Statistical Genomics Lecture 21 Farm CPU Zhiwu Zhang







- Slides: 24

Statistical Genomics Lecture 21: Farm. CPU Zhiwu Zhang Washington State University

Administration Homework 5, due April 13, Wednesday, 3: 10 PM Final exam: May 3, 120 minutes (3: 10 -5: 10 PM), 50 Department seminar (April 4) , Nural Amin

Outline History of method and software development Farm. CPU BLINK

Problems in GWAS Computing difficulties: millions of markers, individuals, and traits False positives, ex: “Amgen scientists tried to replicate 53 high-profile cancer research findings, but could only replicate 6”, Nature, 2012, 483: 531 False negatives

Q EMMA PC PC+K EMMAx Q+K GWAS Stream MLMM P 3 D CMLM SELECT GCTA ECMLM FST-LMM GEMMA Gen. Abel Farm. CPU BLINK

Computing speed t test GLM Speed improvement Power improvement Gen. ABEL Fa. ST-LMM GEMMA P 3 D/EMMAX EMMA CMLM ECMLM Select SUPER MLMM MLM Power | type I error

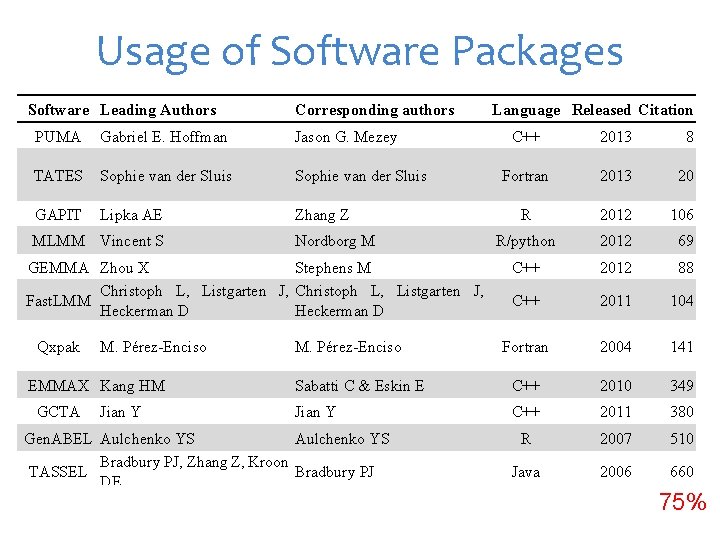
Usage of Software Packages Software Leading Authors Corresponding authors PUMA Gabriel E. Hoffman Jason G. Mezey TATES Sophie van der Sluis GAPIT Lipka AE Zhang Z MLMM Vincent S C++ 2013 8 Fortran 2013 20 R 2012 106 R/python 2012 69 C++ 2012 88 C++ 2011 104 Fortran 2004 141 Sabatti C & Eskin E C++ 2010 349 Jian Y C++ 2011 380 R 2007 510 Java 2006 660 C++ 2007 7037 75% Nordborg M GEMMA Zhou X Stephens M Christoph L, Listgarten J, Fast. LMM Heckerman D Qxpak M. Pérez-Enciso EMMAX Kang HM GCTA Jian Y Language Released Citation M. Pérez-Enciso Gen. ABEL Aulchenko YS Bradbury PJ, Zhang Z, Kroon TASSEL Bradbury PJ DE PLINK Purcell S

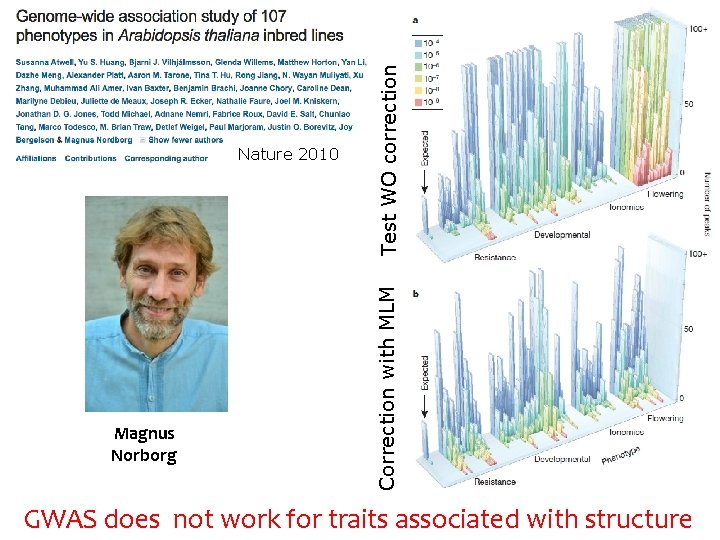
Magnus Norborg Test WO correction Correction with MLM Nature 2010 GWAS does not work for traits associated with structure

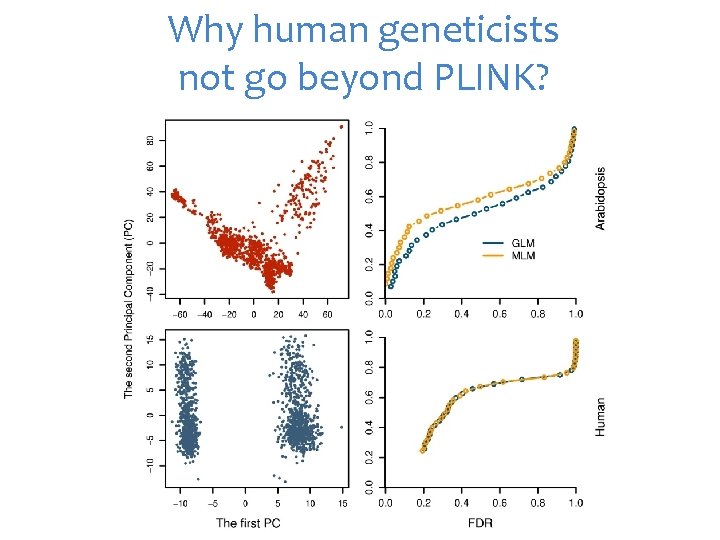
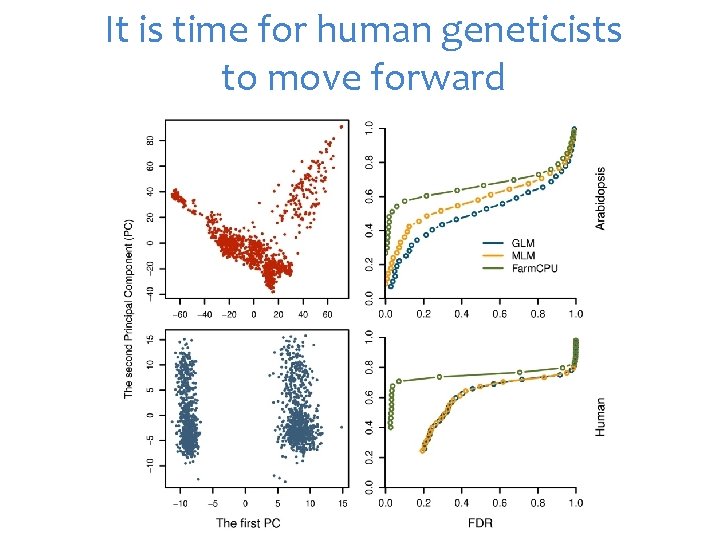
Why human geneticists not go beyond PLINK?

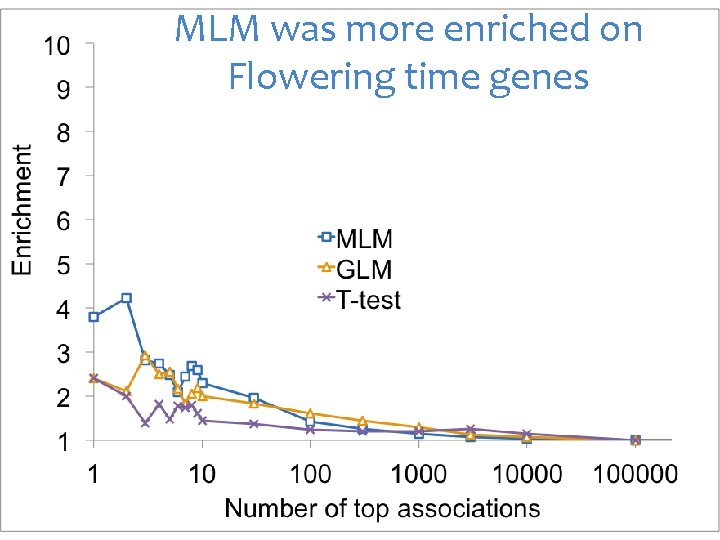
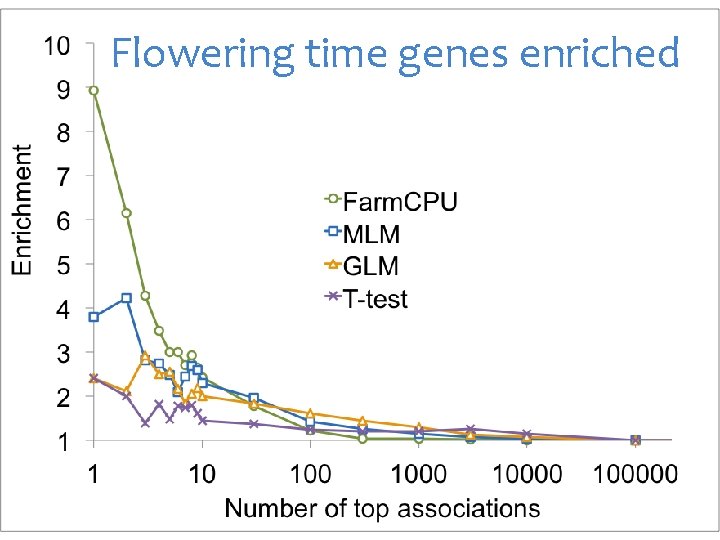
MLM was more enriched on Flowering time genes

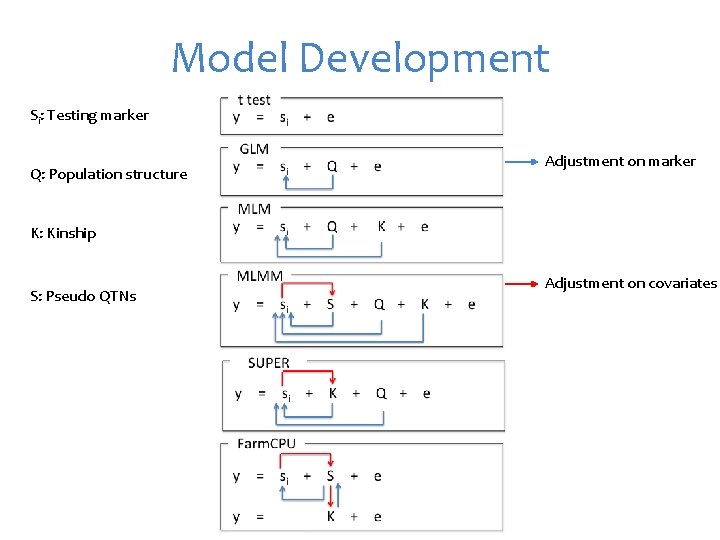
Model Development Si: Testing marker Q: Population structure Adjustment on marker K: Kinship S: Pseudo QTNs Adjustment on covariates

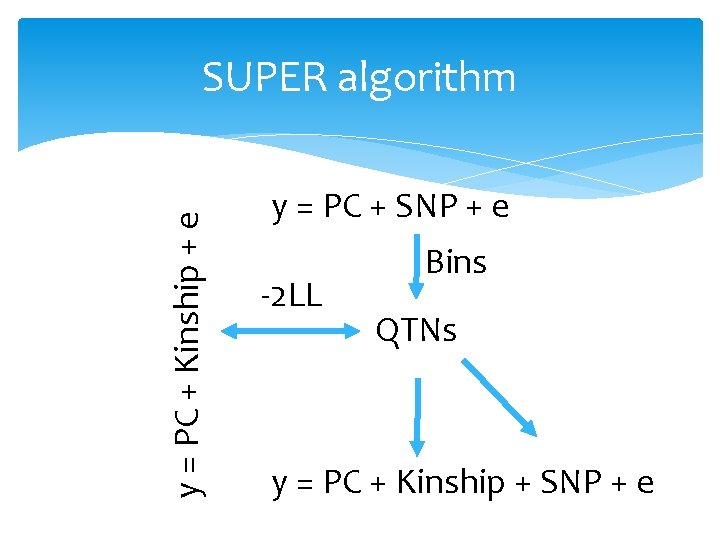
y = PC + Kinship + e SUPER algorithm y = PC + SNP + e -2 LL Bins QTNs y = PC + Kinship + SNP + e

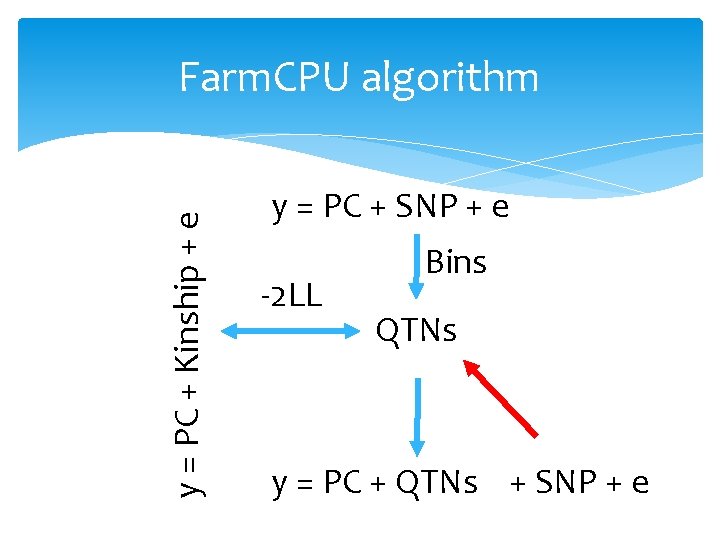
y = PC + Kinship + e Farm. CPU algorithm y = PC + SNP + e -2 LL Bins QTNs y = PC + QTNs + SNP + e

Computing speed t test GLM Speed improvement Power improvement Gen. ABEL BLINK Farm. CPU Fa. ST-LMM GEMMA P 3 D/EMMAX EMMA CMLM ECMLM Select SUPER MLMM MLM Power | type I error

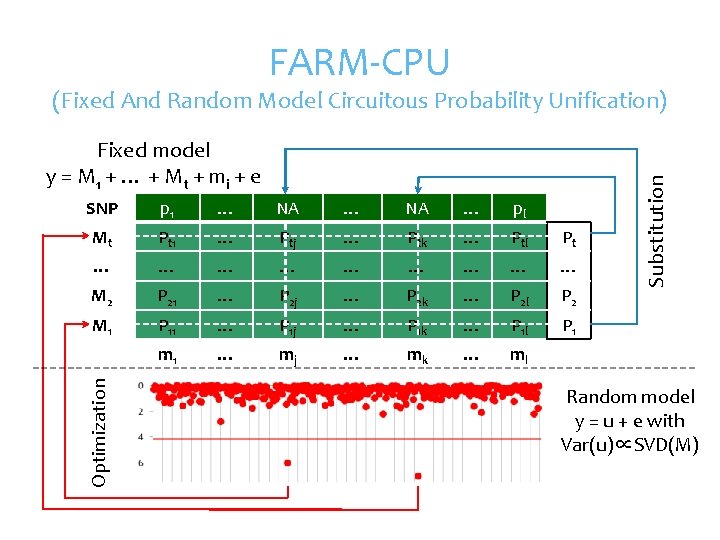
FARM-CPU Fixed model y = M 1 + … + M t + mi + e p 1 … NA … pl Mt Pt 1 … Ptj … Ptk … Ptl Pt … … … … … M 2 P 21 … P 2 j … P 2 k … P 2 l P 2 M 1 P 11 … P 1 j … P 1 k … P 1 l P 1 m 1 … mj … mk … ml Optimization SNP Substitution (Fixed And Random Model Circuitous Probability Unification) Random model y = u + e with Var(u)∝SVD(M)

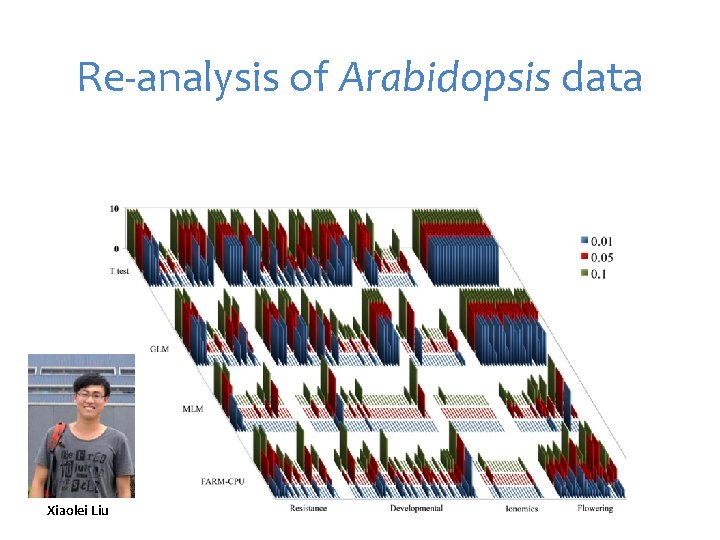
Re-analysis of Arabidopsis data Xiaolei Liu

Flowering time genes enriched

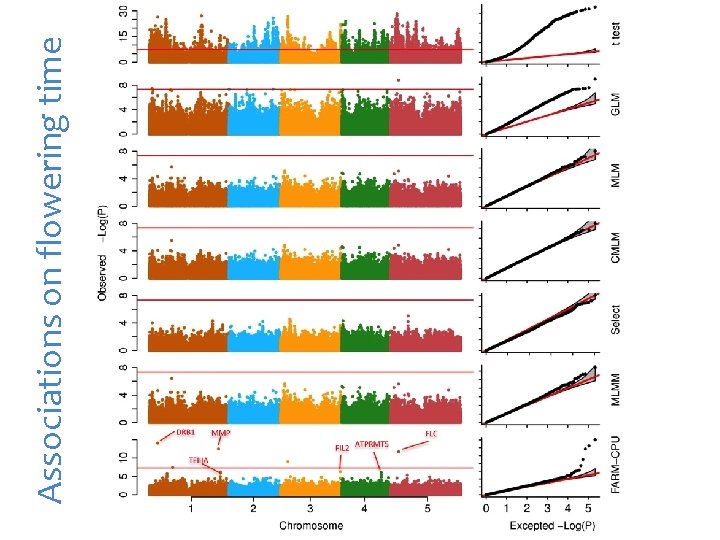
Associations on flowering time

It is time for human geneticists to move forward

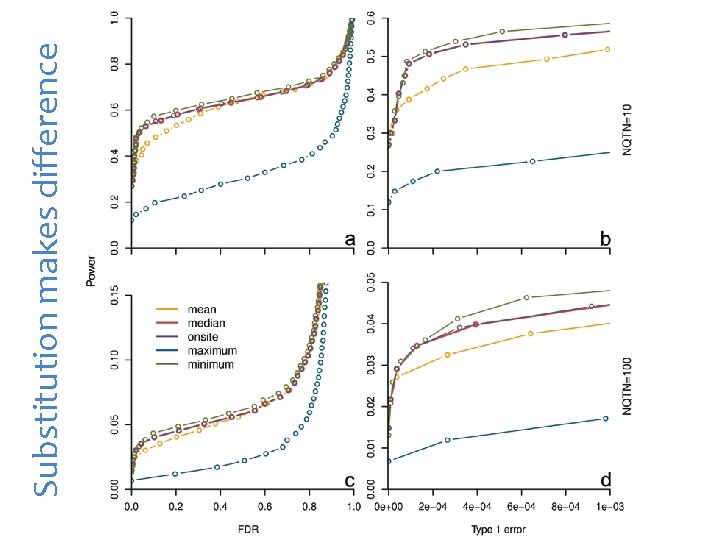
Substitution makes difference

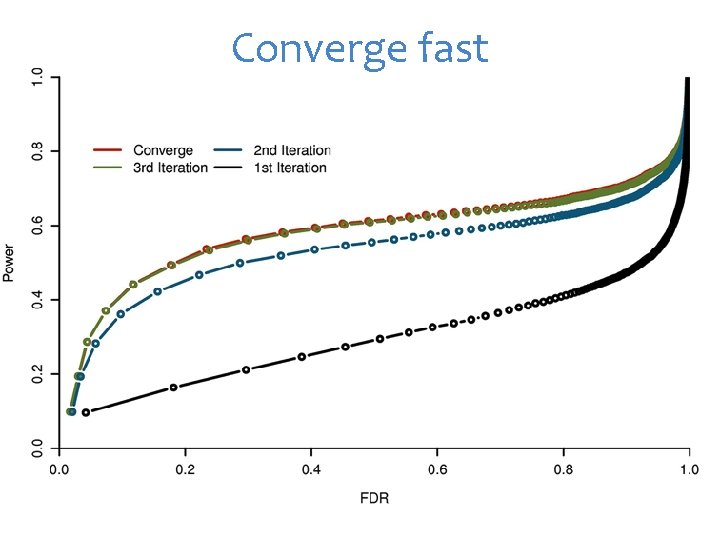
Converge fast

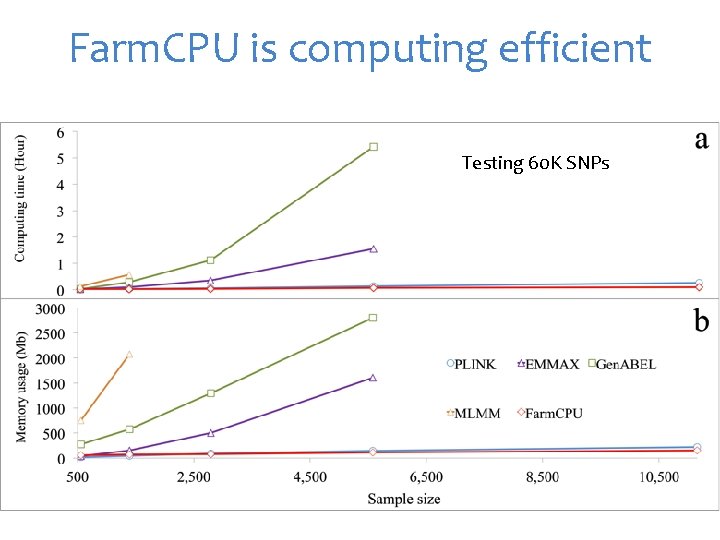
Farm. CPU is computing efficient Testing 60 K SNPs

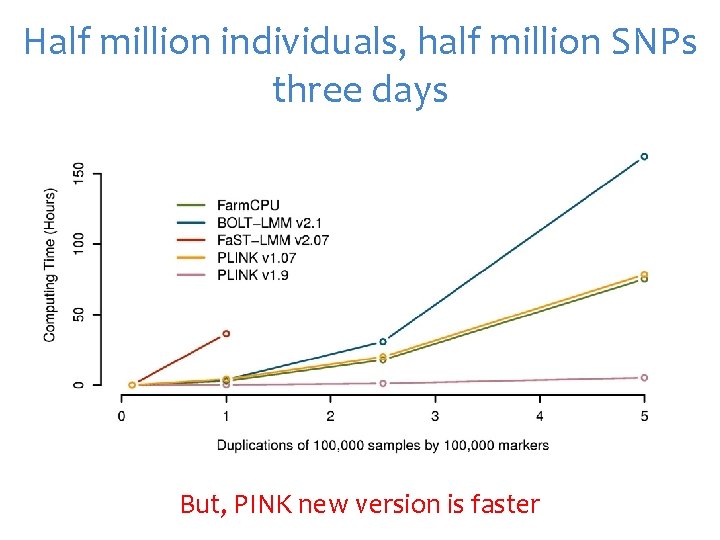
Half million individuals, half million SNPs three days But, PINK new version is faster

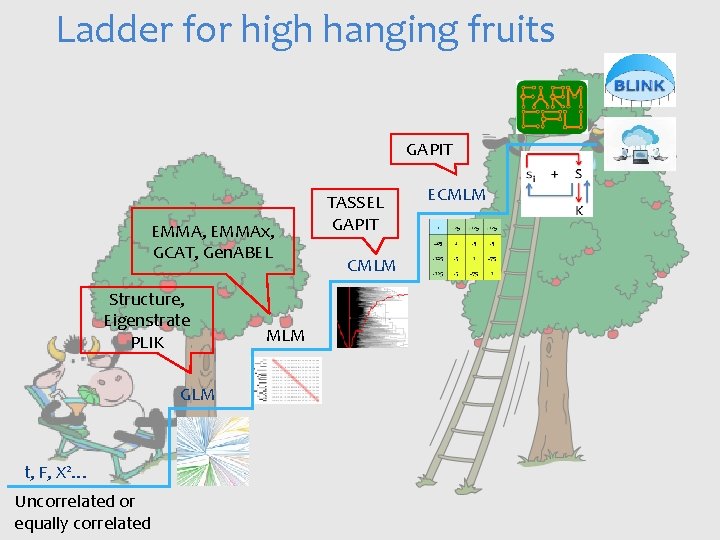
Ladder for high hanging fruits GAPIT EMMA, EMMAx, GCAT, Gen. ABEL Structure, Eigenstrate PLIK GLM t, F, X 2… Uncorrelated or equally correlated MLM TASSEL GAPIT CMLM ECMLM