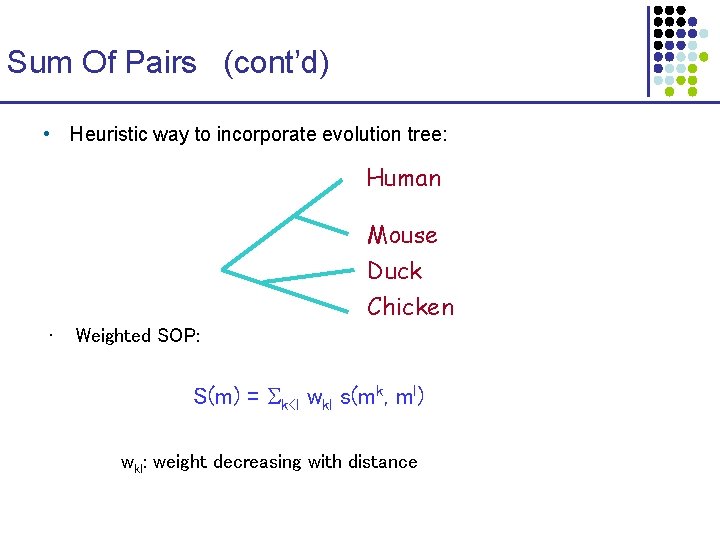
Sum Of Pairs (cont’d) • Heuristic way to incorporate evolution tree: Human Mouse Duck Chicken • Weighted SOP: S(m) = k<l wkl s(mk, ml) wkl: weight decreasing with distance

A Profile Representation T C C C A C G T O E C A A A G G G – – C C C T T T 1 A A A T C C T T C C C 1 . 6 1. 8 . 2 . 8 C C C . 4 1 T G G G G . 4. 2 1 . 6. 2. 2 1 C – – . 8 1. 2. 2. 2 A A G . 6 . 4. 4. 2 • Given a multiple alignment M = m 1…mn § Replace each column mi with profile entry pi • Frequency of each letter in • # gaps • Optional: # gap openings, extensions, closings

Multiple Sequence Alignments Algorithms

1. Multidimensional Dynamic Programming Generalization of Needleman-Wunsh: S(m) = i S(mi) (sum of column scores) F(i 1, i 2, …, i. N): Optimal alignment up to (i 1, …, i. N) F(i 1, i 2, …, i. N) = max(all neighbors of cube)(F(nbr)+S(nbr))

1. Multidimensional Dynamic Programming • Example: in 3 D (three sequences): • 7 neighbors/cell F(i, j, k) = max{ F(i-1, j-1, k-1)+S(xi, xj, xk), F(i-1, j-1, k )+S(xi, xj, - ), F(i-1, j , k-1)+S(xi, -, xk), F(i-1, j , k )+S(xi, -, - ), F(i , j-1, k-1)+S( -, xj, xk), F(i , j-1, k )+S( -, xj, xk), F(i , j , k-1)+S( -, -, xk) }
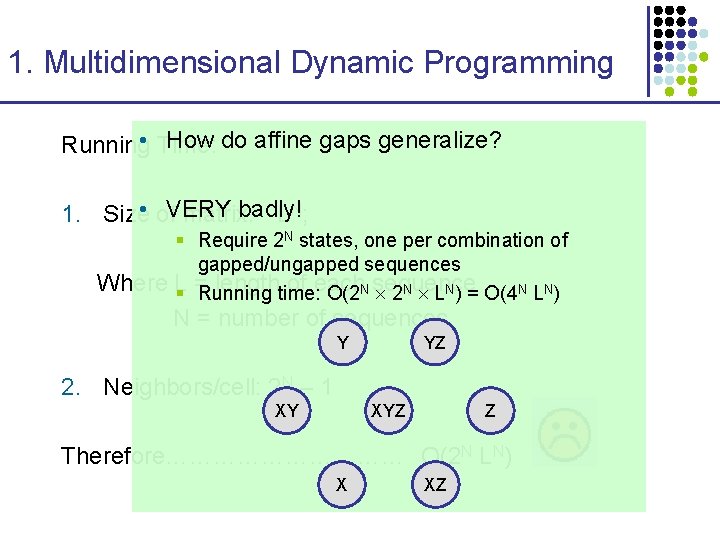
1. Multidimensional Dynamic Programming How do affine gaps generalize? Running • Time: badly! 1. Size • of. VERY matrix: L N; § Require 2 N states, one per combination of gapped/ungapped sequences Where L lengthtime: of each § = Running O(2 N sequence 2 N LN) = O(4 N LN) N = number of sequences Y YZ 2. Neighbors/cell: 2 N – 1 XY XYZ Z Therefore…………… O(2 N LN) X XZ

2. Progressive Alignment x y Example z Profile: (A, C, G, T, -) px = (0. 8, 0. 2, 0, 0, 0) w py = (0. 6, 0, 0. 4) • When evolutionary tree is known: s(px, py) = 0. 8*0. 6*s(A, A) + 0. 2*0. 6*s(C, A) + 0. 8*0. 4*s(A, -) + 0. 2*0. 4*s(C, -) § Align closest first, in the order of the tree § In each step, align two sequences. Result: x, y, or profiles px, py 0. 1, , to generate a new pxy = (0. 7, 0, 0, 0. 2) alignment with associated profile presult s(p , -) = 0. 8*1. 0*s(A, -) + 0. 2*1. 0*s(C, -) x Weighted version: § Tree edges have weights, proportional to the divergence in that edge Result: p = (0. 4, 0. 1, 0, 0, 0. 5) § New profile is a weighted average of two old x-profiles
2. Progressive Alignment x ? y z w • When evolutionary tree is unknown: § Perform all pairwise alignments § Define distance matrix D, where D(x, y) is a measure of evolutionary distance, based on pairwise alignment § Construct a tree (we will describe more in detail later in the course) § Align on the tree

Aligning two alignments • Given two alignments, m 1, m 2, can we find the optimal alignment under SOP scoring, with affine gaps? m 1 x GGGCACTGCAT y GGTTACGTC-- GTAGTCG x ---GTCACGTG y m 1 m 2 z GGGAACTGCAG w GGACGTACC-v GGACCT----- GTCGTCAGTCG z --CGCCAGGGG w --CGCCAGGGA v m 2

Aligning two alignments • Given two alignments, m 1, m 2, can we find the optimal alignment under SOP scoring, with affine gaps? NP-hard! m 1 x GGGCACTGCAT y GGTTACGTC-- GTAGTCG x ---GTCACGTG y m 1 m 2 z GGGAACTGCAG w GGACGTACC-v GGACCT----- GTCGTCAGTCG z --CGCCAGGGG w --CGCCAGGGA v m 2 Optimistic: assume no gap – don’t pay gap-open penalty Pessimistic: assume gap – pay gap-open penalty

Heuristics to improve multiple alignments • Iterative refinement schemes • A*-based search • Consistency • Simulated Annealing • …

Iterative Refinement One problem of progressive alignment: • Initial alignments are “frozen” even when new evidence comes Example: x: y: GAAGTT GAC-TT Frozen! z: w: GAACTG GTACTG Now clear correct y = GA-CTT

Iterative Refinement Algorithm (Barton-Stenberg): 1. 2. 3. Align most similar xi, xj Align xk most similar to (xixj) Repeat 2 until (x 1…x. N) are aligned 4. For j = 1 to N, Remove xj, and realign to x 1…xj-1 xj+1…x. N 5. Repeat 4 until convergence Note: Guaranteed to converge
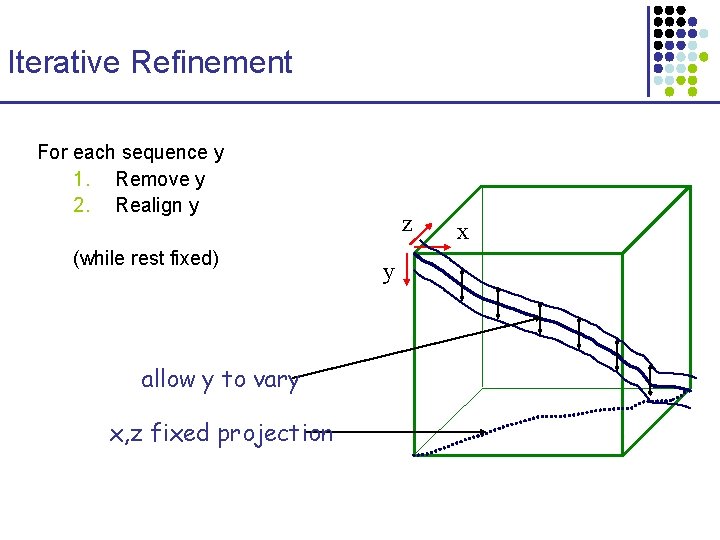
Iterative Refinement For each sequence y 1. Remove y 2. Realign y (while rest fixed) allow y to vary x, z fixed projection z y x

Iterative Refinement Example: align (x, y), (z, w), (xy, zw): x: y: z: w: GAAGTTA GAC-TTA GAACTGA GTACTGA After realigning y: x: y: z: w: GAAGTTA G-ACTTA GAACTGA GTACTGA + 3 matches

Iterative Refinement Example not handled well: x: y 1: y 2: y 3: GAAGTTA GAC-TTA z: w: GAACTGA GTACTGA Realigning any single yi changes nothing

A* for Multiple Alignments Review of the A* algorithm v GOAL START • • • Say that we have a gigantic graph G START: start node GOAL: we want to reach this node with the minimum path Dijkstra: O(Vlog. V + E) – too slow if the number of edges is huge A*: a way of finding the optimal solution faster in practice
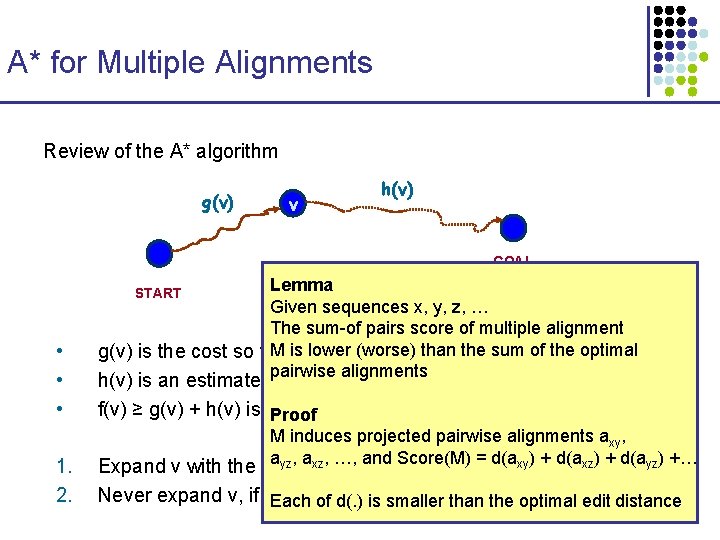
A* for Multiple Alignments Review of the A* algorithm g(v) v h(v) GOAL Lemma Given sequences x, y, z, … The sum-of pairs score of multiple alignment M is lower (worse) than the sum of the optimal g(v) is the cost so far pairwise alignments START • • • 1. 2. h(v) is an estimate of the minimum cost from v to GOAL f(v) ≥ g(v) + h(v) is the minimum cost of a path passing by v Proof M induces projected pairwise alignments axy, ayz, axz, …, and Score(M) = d(axy) + d(axz) + d(ayz) +… Expand v with the smallest f(v) Never expand v, if f(v) ≥ shortest path tothan the goal foundedit so distance far Each of d(. ) is smaller optimal

A* for Multiple Alignments g(v) v h(v) GOAL START • Nodes: Cells in the DPTo matrix compute h(v) • g(v): alignment cost so far For each pair of sequences x, y, • h(v): sum-of-pairs of individual pairwise alignments R Compute F (x, y), the DP matrix of scores of aligning a suffix of x to a suffix of y • Initial minimum alignment cost estimate: sum-of-pairs of global pairwise alignments Then, at position (i 1, i 2, …, i. N), h(v) becomes the sum of (N choose 2) FR scores
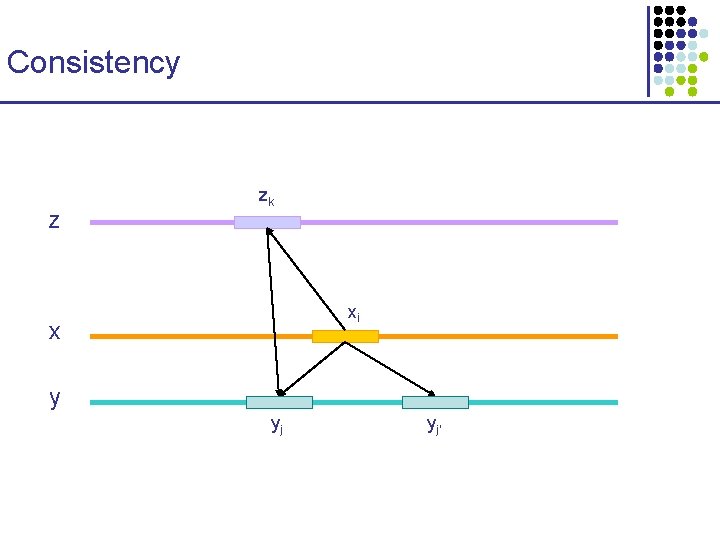
Consistency z zk xi x y yj yj’
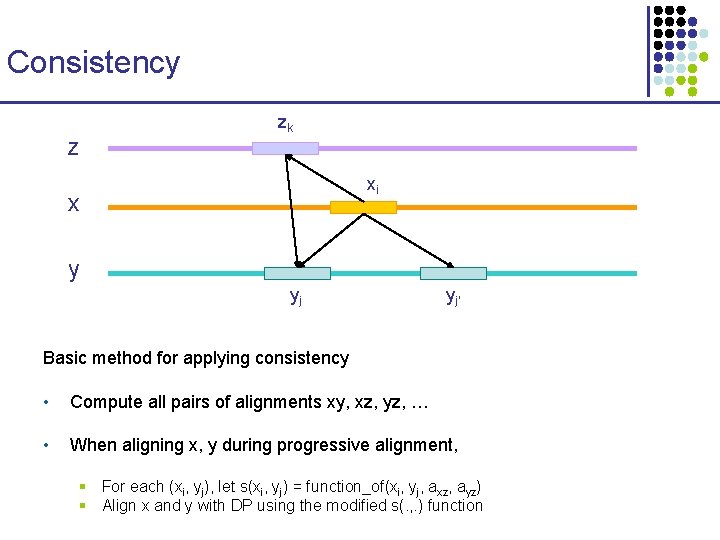
Consistency zk z xi x y yj yj’ Basic method for applying consistency • Compute all pairs of alignments xy, xz, yz, … • When aligning x, y during progressive alignment, § For each (xi, yj), let s(xi, yj) = function_of(xi, yj, axz, ayz) § Align x and y with DP using the modified s(. , . ) function

Some Resources Genome Resources Annotation and alignment genome browser at UCSC http: //genome. ucsc. edu/cgi-bin/hg. Gateway Specialized VISTA alignment browser at LBNL http: //pipeline. lbl. gov/cgi-bin/gateway 2 ABC—Nice Stanford tool for browsing alignments http: //encode. stanford. edu/~asimenos/ABC/ Protein Multiple Aligners http: //www. ebi. ac. uk/clustalw/ CLUSTALW – most widely used http: //phylogenomics. berkeley. edu/cgi-bin/muscle/input_muscle. py MUSCLE – most scalable http: //probcons. stanford. edu/ PROBCONS – most accurate
Whole-genome alignment Rat—Mouse—Human

Next 2 years: 20+ mammals, & many other animals, will be sequenced & aligned
- Slides: 28