seq Record object from scratch from Bio Seq

seq. Record object from scratch from Bio. Seq import Seq simple_seq = Seq("GATC") from Bio. Seq. Record import Seq. Record simple_seq_r = Seq. Record(simple_seq) simple_seq_r. id = (“ 0001”) simple_seq_r. name = (“MFG 1”) simple_seq_r. description = "Made up sequence” print simple_seq_r reading the information from Bio import Seq. IO record = Seq. IO. read("NC_005816. fna", "fasta") print record

quickstart: parsing sequences Simple example: file format from Bio import Seq. IO for seq_record in Seq. IO. parse("ls_orchid. fasta", "fasta"): print(seq_record. id) print(repr(seq_record. seq)) print(len(seq_record))

Sequence I/O Parsing from the web from Bio import Entrez from Bio import Seq. IO Entrez. email = "A. N. Other@example. com" handle = Entrez. efetch(db="nucleotide", rettype="fasta", retmode="text", id="6273291") seq_record = Seq. IO. read(handle, "fasta") handle. close() print("%s with %i features" % (seq_record. id, len(seq_record. features)))

Sequence I/O How to find sequence information from Bio import Seq. IO orchid_dict = Seq. IO. to_dict(Seq. IO. parse("ls_orchid. fasta", ”fasta")) creates Python dictionary with each entry held as a Seq. Record object in memory
Sequence I/O Writing a file from Bio. Seq import Seq from Bio. Seq. Record import Seq. Record from Bio. Alphabet import generic_protein Creating seq. Rec rec 1 = Seq. Record(Seq("MMYQQGCFAGGTVLRLAKDLAENNRGARVLVVCSEITAVTFRGPSETHLDSMVGQALFGD" +"GAGAVIVGSDPDLSVERPLYELVWTGATLLPDSEGAIDGHLREVGLTFHLLKDVPGLISK" +"NIEKSLKEAFTPLGISDWNSTFWIAHPGGPAILDQVEAKLGLKEEKMRATREVLSEYGNM" +"SSAC", generic_protein), id="gi|14150838|gb|AAK 54648. 1|AF 376133_1", description="chalcone synthase [Cucumis sativus]") rec 2 = Seq. Record(Seq("YPDYYFRITNREHKAELKEKFQRMCDKSMIKKRYMYLTEEILKENPSMCEYMAPSLDARQ" +"DMVVVEIPKLGKEAAVKAIKEWGQ", generic_protein), id="gi|13919613|gb|AAK 33142. 1|", description="chalcone synthase [Fragaria vesca subsp. bracteata]") my_records = [rec 1, rec 2, rec 3] Seq. IO. write(my_records, "my_example. faa", "fasta") writing to file
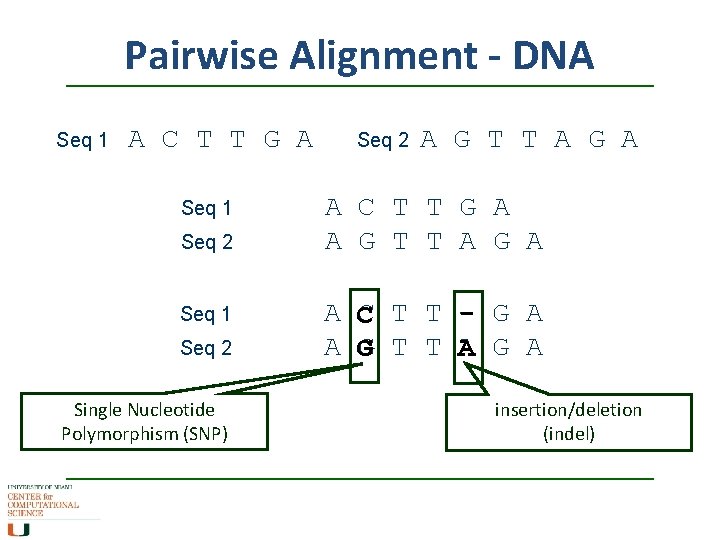
Pairwise Alignment - DNA Seq 1 A C T T G A Seq 1 Seq 2 Single Nucleotide Polymorphism (SNP) Seq 2 A G T T A G A A C T T G A A G T T A G A A C T T - G A A G T T A G A insertion/deletion (indel)
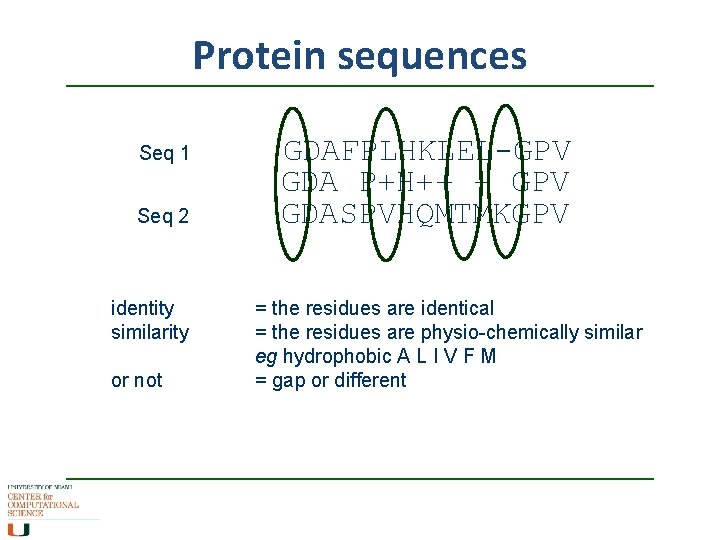
Protein sequences Seq 1 Seq 2 identity similarity or not GDAFPLHKLEL-GPV GDA P+H++ + GPV GDASPVHQMTMKGPV = the residues are identical = the residues are physio-chemically similar eg hydrophobic A L I V F M = gap or different

Describing similarity The sequences are identical 100% similarity The sequences are similar (protein) there is some level of similarity this can range from 99% to 45% even 20% use of adjectives: highly, significantly, somewhat The sequences are homologous implies an evolutionary relationship ie they have a common ancestor requires knowledge of protein or species phylogeny
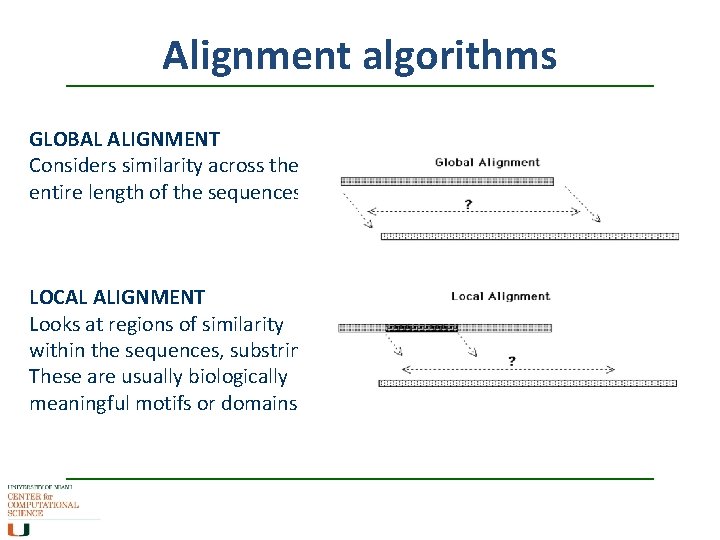
Alignment algorithms GLOBAL ALIGNMENT Considers similarity across the entire length of the sequences LOCAL ALIGNMENT Looks at regions of similarity within the sequences, substrings. These are usually biologically meaningful motifs or domains

Original manual method Start with a 2 D matrix on which you put the query sequence (I) on the vertical, and the test sequence (J) on the horizontal axis. G D I H I F Find the common k-words. G GDIHIF GDVHIF D V H I F
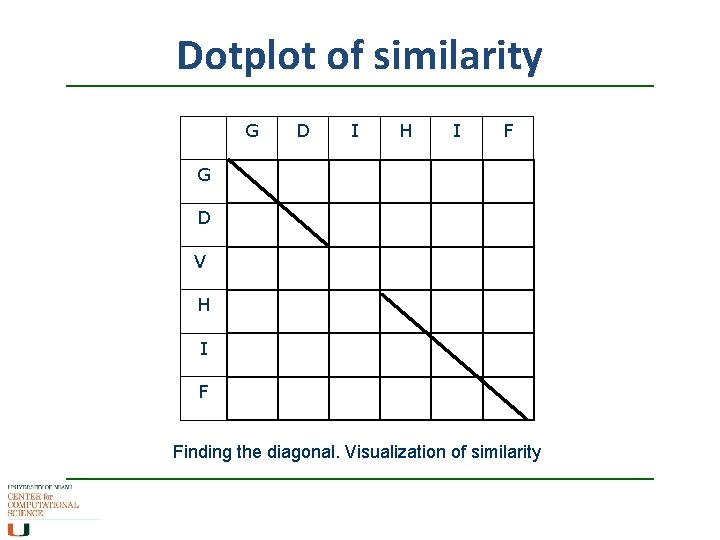
Dotplot of similarity G D I H I F G D V H I F Finding the diagonal. Visualization of similarity

Doesn’t scale well GDIHAL GRVHIF G D I H A L G R V - Visualization of similarity not so good with distantly related sequences - No statistical/numerical measure of similarity + H I F - Slow + +

Computational considerations Sequence orientation Character substitutions Physio-chemical properties which strand? variations protein structure => Biological meaning? Need a system whereby substitutions and gaps are weighted. Different types of scoring systems can be used with different algorithms – The main problem is computational time.

Comparing Strings Called “edit distance” measurements. Hamming distance: the number of different characters between strings of equal lengths; only allows substitutions. Levenshtein distance: the strings do not have to be the same length; allows insertions, deletions, substitutions. Applied to automated spelling corrections…

Dynamic programming A method of solving a large problem by dividing it up into smaller subproblems and solving those, then putting them together to solve the larger problem. Can often have more than one way of aligning two sequences, dynamic programming allows backtracking and testing of various options. The path that most effectively links together the subproblems into an optimal solution is selected as the final alignment. Needs to be based on statistics of scoring system and probability thresholds set May still end up with a few solutions of equal scores – Ultimately need a biologically relevant answer.

Dynamic programing A recursive algorithm To find an LCS of X and Y , we may need to find the LCSs of X and Yn-1 and of Xm-1 and Y. Furthermore, each of these subproblems has the subsubproblem of finding an LCS of Xm-1 and Yn-1. c[i, j] is the length of an LCS of the sequences Xi and Yj. The optimal substructure of the LCS problem gives the recursive formula if i = 0 or j = 0 if i, j > 0 and xi = yj if i, j > 0 and xi ≠ yj LCS … Longest common sequence
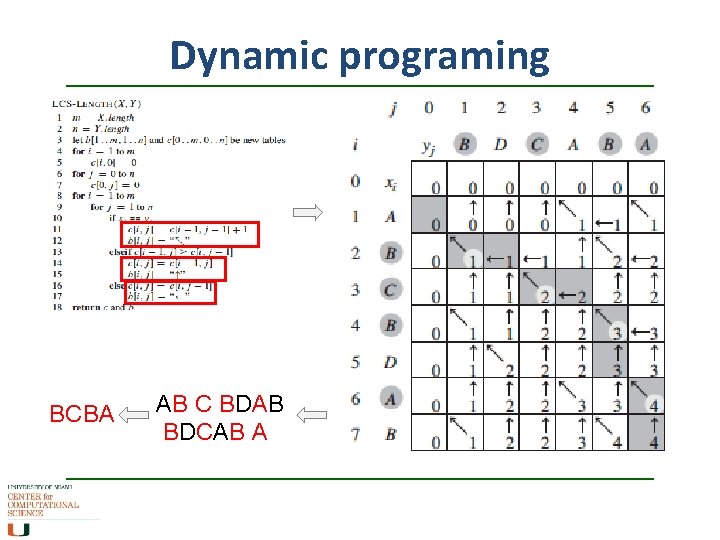
Dynamic programing BCBA AB C BDAB BDCAB A
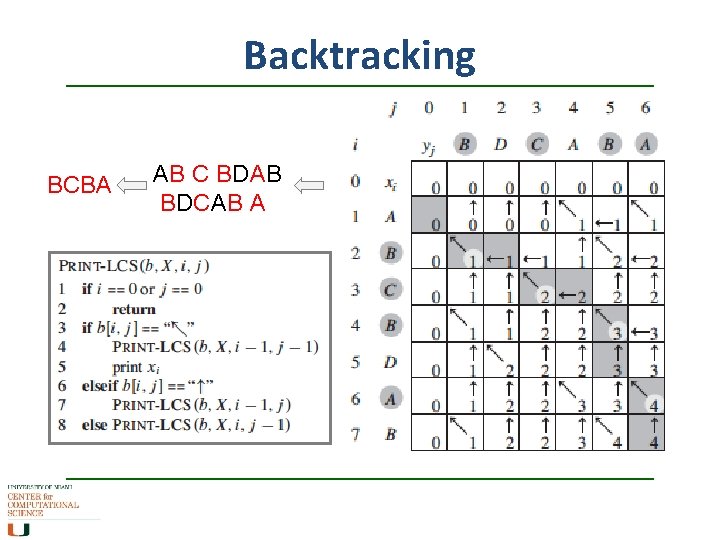
Backtracking BCBA AB C BDAB BDCAB A

Putting it all together • Create a matrix - as in dotplot but “virtual” • Calculate the match/mismatch score step by step as you go along – how many are too much? • Then backtrack and do it again – iterative, to find optimal score
- Slides: 19