Microbiome Analysis 16 S AND METAGENOMICS Welcome Your

Microbiome Analysis 16 S AND METAGENOMICS ‘

Welcome! Your Tutorial Team: Me (16 S theory) Mike Hall (16 S practical) Morgan Langille (metagenomics theory and practical) Special thanks to: Will Hsiao (CBW presentation) 2
Today’s presentation CBW “Analysis of metagenomic data” http: //bioinformatics. ca/workshops/2015/analysis-metagenomic-data-2015 3

Overview Morning session 1. A brief history of molecules and microbes 2. Why 16 S? 3. How 16 S analysis is usually done 4. Assumptions 5. Hands-on practical Afternoon session 1. 16 S vs Metagenomics 2. Metagenome Taxonomic Composition 3. Metagenome Functional Composition 4. PICRUSt: Functional Inference 5. Hands-on practical 4

Learning objectives At the end of the 16 S tutorial, you should be able to do the following: 1. Run a simple QIIME analysis of a data set (https: //www. dropbox. com/s/kpte 51 nm 17 wav 9 o/stool_data. zip) 2. Interpret analysis results 3. Understand the limitations of the standard 16 S analysis pipeline 5

Defining metagenomics Microbiome: Attributed to Joshua Lederberg by Hooper and Gordon (2001): “the collective genome of our indigenous microbes (microflora), the idea being that a comprehensive genetic view of Homo sapiens as a life-form should include the genes in our microbiome” Is also used to mean microbiota, the group of microorganisms found in a particular setting (usage varies: be careful and precise!) Metagenome: Handelsman et al. (1998) “…advances in molecular biology and eukaryotic genomics, which have laid the groundwork for cloning and functional analysis of the collective genomes of soil microflora, which we term the metagenome of the soil. ” Does not encompass marker-gene surveys (e. g. , 16 S) This report says it does. 6

Micro-what? Metagenomics is often defined to encompass only Bacteria and Archaea (and often Archaea are excluded too!) Other small things to consider: ◦ Viruses / phages ◦ Microbial eukaryotes ◦ Worms (helminths, nematodes, …) Lukeš et al. (2015) PLo. S Pathogens 7

The dawn of metagenomics 3. 5 BYA – the Archaean Eon 16 S position 349 (-ish) ? G Archaea A Bacteria 8
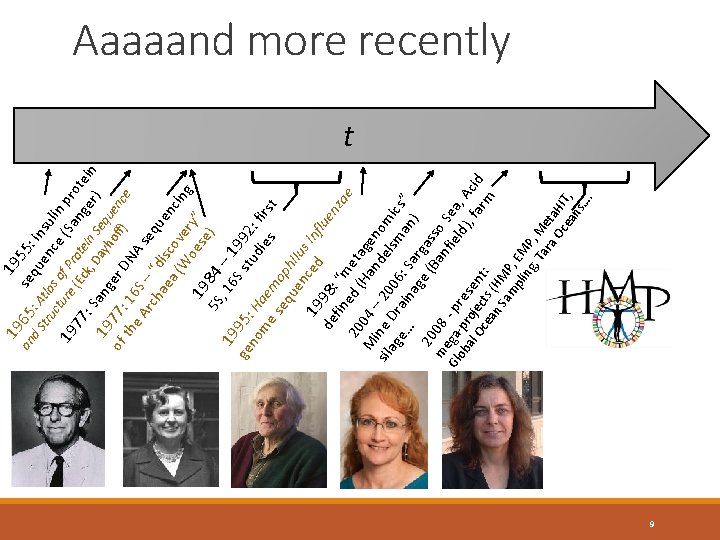
19 an n S HM am P, pli EM ng P , T , M ar et a O a. H ce IT, an s…. 65 se 55: qu in d. S : A en su tru tla ce lin ctu s o f ( p 19 r 77 e (Ec Prote Sang rote k, in e in : 19 Sang Dayh Sequ r) of 77: er D off) enc e th 16 e A S NA rch – “ seq ae disc uen a ( ov W er cing 19 oese y” ) 5 S 84 , – 19 16 1 S s 99 9 ge 5: tu 2: no H die fir a m e st s es m eq oph ue ilu n 19 ced s inf lue de 98: nz fin “m ae 20 ed ( eta M 04 – Han gen d o sil ine 20 ag Dr 06 elsm mic e… ain : S a s” ag arg n) e ( as 20 Ba so 0 m 8 nf Se iel a Gl ega- - pr ob pr e d) , A al oj se , fa cid Oc ect nt rm ea s ( : 19 Aaaaand more recently t 9

The 16 S ribosomal RNA gene THE FIRST WORD IN MICROBIAL BIODIVERSITY 10
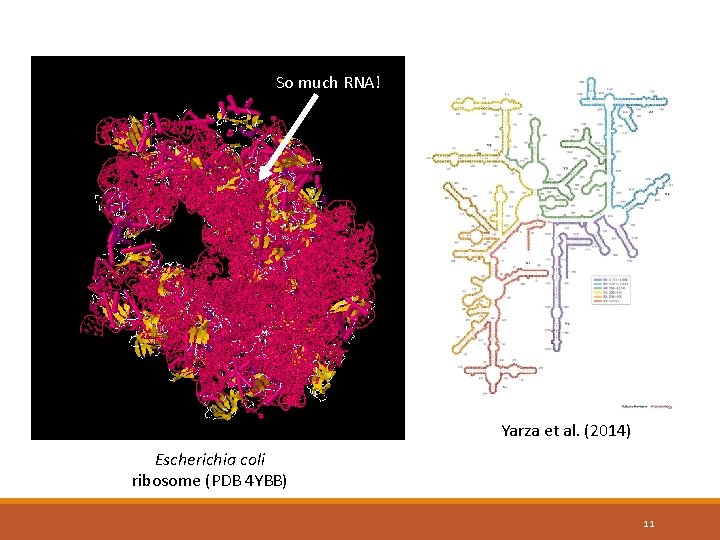
So much RNA! Yarza et al. (2014) Escherichia coli ribosome (PDB 4 YBB) 11

Why 16 S? The “universal phylogenetic marker” (1) Present in all living organisms (2) Single copy* (no recombination) (3) Highly conserved + highly variable regions (4) Huge reference databases 12
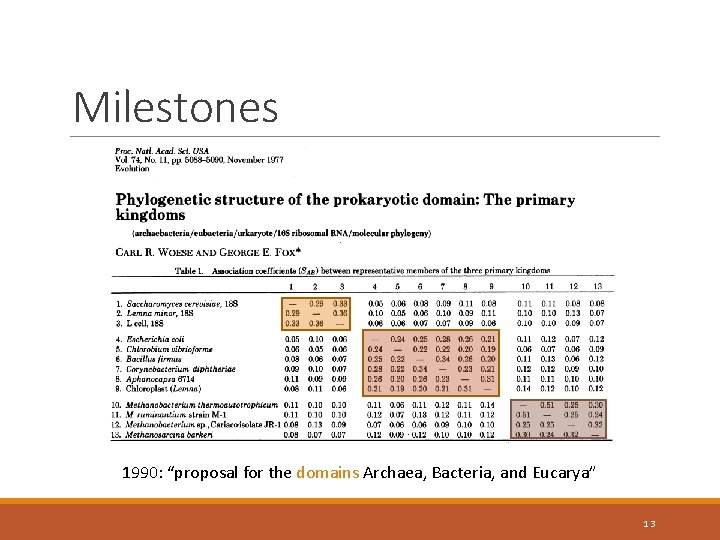
Milestones 1990: “proposal for the domains Archaea, Bacteria, and Eucarya” 13

Milestones 2002: “…as much as 50% of the total surface microbial community…” Nature (1990) 14

Milestones Many critical papers followed (error filtering, clustering approaches, …) PNAS (2006) 15

Milestones + 681 metagenomic samples Huttenhower, Gevers et al. (2012) 16

16 S analysis HOW IT’S DONE 17
Your basic workflow Sample collection DNA extraction Amplification Analysis 18

Sample collection and DNA extraction Defined protocols exist, many kits (e. g. Power. Soil®) Need to consider barriers to DNA recovery and PCR (e. g. humic acids from soil, bile salts from feces) Additional mechanical approaches (e. g. , mechanical lysis of tissues with bead beating) Kits and rogue lab DNA can end up in your sample – need to run negative controls!! ◦ Example from [year redacted]: shocking finding of bacterial DNA in the [location redacted]! However, [taxonomic group redacted] was a known frequent contaminant of DNA extraction kits. 19

Size fractionation http: //www. jove. com/video/52685/automated-gel-size-selection-to-improve-quality-nextgeneration 20

Choosing a PCR strategy Need to consider: ◦ Correct melting temperature (60 -65 degrees C for Illumina protocol) ◦ DNA sequencing read length (influences choice of primers) ◦ Primer specificity! ◦ Comparability with previous studies? [Good luck with that] [but that’s what the Earth Microbiome Project protocol http: //www. earthmicrobiome. org/emp-standard-protocols/16 s/ is meant to achieve] 21
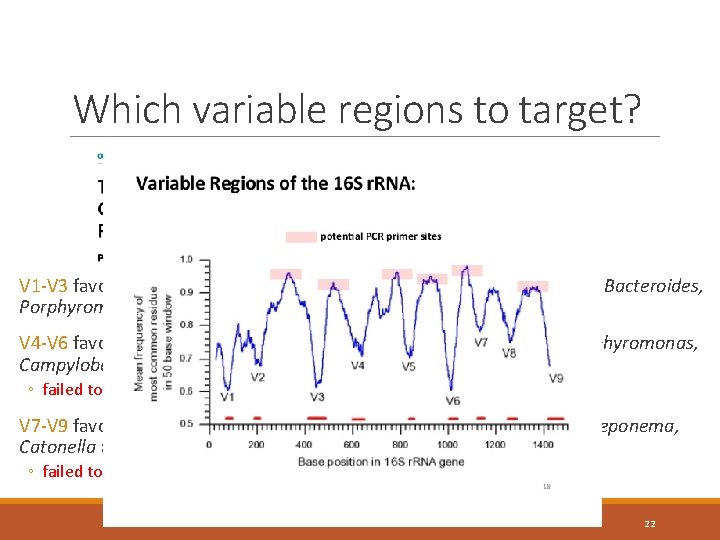
Which variable regions to target? V 1 -V 3 favours Prevotella, Fusobacterium, Streptococcus, Granulicatella, Bacteroides, Porphyromonas and Treponema V 4 -V 6 favours Streptococcus, Treponema, Prevotella, Eubacterium, Porphyromonas, Campylobacter and Enterococcus. ◦ failed to detect Fusobacterium V 7 -V 9 favours Veillonella, Streptococcus, Eubacterium, Enterococcus, Treponema, Catonella and Selenomonas. ◦ failed to detect Selenomonas, TM 7 and Mycoplasma 22

At least there’s no shortage of options… Detailed in silico evaluation of primers, experimental evaluation of two sets Heavily biased recovery of Bacteria, Archaea, and missing groups depending on primer choice. “Out of the 175 primers and 512 primer pairs checked, only 10 can be recommended as broad-range primers. ” 23

Amplification Example: Illumina protocol 24

Analysis (examples mostly from QIIME) 1. Quality Control ◦ Error checking 2. Sample diversity ◦ ◦ Taxonomy agnostic Taxonomy aware 3. Similarity among samples 4. Associations with metadata/groups (ANOSIM, MRPP) 5. Machine-learning classification 6. Functional prediction 25
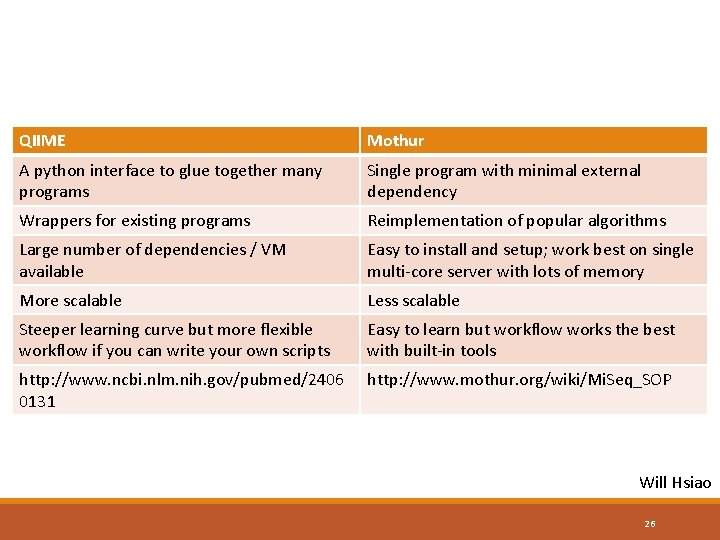
QIIME Mothur A python interface to glue together many programs Single program with minimal external dependency Wrappers for existing programs Reimplementation of popular algorithms Large number of dependencies / VM available Easy to install and setup; work best on single multi-core server with lots of memory More scalable Less scalable Steeper learning curve but more flexible workflow if you can write your own scripts Easy to learn but workflow works the best with built-in tools http: //www. ncbi. nlm. nih. gov/pubmed/2406 0131 http: //www. mothur. org/wiki/Mi. Seq_SOP Will Hsiao 26

“Analysis” #1 Quality Control Quality score filtering: ◦ ◦ Minimal length of consecutive high-quality bases (as % of total read length) Maximal number of consecutive low-quality bases Maximal number of ambiguous bases (N’s) Minimum Phred quality score Other quality filtering tools available ◦ Cutadapt (https: //github. com/marcelm/cutadapt) ◦ Trimmomatic (http: //www. usadellab. org/cms/? page=trimmomatic) ◦ Sickle (https: //github. com/najoshi/sickle) Chimera checking: ◦ UCHIME 27
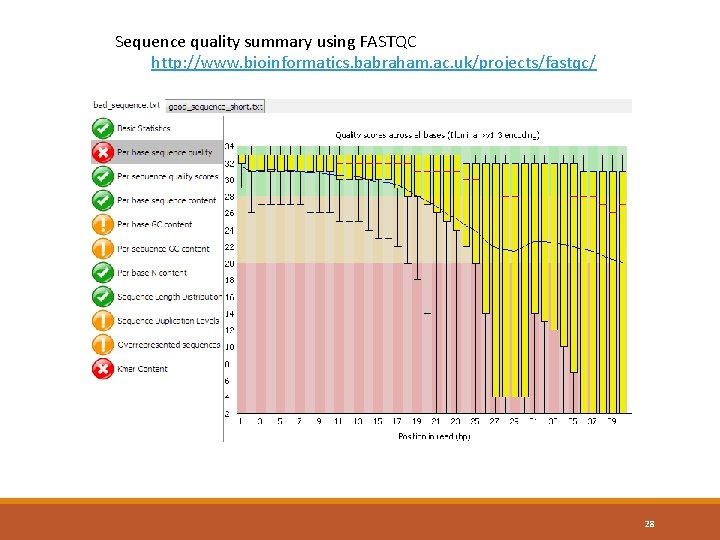
Sequence quality summary using FASTQC http: //www. bioinformatics. babraham. ac. uk/projects/fastqc/ 28

Analysis #2 Within-sample (“alpha”) diversity To describe the diversity of a sample, you need to know what you are counting! Individual sequences? ◦ Most precise, but vulnerable to sequencing error effects – inflation of diversity Clusters of sequences? ◦ Operational taxonomic units (OTUs) – 97% sequence identity as the “species” level of similarity Taxonomic groups? ◦ It’s always reassuring to put names on things, but taxonomic labels can be extremely misleading 29
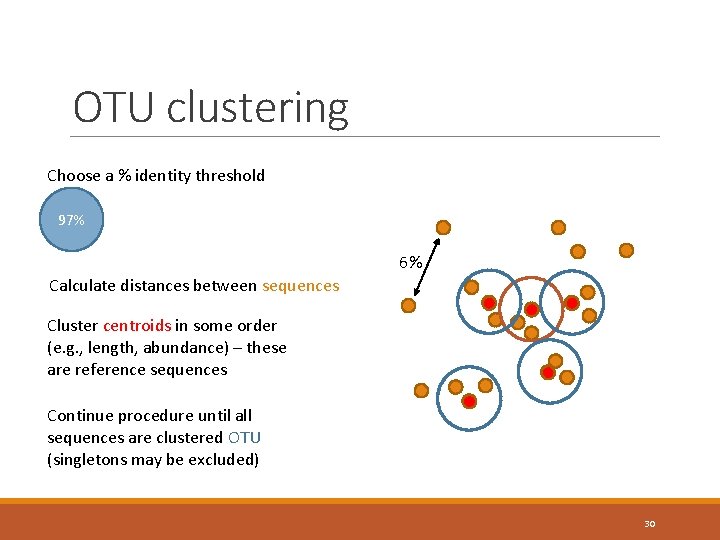
OTU clustering Choose a % identity threshold 97% 6% Calculate distances between sequences Cluster centroids in some order (e. g. , length, abundance) – these are reference sequences Continue procedure until all sequences are clustered OTU (singletons may be excluded) 30

What’s in a name? Akkermansia ? ? ? Ruminococcus Parabacteroides ? ? ? Bacteroides 31

Taxonomic assignment Many choices: BLAST – assign taxonomic label of closest match (simple, possibly too simple) Phylogenetic placement – e. g. Pplacer (Matsen et al. , BMC Bioinformatics 2010) Machine-learning classification, in particular Naïve Bayes e. g. RDP Classifier, Wang et al. (2007) BMC Bioinformatics 32

Example RDP Classifier output GD 6 JEAT 01 AYGPE "Planctomycetes" Planctomycetales Schlesneria Root phylum order genus rootrank 1. 0 Bacteria domain 1. 0 "Planctomycetacia"class 1. 0 Planctomycetaceaefamily 1. 0 0. 96 GD 6 JEAT 01 BEUG 6 Firmicutes Clostridiales Anaerotruncus Root phylum order genus rootrank 1. 0 Bacteria domain 1. 0 0. 32 Clostridia class 0. 26 0. 23 Ruminococcaceae family 0. 22 0. 19 Includes bootstrap support 33

Calculating alpha diversity OTU counts – richness only Simpson index – probability of sampling two individuals of the same type Phylogenetic diversity – sum of branch lengths 34
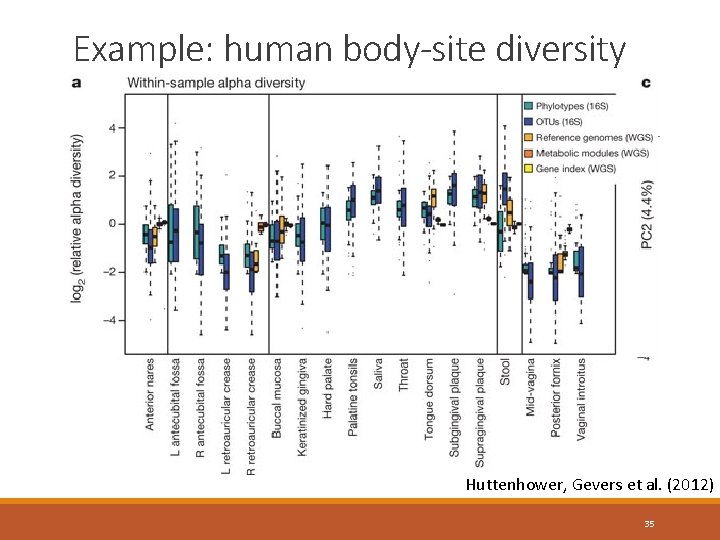
Example: human body-site diversity Huttenhower, Gevers et al. (2012) 35

Analysis #3 Among-sample (“beta”) diversity 1. Perform pairwise comparisons between all samples to build a dissimilarity matrix 2. Summarize the matrix using based on major patterns of covariance or hierarchical similarity 36

Analysis #3 Among-sample (“beta”) diversity Given a pair of samples (described as e. g. OTU abundance), calculate their dissimilarity Beta-diversity measures can be: ◦ non-phylogenetic or phylogenetic ◦ weighted or unweighted There a lot of measures! - Bray-Curtis (weighted, non-phylogenetic) - Jaccard (unweighted, non-phylogenetic) - Weighted Uni. Frac (weighted, phylogenetic) -… 37
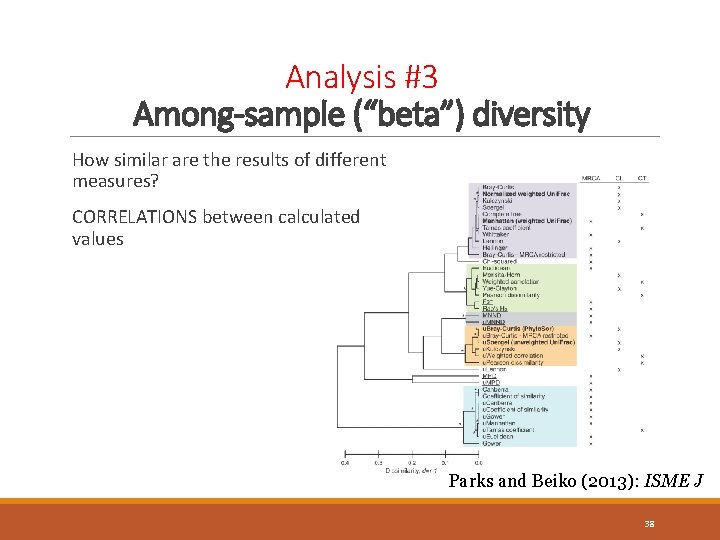
Analysis #3 Among-sample (“beta”) diversity How similar are the results of different measures? CORRELATIONS between calculated values Parks and Beiko (2013): ISME J 38
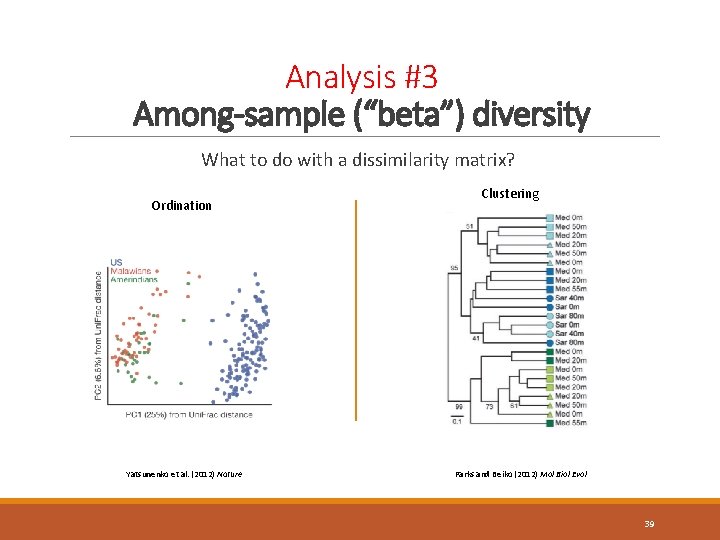
Analysis #3 Among-sample (“beta”) diversity What to do with a dissimilarity matrix? Ordination Yatsunenko et al. (2012) Nature Clustering Parks and Beiko (2012) Mol Biol Evol 39
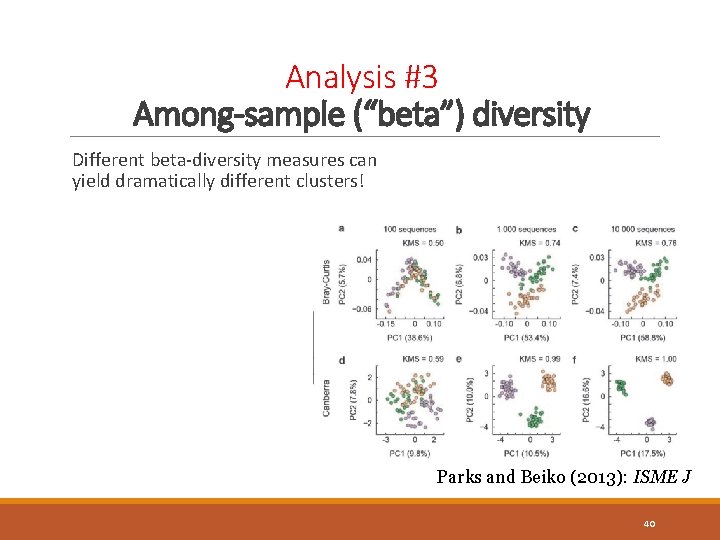
Analysis #3 Among-sample (“beta”) diversity Different beta-diversity measures can yield dramatically different clusters! Parks and Beiko (2013): ISME J 40

Analysis #4 Associations with metadata PERMANOVA: Permutational multivariate analysis of variance ANOSIM: Rank-based analysis of similarity Mantel test: Comparison of between-group vs within-group distances Example: Weighted Uni. Frac distance: root compartment explains 46. 62% of variance (PERMANOVA p<0. 001) Unweighted Uni. Frac: root compartment explains only 18. 07% of variance (PERMANOVA p<0. 001); soil type is more important Good review: Anderson and Walsh (2013) Ecological Monographs 41

Analysis #5 Machine-learning classification Identify aspects of community structure that are predictive of sample attributes Advantages of machine-learning approaches: ◦ Non-linear combinations of variables ◦ Data transformations ◦ Can accommodate many different representations of the data Disadvantages: ◦ Complex, may “overfit” ◦ Can be time consuming ◦ Obfuscation of predictive rules 42
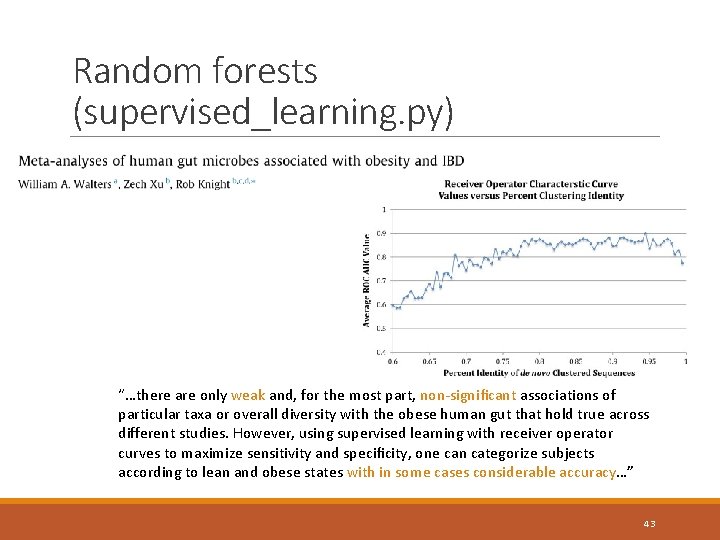
Random forests (supervised_learning. py) “…there are only weak and, for the most part, non-significant associations of particular taxa or overall diversity with the obese human gut that hold true across different studies. However, using supervised learning with receiver operator curves to maximize sensitivity and specificity, one can categorize subjects according to lean and obese states with in some cases considerable accuracy…” 43

Tree-based classifications Nested clade analysis and feature selection Classification of plaque samples using support vector machines Ning and Beiko (2015): Microbiome 44

Analysis #6 Functional prediction PICRUSt: Langille et al (2013) Nat Biotechnol Morgan can tell you about this… 45

Assumptions THAT ARE OFTEN FALSE 46

Do not assume that #1: 16 S is an effective proxy for microbial diversity. #2: All 16 S studies are created equal, with results that are comparable. #3: Rarefaction is a good idea. #4: 16 S OTUs describe ecologically cohesive units (“species”? ). #5: The 16 S tree is the “Tree of Life”. 47

Assumption #1 16 S is an effective proxy for microbial diversity. Variation: Coenye and Vandamme (2003) Estimating copy number: Kembel et al. (2012) and PICRUSt (coming up later) rrn. DB: Stoddard et al. NAR (2014) 48

Assumption #1 16 S is an effective proxy for microbial diversity. Alternative marker genes: cpn 60, rpo. B, … Smaller reference databases! Protein-coding genes! 49

Assumption #2 All 16 S studies are created equal. Effects of sequencing platform, V region, amplicon vs metagenomics Tremblay et al. (2015) Front Microbiol 50

Assumption #3 Rarefaction is a good idea. Example of statistics before and after rarefaction: Loss of statistical power Random subsampling can increase false-positive differences Arbitrary minimum library size chosen for downsampling Alternatives e. g. Negative Binomial fitting (e. g. , De. Seq 2) Mc. Murdie and Holmes (2014) PLo. S Comp Biol 51
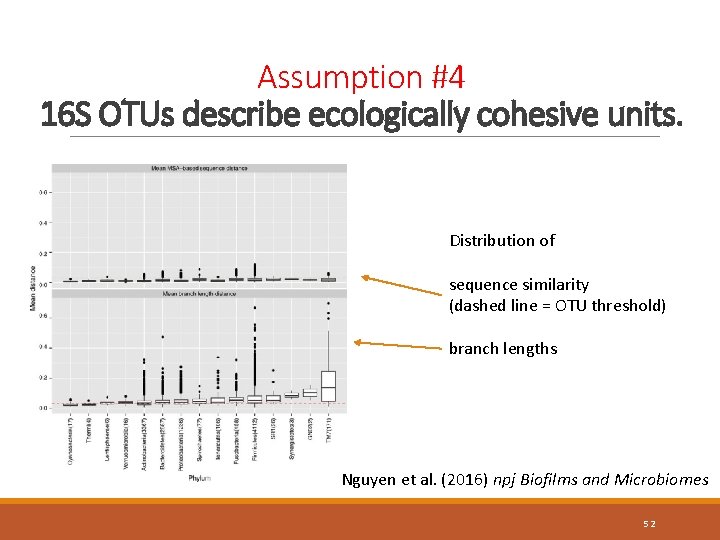
Assumption #4 16 S OTUs describe ecologically cohesive units. Distribution of sequence similarity (dashed line = OTU threshold) branch lengths Nguyen et al. (2016) npj Biofilms and Microbiomes 52
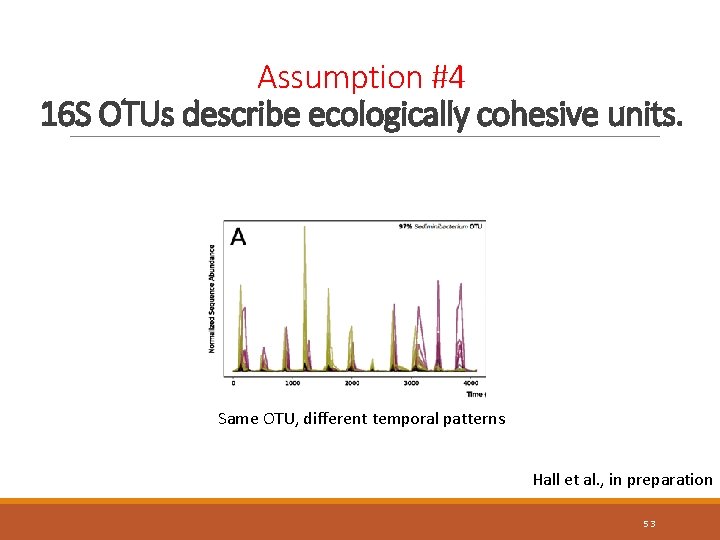
Assumption #4 16 S OTUs describe ecologically cohesive units. Same OTU, different temporal patterns Hall et al. , in preparation 53
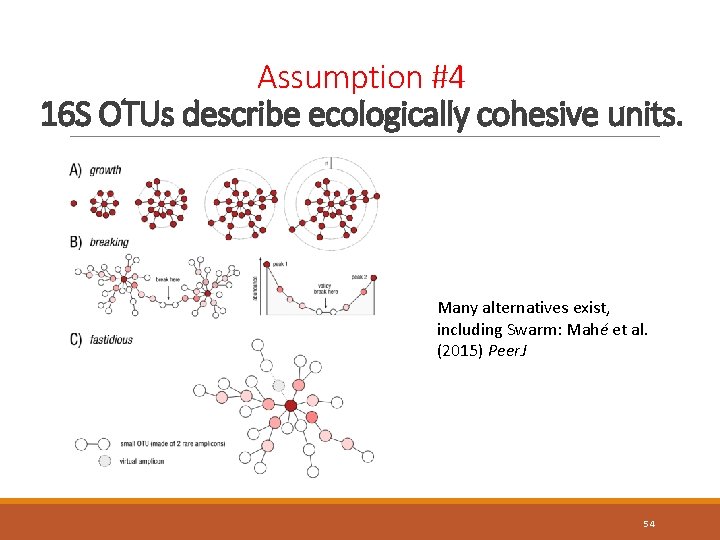
Assumption #4 16 S OTUs describe ecologically cohesive units. Many alternatives exist, including Swarm: Mahé et al. (2015) Peer. J 54

Assumption #5 The 16 S tree is the “Tree of Life”. 16 S is limited for several reasons: Limited resolving power Subject to compositional bias Subject to recombination and lateral transfer Models typically applied to proteincoding genes do not make sense for noncoding RNA 55

Moving On ADVENTURES IN “MULTI-OMICS” 56

Multi-omics? ? 16 S can profile the biodiversity of a microbial sample… But we need the metagenome to shine a light on function… The metatranscriptome tells us what is expressed under specific conditions… And the metaproteome can quantify the relative abundance of different enzymes… While the metabolome focuses on the products of metabolism. What do we really need? 57
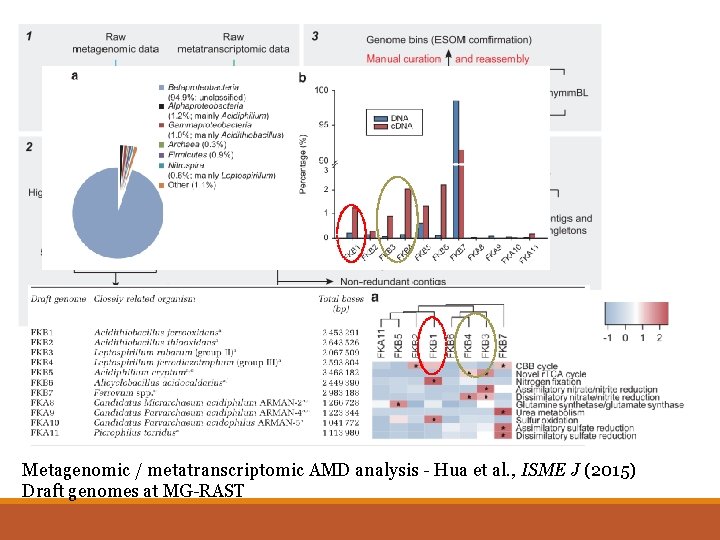
Metagenomic / metatranscriptomic AMD analysis - Hua et al. , ISME J (2015) Draft genomes at MG-RAST
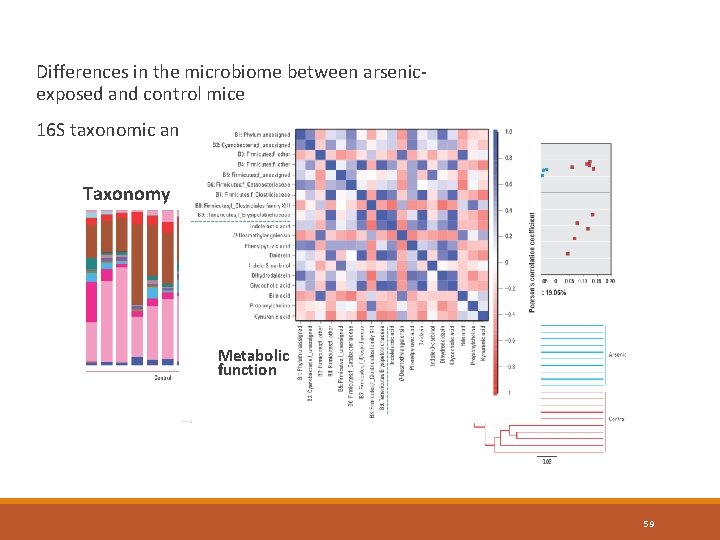
Differences in the microbiome between arsenicexposed and control mice 16 S taxonomic analysis + metabolomics Taxonomy Metabolic function 59

Hands on! LET’S MAKE SCIENCE HAPPEN 60
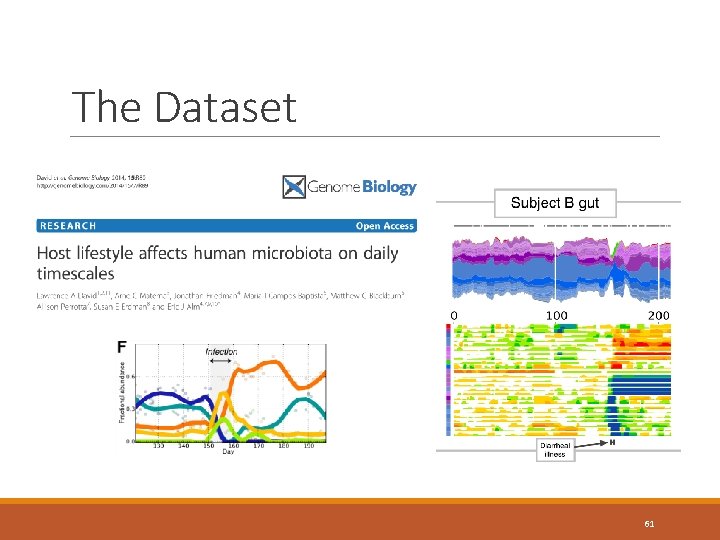
The Dataset 61

Workflow 1. Retrieve data 2. Cluster sequences 3. Taxonomic classification 4. Phylogenetic tree construction 5. OTU table creation 6. Downstream visualization / analysis 62

Presentations http: //www. slideshare. net/Mick. Watson/studying-the-microbiome http: //bioinformatics. ca/metagenomics 2015 module 2 pptx FIN 63
- Slides: 63