Statistical Genomics Lecture 20 MLMM Zhiwu Zhang Washington










![Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr] Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr]](https://slidetodoc.com/presentation_image_h2/4cca72d201ffeeed8a886d99423816b8/image-21.jpg)

- Slides: 23

Statistical Genomics Lecture 20: MLMM Zhiwu Zhang Washington State University

Outline Stepwise regression Criteria MLMM Power vs FDR and Type I error Replicate and mean

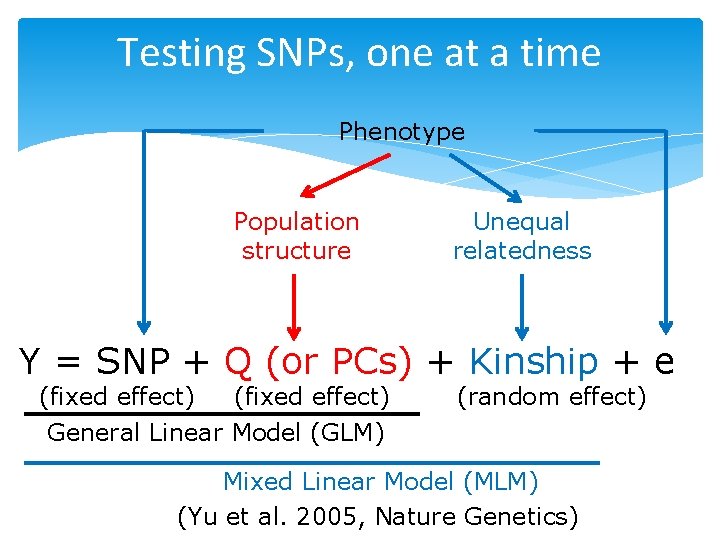
Testing SNPs, one at a time Phenotype Population structure Unequal relatedness Y = SNP + Q (or PCs) + Kinship + e (fixed effect) General Linear Model (GLM) (random effect) Mixed Linear Model (MLM) (Yu et al. 2005, Nature Genetics)

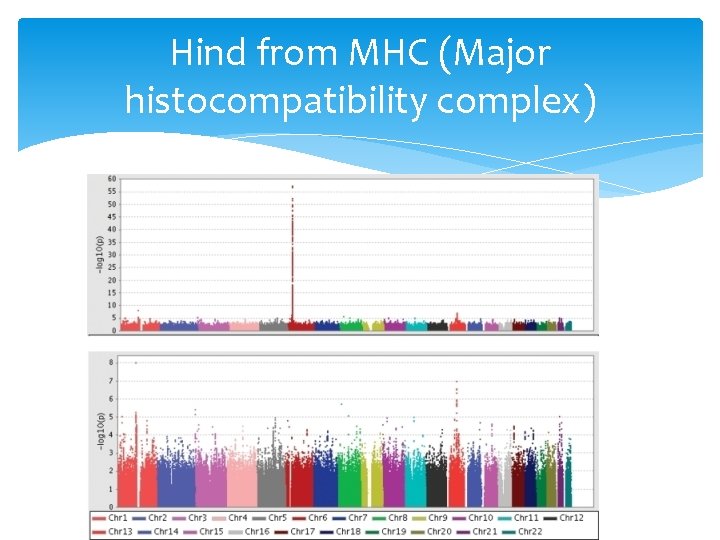
Hind from MHC (Major histocompatibility complex)

Stepwise regression Choose m predictive variables from M (M>>m) variables The challenges : 1. Choosing m from M is an NP problem 2. Option: approximation 3. Non unique criteria

Stepwise regression procedures 1. 2. 3. 4. 5. 6. 7. sequence of F-tests or t-tests Adjusted R-square Akaike information criterion (AIC) Bayesian information criterion (BIC) Mallows's Cp PRESS false discovery rate (FDR) Why so many?

Stepwise regression Forward Test M variables one at a time Fit the most significant variable as covariate Test rest variables one at a time Yes Is the most influential variable significant No End

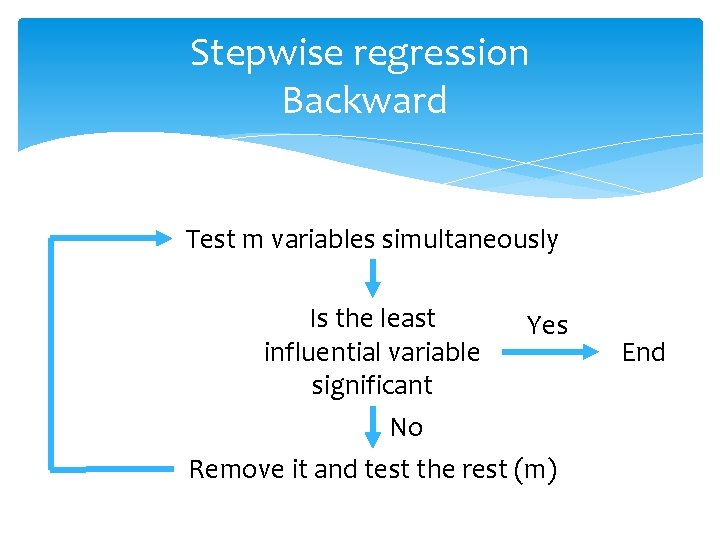
Stepwise regression Backward Test m variables simultaneously Is the least Yes influential variable significant No Remove it and test the rest (m) End

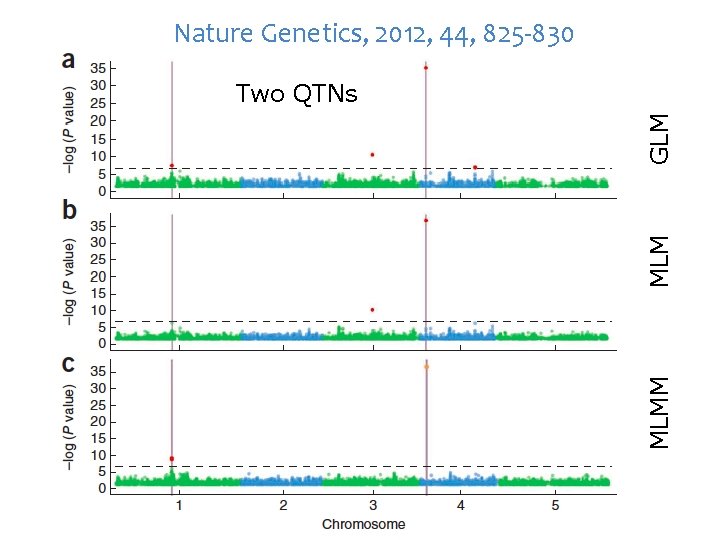
Nature Genetics, 2012, 44, 825 -830 MLMM MLM GLM Two QTNs

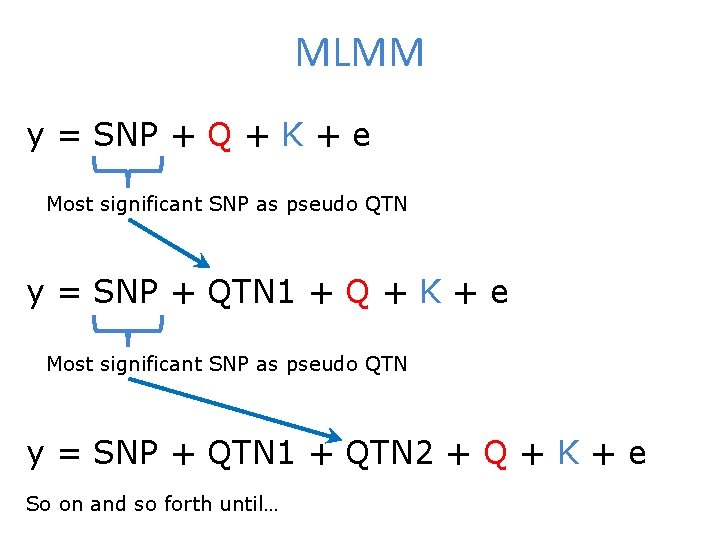
MLMM y = SNP + Q + K + e Most significant SNP as pseudo QTN y = SNP + QTN 1 + QTN 2 + Q + K + e So on and so forth until…

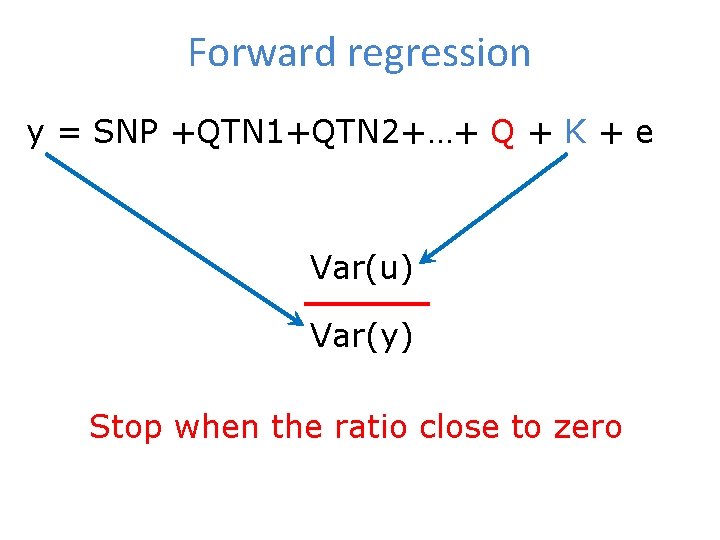
Forward regression y = SNP +QTN 1+QTN 2+…+ Q + K + e Var(u) Var(y) Stop when the ratio close to zero

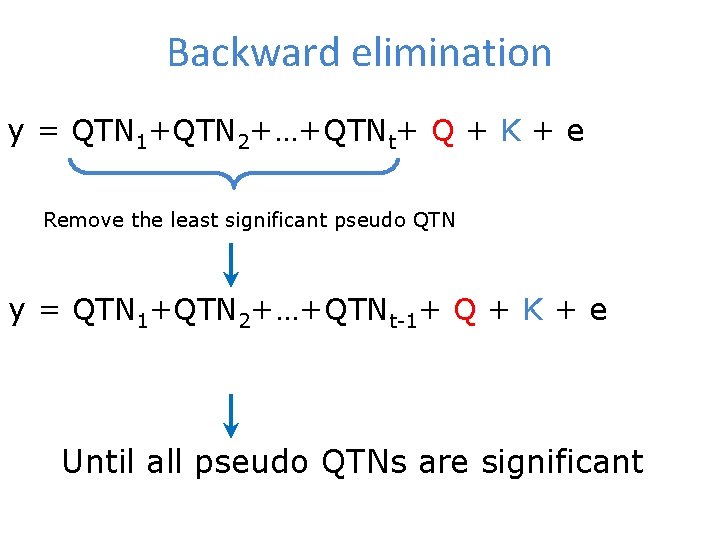
Backward elimination y = QTN 1+QTN 2+…+QTNt+ Q + K + e Remove the least significant pseudo QTN y = QTN 1+QTN 2+…+QTNt-1+ Q + K + e Until all pseudo QTNs are significant

Final p values Pseudo QTNs: y = QTN 1+QTN 2+…+ Q + K + e Other markers: y = SNP +QTN 1+QTN 2+…+ Q + K + e

MLMM R on Git. Hub

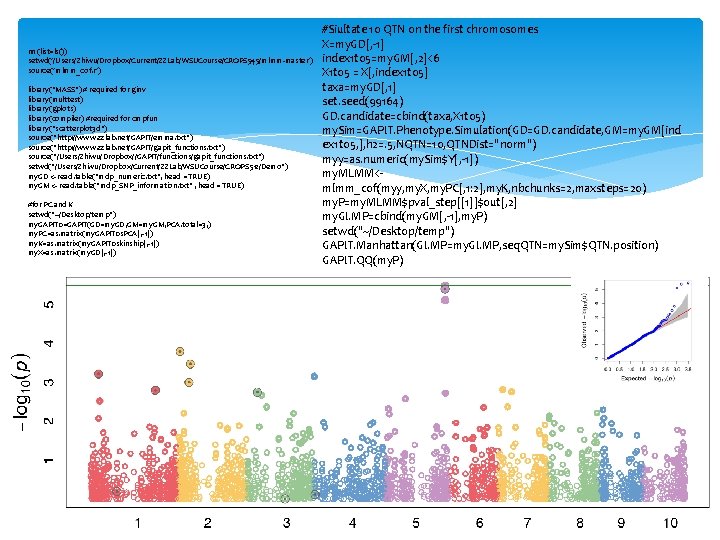
rm(list=ls()) setwd('/Users/Zhiwu/Dropbox/Current/ZZLab/WSUCourse/CROPS 545/mlmm-master') source('mlmm_cof. r') library("MASS") # required for ginv library(multtest) library(gplots) library(compiler) #required for cmpfun library("scatterplot 3 d") source("http: //www. zzlab. net/GAPIT/emma. txt") source("http: //www. zzlab. net/GAPIT/gapit_functions. txt") source("/Users/Zhiwu/Dropbox//GAPIT/functions/gapit_functions. txt") setwd("/Users/Zhiwu/Dropbox/Current/ZZLab/WSUCourse/CROPS 512/Demo") my. GD <- read. table("mdp_numeric. txt", head = TRUE) my. GM <- read. table("mdp_SNP_information. txt" , head = TRUE) #for PC and K setwd("~/Desktop/temp") my. GAPIT 0=GAPIT(GD=my. GD, GM=my. GM, PCA. total=3, ) my. PC=as. matrix(my. GAPIT 0$PCA[, -1]) my. K=as. matrix(my. GAPIT 0$kinship[, -1]) my. X=as. matrix(my. GD[, -1]) #Siultate 10 QTN on the first chromosomes X=my. GD[, -1] index 1 to 5=my. GM[, 2]<6 X 1 to 5 = X[, index 1 to 5] taxa=my. GD[, 1] set. seed(99164) GD. candidate=cbind(taxa, X 1 to 5) my. Sim=GAPIT. Phenotype. Simulation(GD=GD. candidate, GM=my. GM[ind ex 1 to 5, ], h 2=. 5, NQTN=10, QTNDist="norm") myy=as. numeric(my. Sim$Y[, -1]) my. MLMM<mlmm_cof(myy, my. X, my. PC[, 1: 2], my. K, nbchunks=2, maxsteps=20) my. P=my. MLMM$pval_step[[1]]$out[, 2] my. GI. MP=cbind(my. GM[, -1], my. P) setwd("~/Desktop/temp") GAPIT. Manhattan(GI. MP=my. GI. MP, seq. QTN=my. Sim$QTN. position) GAPIT. QQ(my. P)

GAPIT. FDR. Type. I Function my. GWAS=cbind(my. GM, my. P, NA) my. Stat=GAPIT. FDR. Type. I(WS=c(1 e 0, 1 e 3, 1 e 4, 1 e 5), GM=my. GM, seq. QTN=my. Sim$QTN. position, GWAS=my. GWAS)

Return

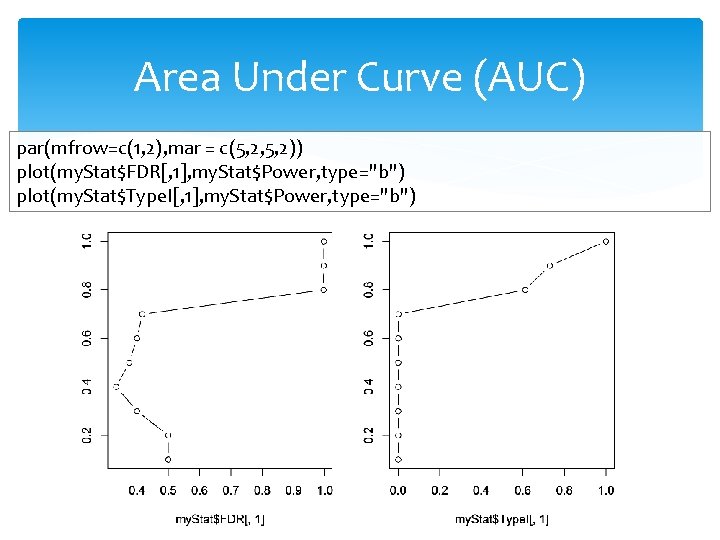
Area Under Curve (AUC) par(mfrow=c(1, 2), mar = c(5, 2, 5, 2)) plot(my. Stat$FDR[, 1], my. Stat$Power, type="b") plot(my. Stat$Type. I[, 1], my. Stat$Power, type="b")

Replicates nrep=10 set. seed(99164) stat. Rep=replicate(nrep, { my. Sim=GAPIT. Phenotype. Simulation(GD=GD. candidate, GM=my. GM[index 1 to 5, ], h 2=. 5, NQTN=10, QTNDist="norm") myy=as. numeric(my. Sim$Y[, -1]) my. MLMM<-mlmm_cof(myy, my. X, my. PC[, 1: 2], my. K, nbchunks=2, maxsteps=20) my. P=my. MLMM$pval_step[[1]]$out[, 2] my. GWAS=cbind(my. GM, my. P, NA) my. Stat=GAPIT. FDR. Type. I(WS=c(1 e 0, 1 e 3, 1 e 4, 1 e 5), GM=my. GM, seq. QTN=my. Sim$QT N. position, GWAS=my. GWAS) })

str(stat. Rep)
![Means over replicates powerstat Rep2 FDR s fdrseq3 lengthstat Rep 7 fdrstat Reps fdr Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr]](https://slidetodoc.com/presentation_image_h2/4cca72d201ffeeed8a886d99423816b8/image-21.jpg)
Means over replicates power=stat. Rep[[2]] #FDR s. fdr=seq(3, length(stat. Rep), 7) fdr=stat. Rep[s. fdr] fdr. mean=Reduce ("+", fdr) / length(fdr) #Type. I s. t 1=seq(4, length(stat. Rep), 7) t 1=stat. Rep[s. t 1] t 1. mean=Reduce ("+", t 1) / length(t 1)

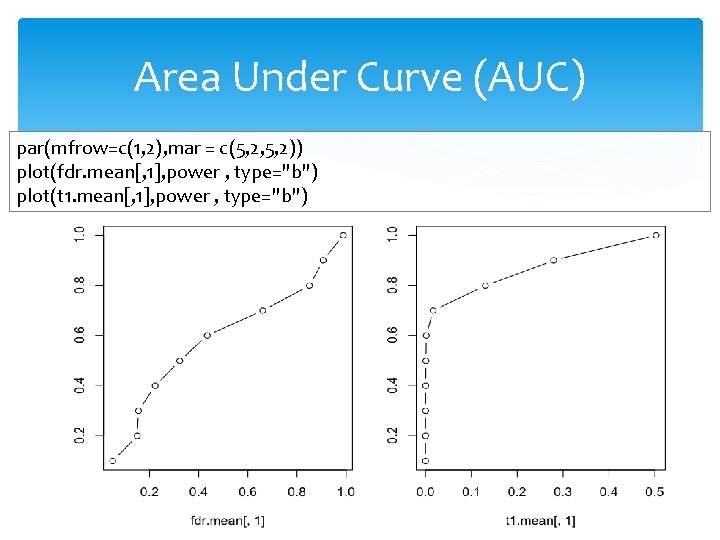
Area Under Curve (AUC) par(mfrow=c(1, 2), mar = c(5, 2, 5, 2)) plot(fdr. mean[, 1], power , type="b") plot(t 1. mean[, 1], power , type="b")

Highlight Stepwise regression Criteria MLMM Power vs FDR and Type I error Replicate and mean