Metagenomics Bench and data analysis concepts historical milestones

Metagenomics Bench and data analysis: concepts, historical milestones and next advances Center of Astrobiology, Madrid Laboratory of Molecular Adaptation Eduardo González-Pastor TGAC Norwich, 2014 Metagenomics: From the Bench to Data Analysis

OUTLINE 1. Introduction • What is the metagenome? • Why and how to study the metagenome? Sequence Functional analysis 2. Functional metagenomic approach to search for novel mechanisms of adaptation to extreme environments • Metal and acid resistance mechanisms in microbial communities from the Rio Tinto (Spain)

What is the metagenome? metagenome: the genomes of all the microorganisms (virus included) of an environmental sample, and it is studied using culture independent techniques “metagenomics” Handelsman, J. ; Rondon, M. R. ; Brady, S. F. ; Clardy, J. ; Goodman, R. M. (1998). "Molecular biological access to the chemistry of unknown soil microbes: A new frontier for natural products". Chemistry & Biology 5 (10): 245– 249.

Why to study the metagenome? Only a small percentage of the microorganisms can be cultured (around 1%) (Pace et al. , 1985). For instance, soil microbial communities could contain between 5, 000 and 20, 000 different species, but only few can be isolated and cultured (50 -200) The study of the metagenome provides culture independent information about the microorganisms of an environmental sample.
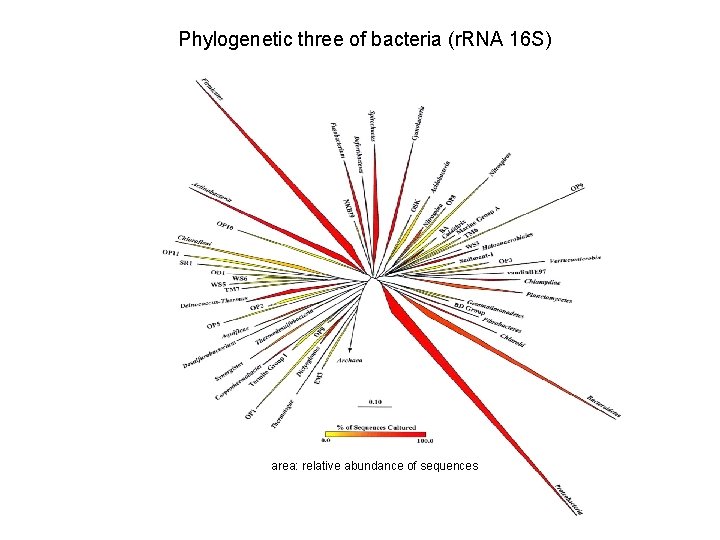
Phylogenetic three of bacteria (r. RNA 16 S) area: relative abundance of sequences
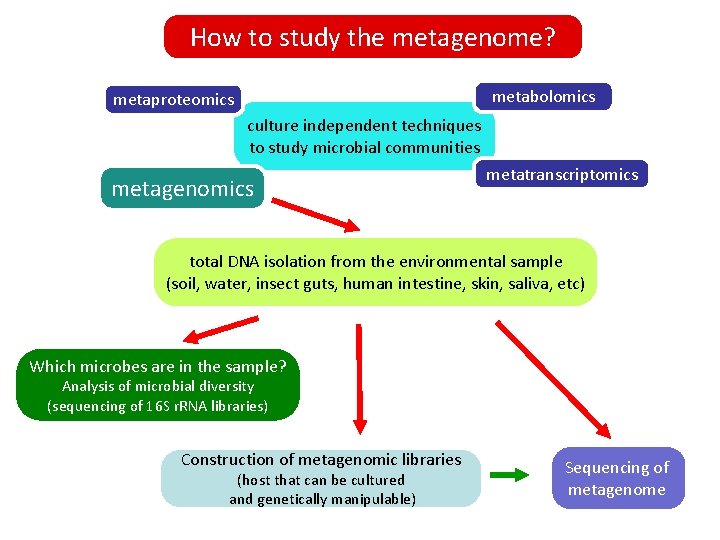
How to study the metagenome? metabolomics metaproteomics culture independent techniques to study microbial communities metagenomics metatranscriptomics total DNA isolation from the environmental sample (soil, water, insect guts, human intestine, skin, saliva, etc) Which microbes are in the sample? Analysis of microbial diversity (sequencing of 16 S r. RNA libraries) Construction of metagenomic libraries (host that can be cultured and genetically manipulable) Sequencing of metagenome
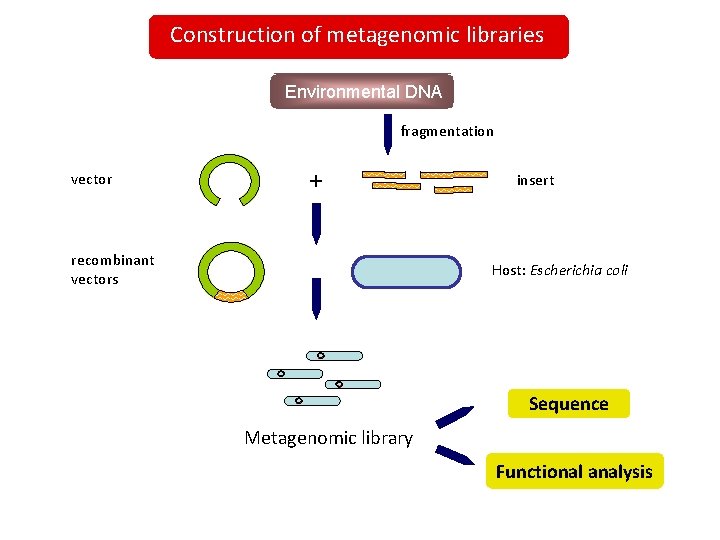
Construction of metagenomic libraries Environmental DNA fragmentation vector + recombinant vectors insert Host: Escherichia coli Sequence Metagenomic library Functional analysis

Selecting the appropiate protocol 1. Sample type • Liquid, solid (soil, sediment, etc), faeces 2. DNA/RNA • • • From raw sample After matrix/cell separation Extraction of DNA or DNA/RNA together 3. Vector type • • • Short insert (phagemid or plasmid) Large insert (fosmid of cosmid) Mega-large insert (p. BAC) 4. DNA digestion • • Enzymatic Physical 5. Microbial host • • • Escherichia coli Pseudomonas putida Bacillus subtilis Streptomyces Pichia pastoris Extraction method
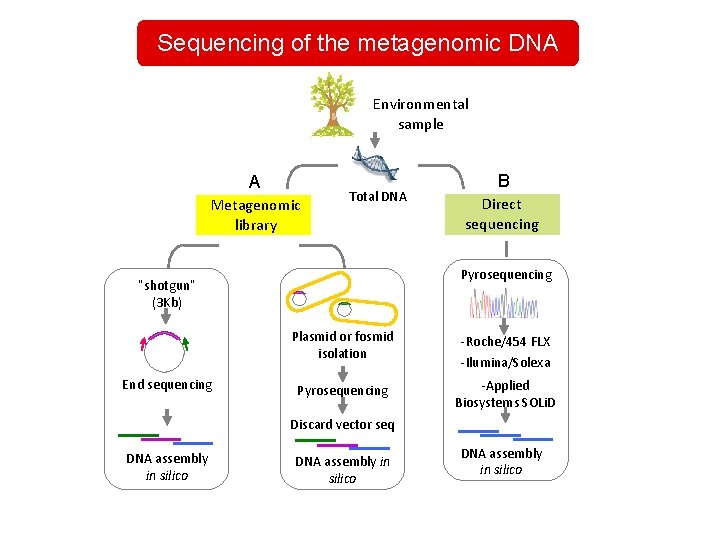
Sequencing of the metagenomic DNA Environmental sample A Metagenomic library Total DNA Direct sequencing Pyrosequencing “shotgun” (3 Kb) End sequencing B Plasmid or fosmid isolation -Roche/454 FLX -Ilumina/Solexa Pyrosequencing -Applied Biosystems SOLi. D Discard vector seq DNA assembly in silico

Sequencing of the metagenomic DNA Bioinformatic analysis: • gene annotation • genome and metabolism reconstruction of microbial communities, • comparation of microbial communities from different environments
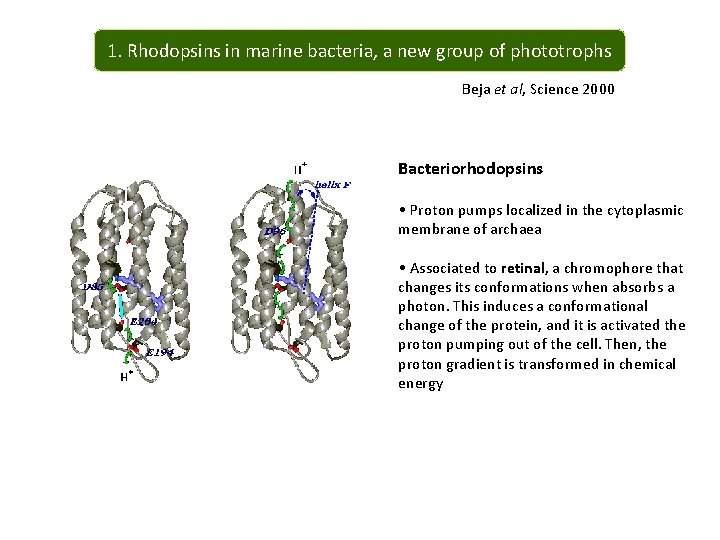
1. Rhodopsins in marine bacteria, a new group of phototrophs Beja et al, Science 2000 Bacteriorhodopsins • Proton pumps localized in the cytoplasmic membrane of archaea • Associated to retinal, a chromophore that changes its conformations when absorbs a photon. This induces a conformational change of the protein, and it is activated the proton pumping out of the cell. Then, the proton gradient is transformed in chemical energy
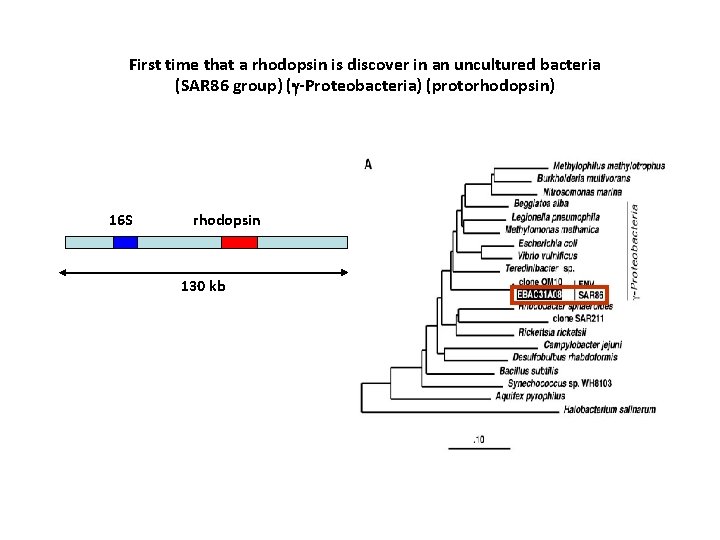
First time that a rhodopsin is discover in an uncultured bacteria (SAR 86 group) (g-Proteobacteria) (protorhodopsin) 16 S rhodopsin 130 kb
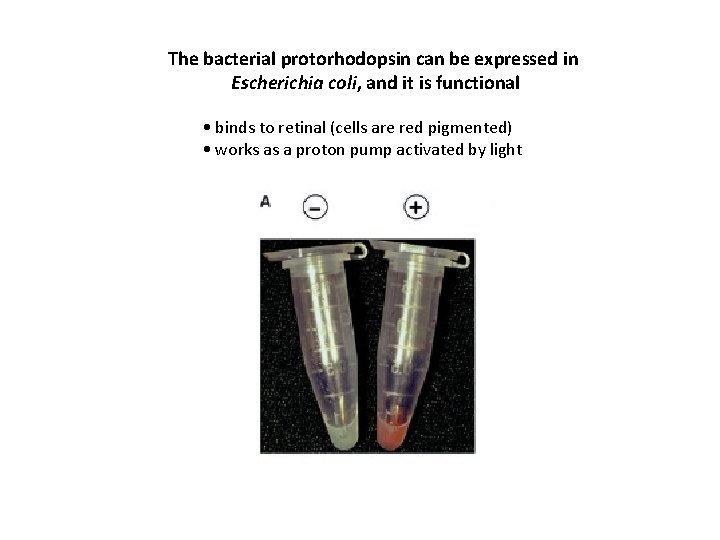
The bacterial protorhodopsin can be expressed in Escherichia coli, and it is functional • binds to retinal (cells are red pigmented) • works as a proton pump activated by light

2. Sequencing of the microbial communities from the Sargasso sea Venter et al. , Science 2004 Microorganisms were collected from the Sargasso sea Metagenomic DNA is fractionated and libraries are constructed with inserts from 2 -6 kbp (“shotgun” sequencing, pairwise-end sequencing) • Weatherbird II: 1. 66 million sequences (1. 36 Gbp) • Sorcerer II: 325, 561 sequences (265 Mbp) 1800 species o phylotypes (148 new)
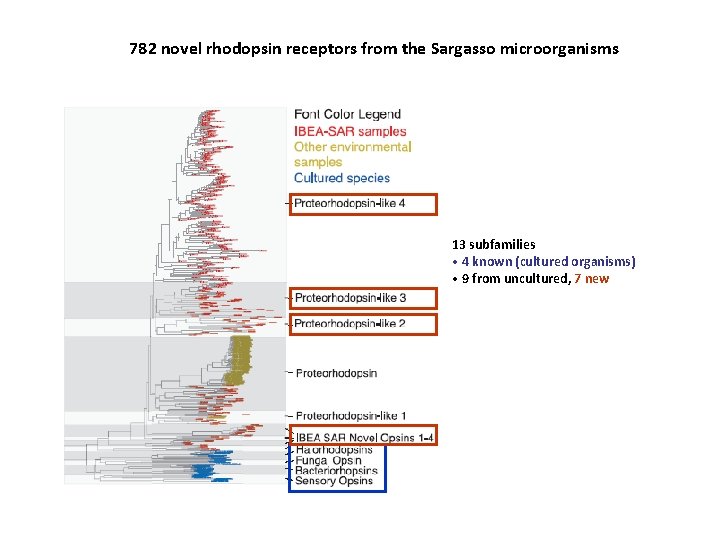
782 novel rhodopsin receptors from the Sargasso microorganisms 13 subfamilies • 4 known (cultured organisms) • 9 from uncultured, 7 new
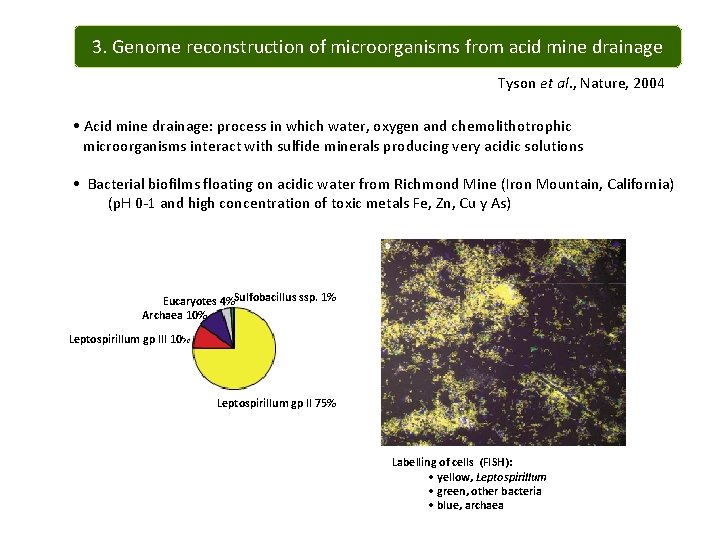
3. Genome reconstruction of microorganisms from acid mine drainage Tyson et al. , Nature, 2004 • Acid mine drainage: process in which water, oxygen and chemolithotrophic microorganisms interact with sulfide minerals producing very acidic solutions • Bacterial biofilms floating on acidic water from Richmond Mine (Iron Mountain, California) (p. H 0 -1 and high concentration of toxic metals Fe, Zn, Cu y As) Eucaryotes 4%Sulfobacillus ssp. 1% Archaea 10% Leptospirillum gp III 10% Leptospirillum gp II 75% Labelling of cells (FISH): • yellow, Leptospirillum • green, other bacteria • blue, archaea
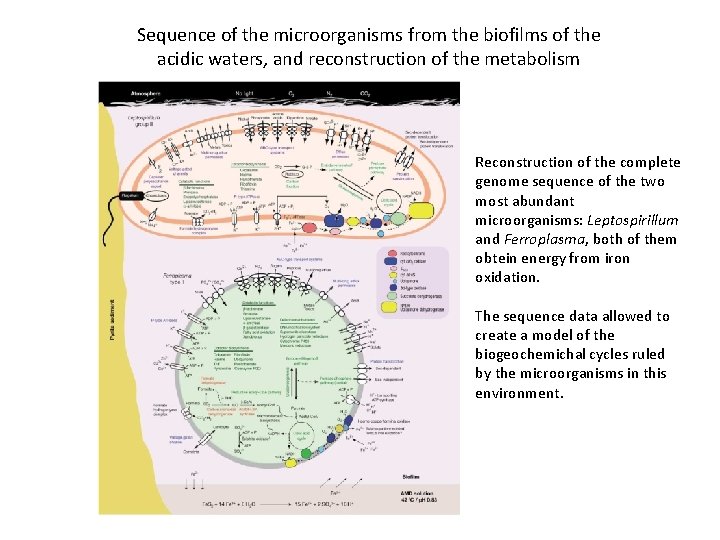
Sequence of the microorganisms from the biofilms of the acidic waters, and reconstruction of the metabolism Reconstruction of the complete genome sequence of the two most abundant microorganisms: Leptospirillum and Ferroplasma, both of them obtein energy from iron oxidation. The sequence data allowed to create a model of the biogeochemichal cycles ruled by the microorganisms in this environment.

4. Comparative metagenomics of microbial communities Tringe et al. , Science 2006 • Comparison of unassembled sequence data obtained from shotgun sequencing DNA isolated from different environments. • Quantitative gene content analysis (abundance or absence) reveals habitat specific fingerprints that reflects known characteristics of the sampled environment • Identification of genes or metabolic pathways specific for a particular environment.
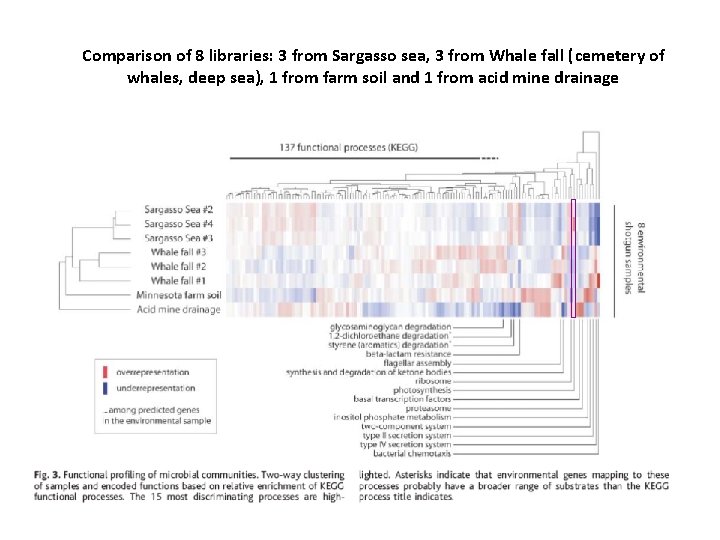
Comparison of 8 libraries: 3 from Sargasso sea, 3 from Whale fall (cemetery of whales, deep sea), 1 from farm soil and 1 from acid mine drainage
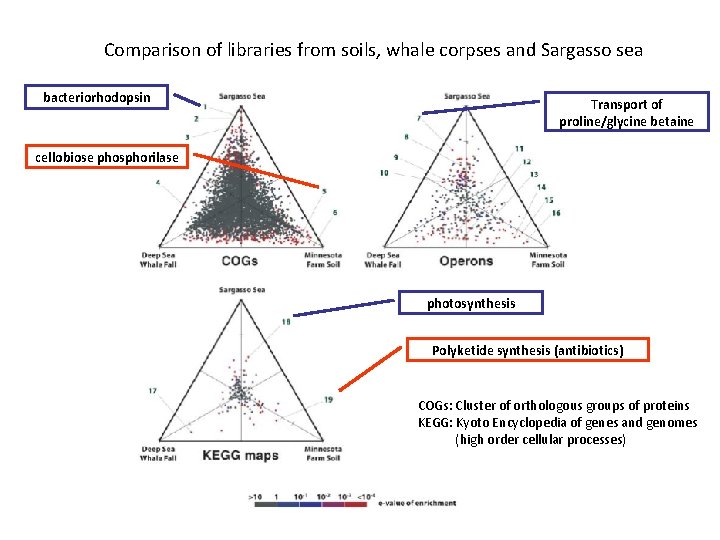
Comparison of libraries from soils, whale corpses and Sargasso sea bacteriorhodopsin Transport of proline/glycine betaine cellobiose phosphorilase photosynthesis Polyketide synthesis (antibiotics) COGs: Cluster of orthologous groups of proteins KEGG: Kyoto Encyclopedia of genes and genomes (high order cellular processes)

Functional metagenomics: search of genes expressing a function • Screening of metagenomic libraries to search for a particular function (resistance to some compounds, fluorescence, etc). • Many compounds like antibiotics, quorum sensing inhibitors or inducers, enzymes of commercial interest, pigments, etc, have been discovered. The ISME Journal, 9 October 2008; Functional metagenomics reveals diverse b -lactamases in a remote Alaskan soil Heather K Allen 1, 2, Luke A Moe 1, Jitsupang Rodbumrer 1, 3, Andra Gaarder 1 and Jo Handelsman 1

2. Functional metagenomic approach to search for novel mechanisms of adaptation to extreme environments

Study of life in extreme environments Which are the limits of life? Search for novel molecular mechanisms of adaptation of the microorganisms to extreme conditions (toxic metales, acidic p. H, low and high temperatures, high radiation and high salt concentrations) Biotechnological aplications, bioremediation, biomining… Bias in the known mechanisms of adaptation, most from cultured microorganisms Functional Metagenomic approach (culture independent)

OUTLINE 1. Search for metal resistance genes in microorganisms from the Río Tinto • Nickel resistance genes from rhizosphere communities 2. Search for acid p. H resistance genes in microorganisms from the Río Tinto 3. Construction of nickel resistant transgenic plants 4. Future: search for adaptation mechanisms in microorganisms from rhizosphere and phyllosphere of Antartic plants, and from hypersaline environments
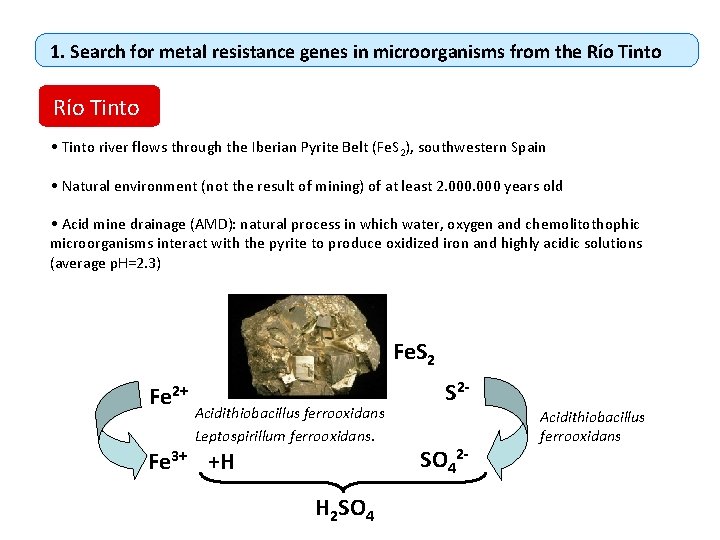
1. Search for metal resistance genes in microorganisms from the Río Tinto • Tinto river flows through the Iberian Pyrite Belt (Fe. S 2), southwestern Spain • Natural environment (not the result of mining) of at least 2. 000 years old • Acid mine drainage (AMD): natural process in which water, oxygen and chemolitothophic microorganisms interact with the pyrite to produce oxidized iron and highly acidic solutions (average p. H=2. 3) Fe. S 2 Fe 2+ Acidithiobacillus ferrooxidans Leptospirillum ferrooxidans. Fe 3+ +H H 2 SO 4 S 2 SO 42 - Acidithiobacillus ferrooxidans

Acid water and oxidation increase the solubility of other metals and metalloids As 380 ppm Cr 380 ppm Cu Zn 110 ppm 220 ppm Ni 10 ppm Complex microbial communities. (High diversity of eukaryotes, but low diversity of bacteria and archaea in the planktonic phase)

Metagenomic libraries • planktonic phase: highly enriched in toxic metals, very low p. H, low bacterial diversity (less than ten species) • rhizosphere from the endemic heather, Erica andevalensis: less enriched in heavy metals, p. H ~ 4 -5, high bacterial diversity (root exudates are enriched in nutrients)
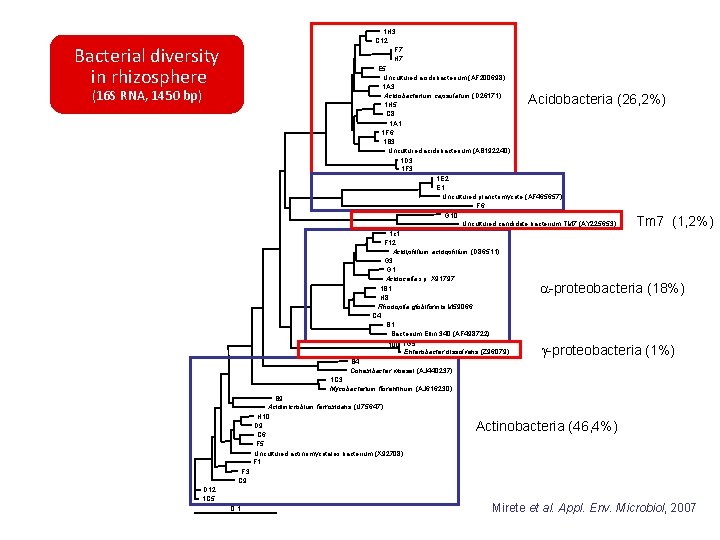
1 H 3 C 12 F 7 H 7 E 5 Uncultured acidobacterium (AF 200698) 1 A 3 Acidobacterium capsulatum (D 26171) 1 H 5 C 8 1 A 1 1 F 6 1 B 3 Uncultured acidobacterium (AB 192240) 1 D 3 1 F 3 1 E 2 E 1 Uncultured planctomycete (AF 465657) F 6 G 10 Uncultured candidate bacterium TM 7 (AY 225653) 1 c 1 F 12 Acidiphilium acidophilum (D 86511) G 3 G 1 Acidocella sp. X 91797 1 B 1 H 8 Rhodopila globiformis M 59066 C 4 B 1 Bacterium Ellin 340 (AF 498722) 100 1 G 5 Enterobacter dissolvens (Z 96079) Bacterial diversity in rhizosphere (16 S RNA, 1450 bp) Acidobacteria (26, 2%) Tm 7 (1, 2%) a-proteobacteria (18%) B 4 Conexibacter woesei (AJ 440237) g-proteobacteria (1%) 1 C 3 Mycobacterium florentinum (AJ 616230) B 9 Acidimicrobium ferroxidans (U 75647) H 10 D 9 C 6 F 5 Uncultured actinomycetales bacterium (X 92708) F 1 Actinobacteria (46, 4%) F 3 C 9 D 12 1 C 5 0. 1 Mirete et al. Appl. Env. Microbiol, 2007
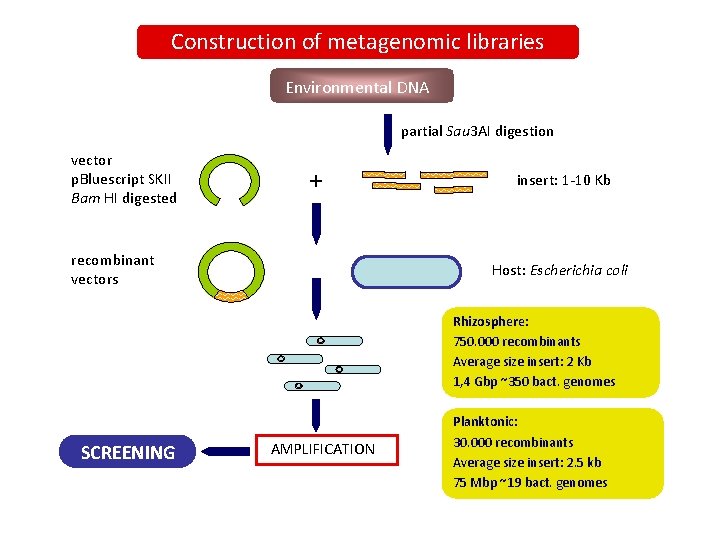
Construction of metagenomic libraries Environmental DNA partial Sau 3 AI digestion vector p. Bluescript SKII Bam HI digested + recombinant vectors insert: 1 -10 Kb Host: Escherichia coli Rhizosphere: 750. 000 recombinants Average size insert: 2 Kb 1, 4 Gbp ~350 bact. genomes SCREENING AMPLIFICATION Planktonic: 30. 000 recombinants Average size insert: 2. 5 kb 75 Mbp ~19 bact. genomes
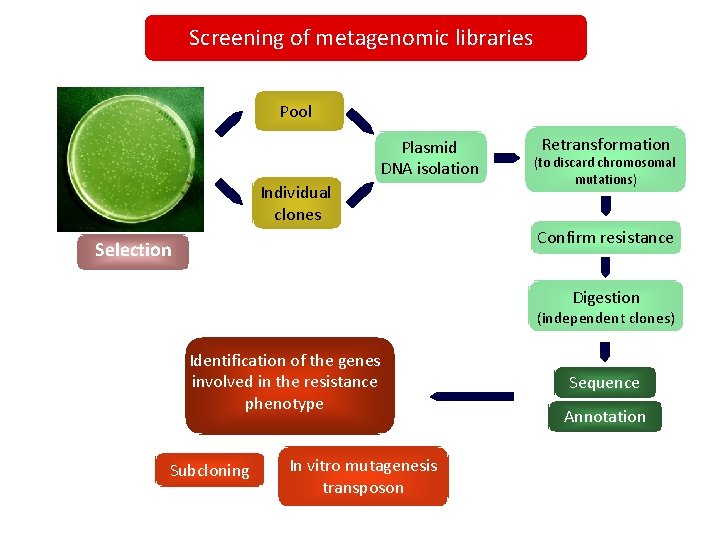
Screening of metagenomic libraries Pool Plasmid DNA isolation Individual clones Retransformation (to discard chromosomal mutations) Confirm resistance Selection Digestion (independent clones) Identification of the genes involved in the resistance phenotype Subcloning In vitro mutagenesis transposon Sequence Annotation
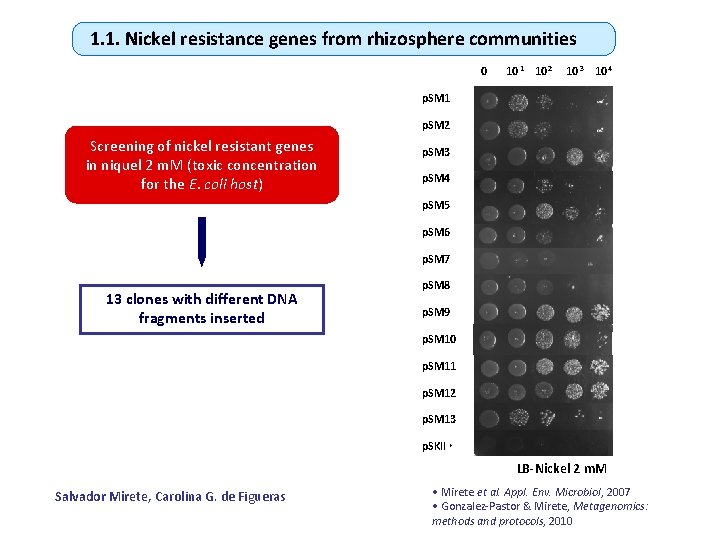
1. 1. Nickel resistance genes from rhizosphere communities 0 10 -1 10 -2 10 -3 10 -4 p. SM 1 p. SM 2 Screening of nickel resistant genes in niquel 2 m. M (toxic concentration for the E. coli host) p. SM 3 p. SM 4 p. SM 5 p. SM 6 p. SM 7 13 clones with different DNA fragments inserted p. SM 8 p. SM 9 p. SM 10 p. SM 11 p. SM 12 p. SM 13 p. SKII + LB-Nickel 2 m. M Salvador Mirete, Carolina G. de Figueras • Mirete et al. Appl. Env. Microbiol, 2007 • Gonzalez-Pastor & Mirete, Metagenomics: methods and protocols, 2010
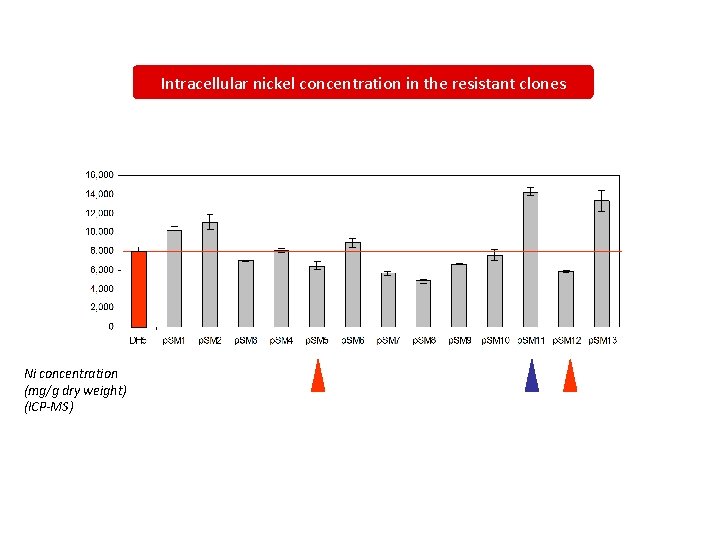
Intracellular nickel concentration in the resistant clones Ni concentration (mg/g dry weight) (ICP-MS)
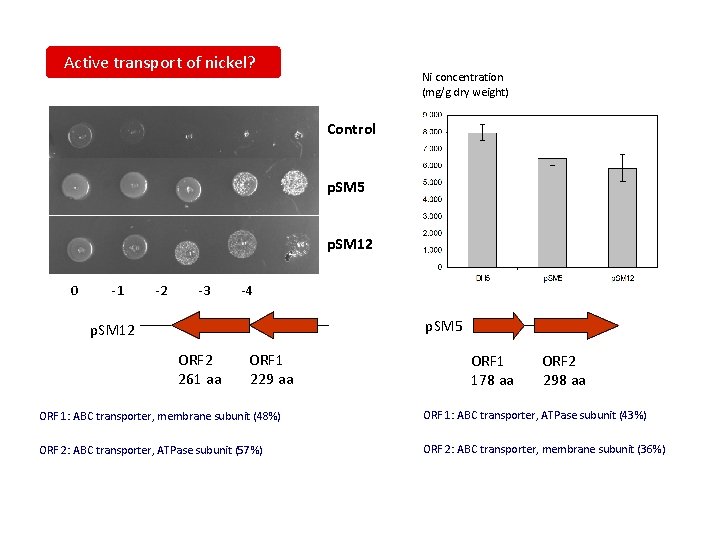
Active transport of nickel? Ni concentration (mg/g dry weight) Control p. SM 5 p. SM 12 0 -1 -2 -3 -4 p. SM 5 p. SM 12 ORF 2 261 aa ORF 1 229 aa ORF 1 178 aa ORF 2 298 aa ORF 1: ABC transporter, membrane subunit (48%) ORF 1: ABC transporter, ATPase subunit (43%) ORF 2: ABC transporter, ATPase subunit (57%) ORF 2: ABC transporter, membrane subunit (36%)
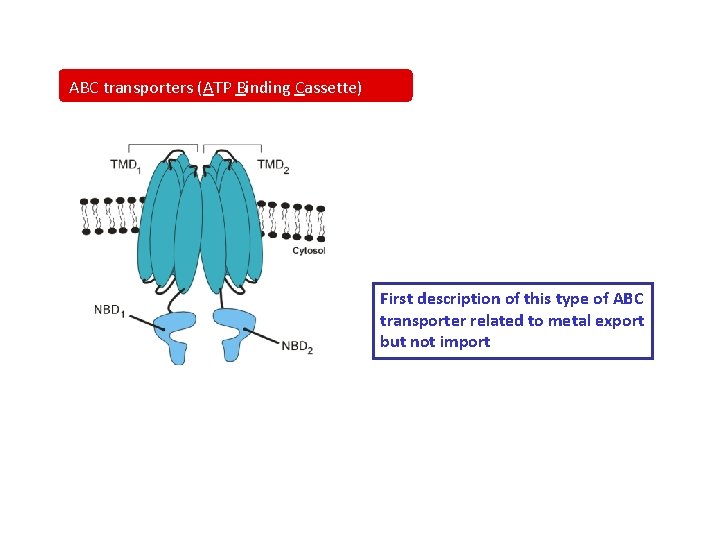
ABC transporters (ATP Binding Cassette) First description of this type of ABC transporter related to metal export but not import
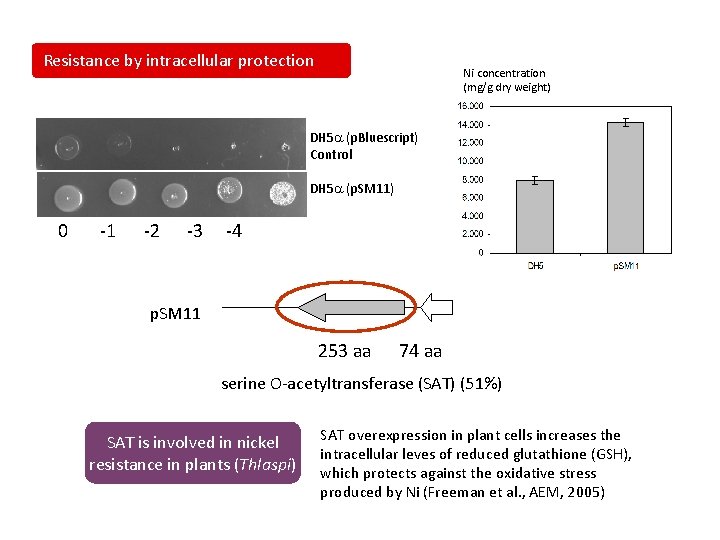
Resistance by intracellular protection Ni concentration (mg/g dry weight) DH 5 a (p. Bluescript) Control DH 5 a (p. SM 11) 0 -1 -2 -3 -4 p. SM 11 253 aa 74 aa serine O-acetyltransferase (SAT) (51%) SAT is involved in nickel resistance in plants (Thlaspi) SAT overexpression in plant cells increases the intracellular leves of reduced glutathione (GSH), which protects against the oxidative stress produced by Ni (Freeman et al. , AEM, 2005)
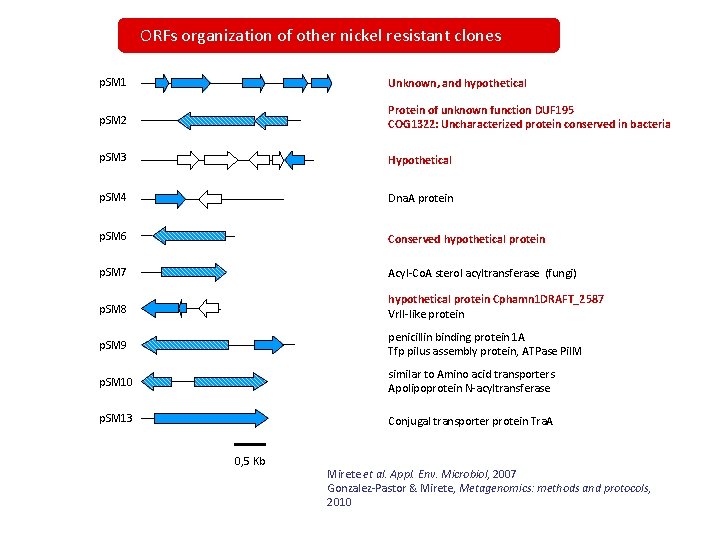
ORFs organization of other nickel resistant clones p. SM 1 Unknown, and hypothetical p. SM 2 Protein of unknown function DUF 195 COG 1322: Uncharacterized protein conserved in bacteria p. SM 3 Hypothetical p. SM 4 Dna. A protein p. SM 6 Conserved hypothetical protein p. SM 7 Acyl-Co. A sterol acyltransferase (fungi) p. SM 8 hypothetical protein Cphamn 1 DRAFT_2587 Vrl. I-like protein p. SM 9 penicillin binding protein 1 A Tfp pilus assembly protein, ATPase Pil. M p. SM 10 similar to Amino acid transporters Apolipoprotein N-acyltransferase p. SM 13 Conjugal transporter protein Tra. A 0, 5 Kb Mirete et al. Appl. Env. Microbiol, 2007 Gonzalez-Pastor & Mirete, Metagenomics: methods and protocols, 2010
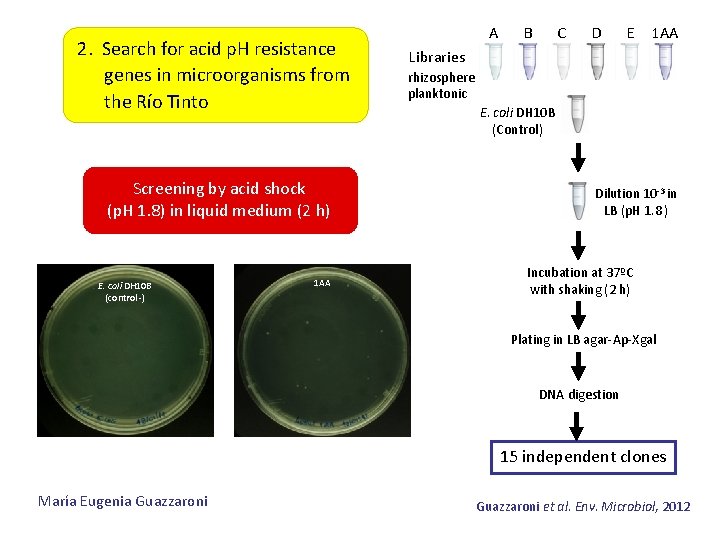
2. Search for acid p. H resistance genes in microorganisms from the Río Tinto Screening by acid shock (p. H 1. 8) in liquid medium (2 h) E. coli DH 10 B (control -) 1 AA A B C D E 1 AA Libraries rhizosphere planktonic E. coli DH 10 B (Control) Dilution 10 -3 in LB (p. H 1. 8 ) Incubation at 37ºC with shaking (2 h) Plating in LB agar-Ap-Xgal DNA digestion 15 independent clones María Eugenia Guazzaroni et al. Env. Microbiol, 2012
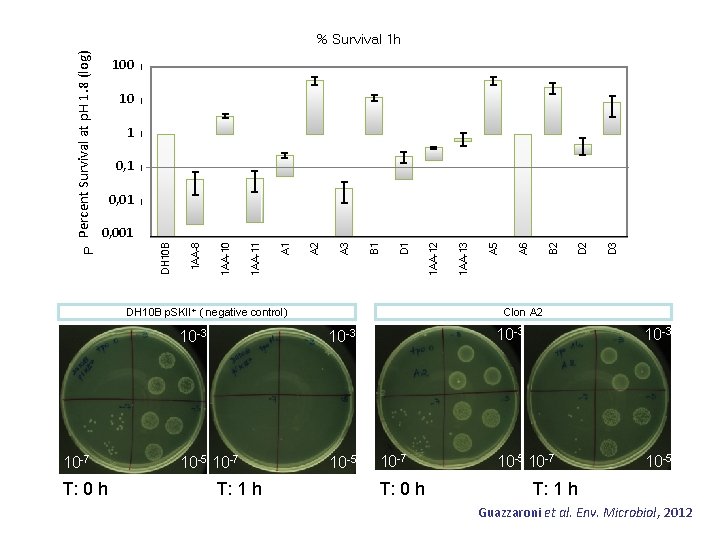
100 100. 000 10 10. 000 1 1. 000 0, 1 0. 100 T: 0 h 10 -3 10 -5 10 -7 10 -5 T: 1 h D 3 D 2 B 2 A 6 A 5 1 AA-13 Clon A 2 DH 10 B p. SKII+ ( negative control) 10 -7 1 AA-12 D 1 B 1 A 3 A 2 A 1 1 AA-10 0, 001 0. 001 1 AA-8 0, 01 0. 010 DH 10 B Percent Survival 1. 8 (log) Percent Survival at p. Hat 1. 8 p. H (log) % Survival 1 h 10 -7 T: 0 h 10 -3 10 -5 10 -7 10 -5 T: 1 h Guazzaroni et al. Env. Microbiol, 2012
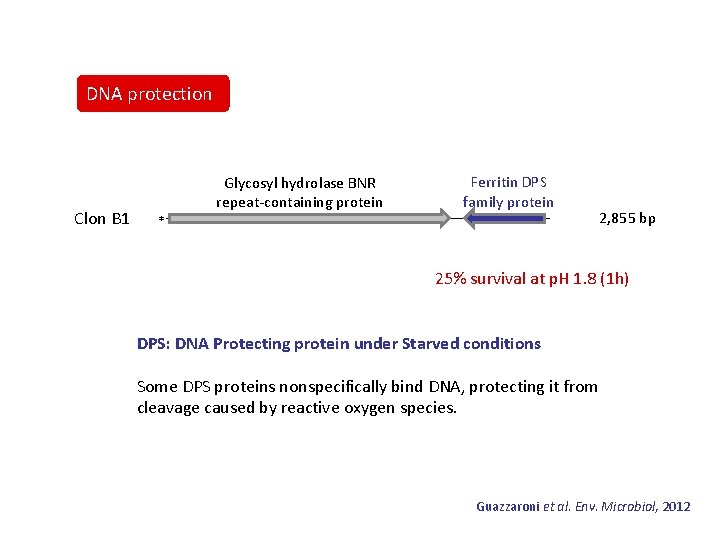
DNA protection Clon B 1 Glycosyl hydrolase BNR repeat-containing protein Ferritin DPS family protein * 2, 855 bp 25% survival at p. H 1. 8 (1 h) DPS: DNA Protecting protein under Starved conditions Some DPS proteins nonspecifically bind DNA, protecting it from cleavage caused by reactive oxygen species. Guazzaroni et al. Env. Microbiol, 2012

A chaperon involved in acid p. H resistance ATP-dependent Clp protease, ATP-binding proteolytic subunit Clp. P subunit Clp. X Clon B 2 1, 701 bp * 32 % survival at p. H 1. 8 (1 h) Clp. PX: a two component protease involved in removing heat-damaged proteins (heat shock). Not previously reported to be involved in acid p. H tolerance • Clp. P is the proteolytic subunit • Clp. X is the ATP-binding subunit and works as a molecular chaperone. Guazzaroni et al. Env. Microbiol, 2012
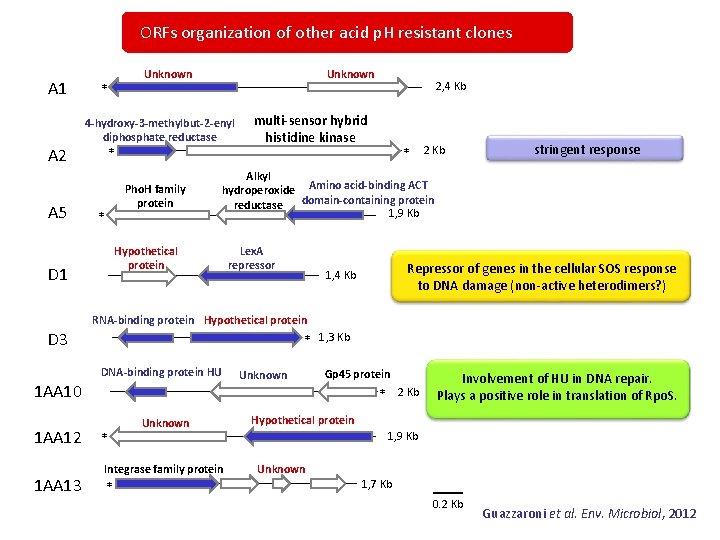
ORFs organization of other acid p. H resistant clones A 1 A 2 A 5 Unknown * Unknown 4 -hydroxy-3 -methylbut-2 -enyl diphosphate reductase * Pho. H family protein * multi-sensor hybrid histidine kinase * 2 Kb stringent response Alkyl hydroperoxide Amino acid-binding ACT domain-containing protein reductase 1, 9 Kb Hypothetical protein D 1 2, 4 Kb Lex. A repressor Repressor of genes in the cellular SOS response to DNA damage (non-active heterodimers? ) 1, 4 Kb RNA-binding protein Hypothetical protein D 3 * 1, 3 Kb DNA-binding protein HU 1 AA 10 1 AA 12 1 AA 13 Unknown Gp 45 protein 2 Kb * Unknown Hypothetical protein 1, 9 Kb * Integrase family protein * Involvement of HU in DNA repair. Plays a positive role in translation of Rpo. S. Unknown 1, 7 Kb 0. 2 Kb Guazzaroni et al. Env. Microbiol, 2012
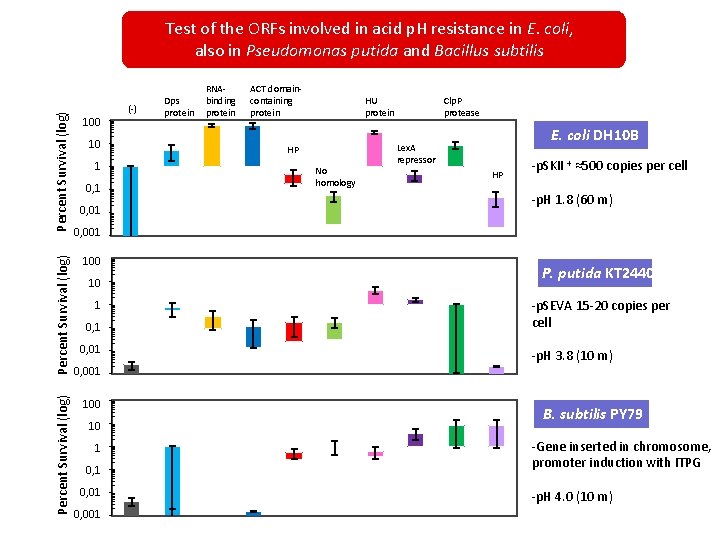
Percent Survival (log) Test of the ORFs involved in acid p. H resistance in E. coli, also in Pseudomonas putida and Bacillus subtilis 100. 000 100 Percent Survival (log) 10. 000 10 1. 0001 0, 1 0. 100 0, 01 0. 010 RNAbinding protein ACT domaincontaining protein HU protein HP No homology Clp. P protease E. coli DH 10 B Lex. A repressor HP -p. SKII + ≈500 copies per cell -p. H 1. 8 (60 m) 0, 001 0. 001 100 10. 000 10 1 1. 000 0, 1 0. 100 0, 01 0. 010 0, 001 0. 001 100. 000 100 Percent Survival (log) (-) Dps protein 10. 000 10 P. putida KT 2440 -p. SEVA 15 -20 copies per cell -p. H 3. 8 (10 m) B. subtilis PY 79 0, 1 0. 100 -Gene inserted in chromosome, promoter induction with ITPG 0, 01 0. 010 -p. H 4. 0 (10 m) 1. 000 1 0, 001 0. 001
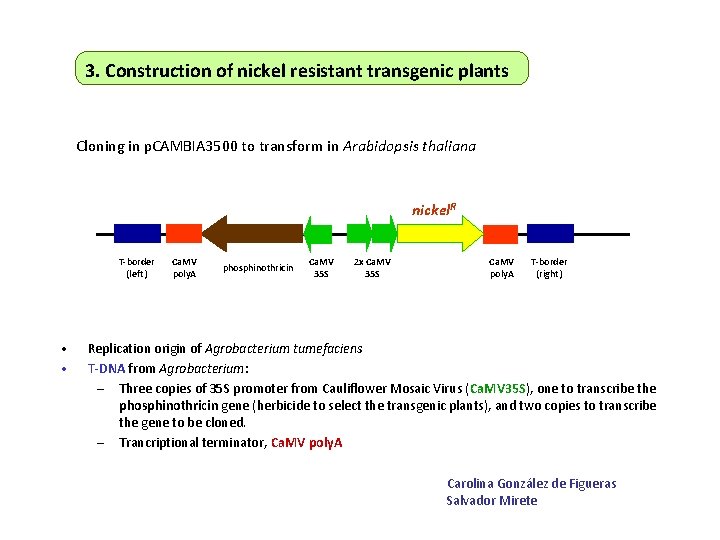
3. Construction of nickel resistant transgenic plants Cloning in p. CAMBIA 3500 to transform in Arabidopsis thaliana nickel. R T-border (left) • • Ca. MV poly. A phosphinothricin Ca. MV 35 S 2 x Ca. MV 35 S Ca. MV poly. A T-border (right) Replication origin of Agrobacterium tumefaciens T-DNA from Agrobacterium: – Three copies of 35 S promoter from Cauliflower Mosaic Virus (Ca. MV 35 S), one to transcribe the phosphinothricin gene (herbicide to select the transgenic plants), and two copies to transcribe the gene to be cloned. – Trancriptional terminator, Ca. MV poly. A Carolina González de Figueras Salvador Mirete

3. Construction of nickel resistant transgenic plants p. SM 6: Conserved hypothetical protein p. SM 7: Acyl-Co. A sterol acyltransferase (fungi). This enzyme solubilizes the sterol from the membrane, and is accumulated in the cytoplasm. Could the Ni resistance be explained by changes in membrane permeability? Ni concentration (mg/g dry weight)
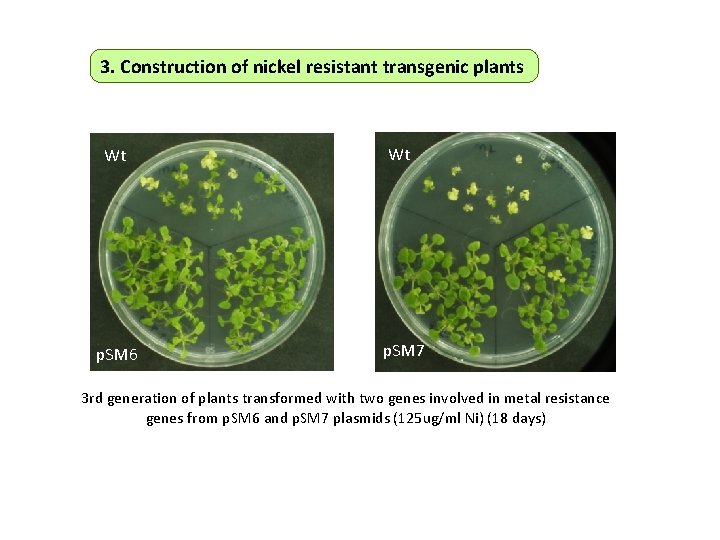
3. Construction of nickel resistant transgenic plants Wt Wt p. SM 6 p. SM 7 3 rd generation of plants transformed with two genes involved in metal resistance genes from p. SM 6 and p. SM 7 plasmids (125 ug/ml Ni) (18 days)
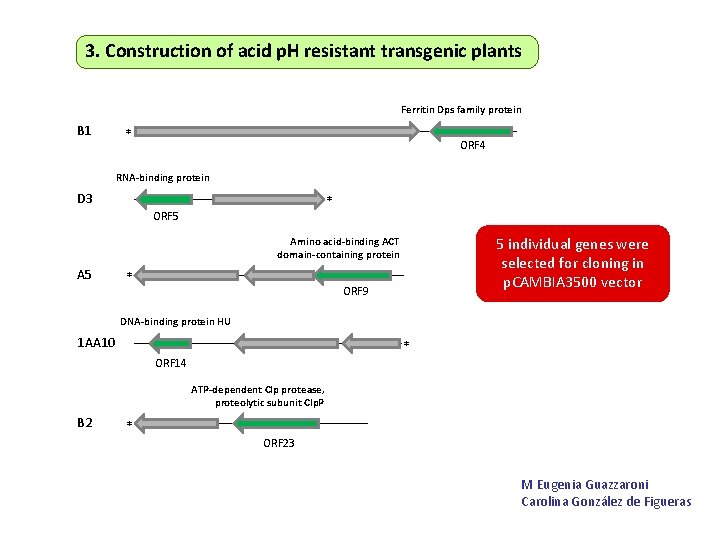
3. Construction of acid p. H resistant transgenic plants Ferritin Dps family protein B 1 * ORF 4 RNA-binding protein D 3 * ORF 5 5 individual genes were selected for cloning in p. CAMBIA 3500 vector Amino acid-binding ACT domain-containing protein A 5 * ORF 9 DNA-binding protein HU 1 AA 10 * ORF 14 ATP-dependent Clp protease, proteolytic subunit Clp. P B 2 * ORF 23 M Eugenia Guazzaroni Carolina González de Figueras

4. Search for adaptation mechanisms in microorganisms from rhizosphere and phyllosphere of Antartic plants Colobanthus quitensis Deschampsia antartica • Microbial diversity from rhizosphere and phyllosphere • Metagenomics: - sequence - funtional (genes involved in cold and radiation adaptacion) Verónica Morgante

4. Search for adaptation mechanisms in microorganisms from hypersaline environments (collaboration Ramón Rosselló-Móra) Hipersaline antarctic ponds (Bratina Island) Salt flats Añana (Spain) Coast Salt flats Boyeruca (Chile), Es Trenc (Mallorca) Rhizosphere and phyllosphere Salicornia Calonecris diomedea (nostril salt glands) • Microbial and viral diversity • Functional diversit: salt resistance, UV radiation resistance, low temperatures, etc (functional metagenomics, sequencing, and metatranscriptomic in experiments with mesocosms)

CONCLUSIONS Ø Small insert metagenomic libraries have been useful to retrieve genes involved in resistance to toxic metals and acidic p. H. - genes previously described (chaperons, transporters, DNA binding proteins…) - hypothetical and unknown genes not previously assigned to be resistant to these conditions, and now they can be annotated

The team…… Carolina González de Figueras M. Eugenia Guazzaroni Salvador Mirete Castañeda Verónica Morgante Maria Lamprecht Olga Zafra Collaborators from CAB Manuel Gómez Marina Postigo M. Paz Martín
- Slides: 50