Lecture 14 RNA Seq analysis Based on Anders

Lecture 14 RNA – Seq analysis Based on Anders and Huber paper

Some background • Cole (one of my ex students who was working on RNA-seq) said: • “RNA sequencing is a high-throughput tool for investigating gene expression, made possible with rapid advances in the speed and efficiency of sequencing technologies. • Unlike microarrays, RNA-seq benefits from a highly dynamic range of signal detection, identifying both rare and common transcripts with no a priori knowledge of the organism’s genome or transcriptome. • The additional information captured in RNA-seq libraries has revolutionized our understanding of cancer, stem cell differentiation, and plant genetics. ”

More Background • An RNA-seq run reads and quantifies the transcriptome (complete set of m. RNA) in a single sequencing run. • RNA is extracted from tissue, cleaved into fragments a few hundred nucleotides long, and then converted to a complementary DNA (c. DNA) library (Wilhelm & Landry, 2009). • Sequencing adaptors are ligated to both ends of each fragment, and the products are sequenced using any highthroughput method such as 454, SOLi. D, or Ion Torrent.

Comparison with Microarrays: advantages • New sequences can be discovered. • RNA-seq, on the other hand, determines all sequences empirically. • This has proved invaluable in non-model species with large genomes, • False positives from cross-hybridization are not an issue in RNA-seq. • Quantification is possible even at extremely low and high expression levels. • Whereas microarrays have a dynamic range of one to a few hundred fold, RNA-seq boasts a dynamic range of >8, 000 fold (Wang, Gerstein, & Snyder, 2009).

Comparison with Microarrays: disadvantages • Considerably more processing power is required to handle millions of RNA-seq reads, and chemical manipulation of RNA and c. DNA can introduce artifacts. • Slower than microarrays when the genome is known. But as sequencing costs have plummeted and computing power has increased, RNA-seq is now the transcriptomics method of choice for most applications.
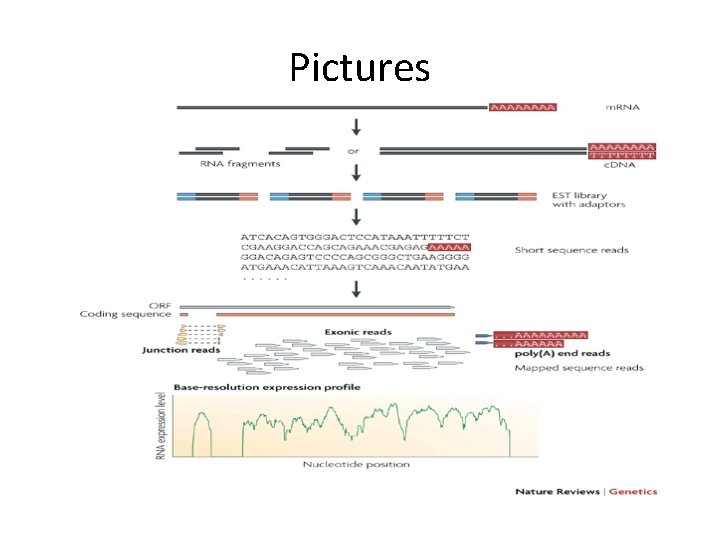
Pictures

Data structure • So here you have a sequence and for each sequence you have the number of READS • The data is the COUNT of the sequences read. • NOT continuous like expression data • So, normal and other related distributions cannot be used. • General modeling is done using the Poisson distribution

Poisson Distribution • Generally used to model count data • The mass function is given by • P(Y=y)=f(y)= • • • Properties: Has a range from 0 to positive infinity Mean, E(Y)= m Variance, = m Hence, mean and Variance are same.

Issues with Poisson • The property that requires that mean and Variance are the same is problematic for RNA-seq data, where Variance is often much larger than the mean. • This is called the over-dispersion problem. • Common in litter studies where over-dispersion is induced by auto-correlation.

Solutions: The NB Distribution • To try and address this question ne distribution that is used is the Negative Binomial Distribution. • It is used to model the number of trials till the rth success and is related to the geometric distribution. Model: P(Y=y)=f(y) =

Properties of the NB distribution So, the mean and variance are related by a proportionality constant

Theoretical Background • To model over-dispersion in Poisson regression one generally adds a random effect qi to represent the unobserved heterogenity. • So the conditional distribution of Yi given qi is indeed Poisson with mean and variance miqi. • Idea is: if we knew and observed qi the data would be Poisson. But, we don’t know it, so if we assume a assume that qi has a gamma distribution with both parameters a=b=1/s 2 which represents the variance of the unobserved. • Then the unconditional distribution is given by:

Theory: • The form Is of negative binomial r=a, p= b/(m+b) The mean and variance are related with a proportionality constant. • This is the form used in the Anders and Huber paper laying the basic theory for D-seq. • •

The Theory of DESeq 2 • plot. MA(res, main="DESeq 2", ylim=c(-2, 2))

Theory •

The Data set • • Steps for analysis: We have a set of FASTQ files that contain the nucleotide sequence of each read and a quality score at each position. Our first task is to align the reads to a reference genome or transcriptome. It is important to know if the sequencing experiment was single-end or paired-end, as the alignment software will require the user to specify both FASTQ files for a paired-end experiment. The output of this alignment step is commonly stored in a file format called SAM/BAM. R can read the BAM files A number of software programs exist to align reads to a reference genome. Pick the ones that are relevant and used in the field of choice

Data Prep: • • Example of such an aligner is STAR. We should create and have a file in the current directory called ”files” with each line containing an identifier for each experiment. We put all the FASTQ files in a subdirectory fastq. If we have performed a single-end experiment, we would only have one file per ID else we would have two files We should create subdirectory, aligned, where STAR or other aligners will output its alignment files. The alignment and conversion is done outside the R environment. For R we will assume we have the BAM files and the descriptor files

Graphs and results from DESeq 2 Using Airway data from airway library in R

Highlights of DESeq 2 • 1. estimation of size factors sj by estimate. Size. Factors • 2. estimation of dispersion αi by estimate. Dispersions • 3. negative binomial GLM fitting for βi and Wald statistics by nbinom. Wald. Test • Where: – Kij ∼ NB(µij, αi) – µij = sjqij – log 2(qij) = xj. βi
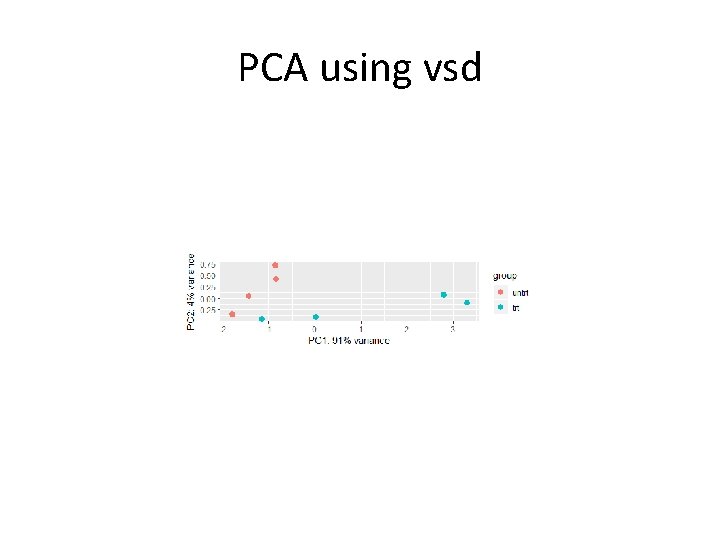
PCA using vsd
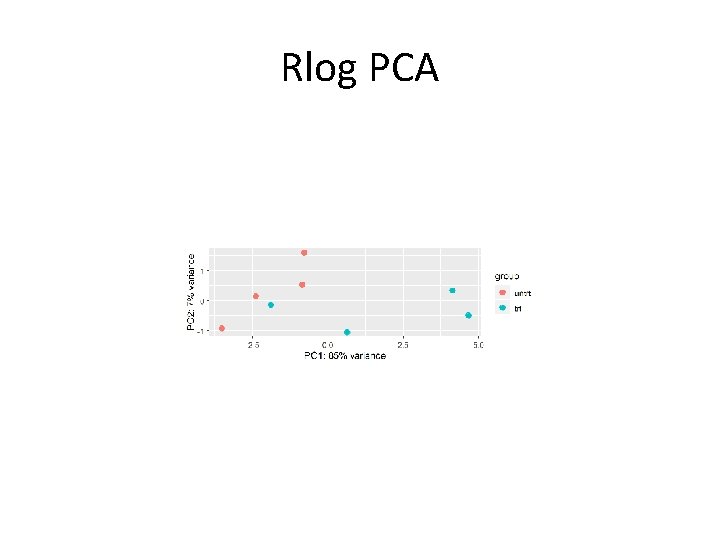
Rlog PCA

• > summary(res) out of 16 with nonzero total read count adjusted p-value < 0. 1 LFC > 0 (up) : 2, 12% LFC < 0 (down) : 2, 12% outliers [1] : 0, 0% low counts [2] : 0, 0% (mean count < 0)
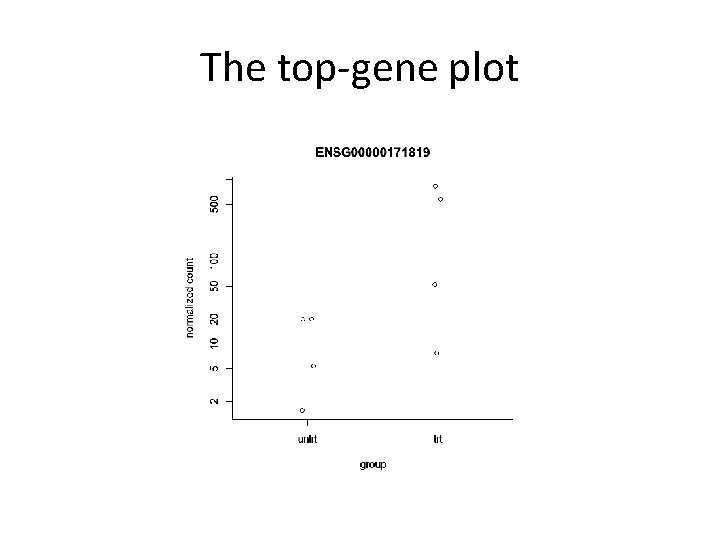
The top-gene plot

Heat Map for top 30
- Slides: 24